6XG4
 
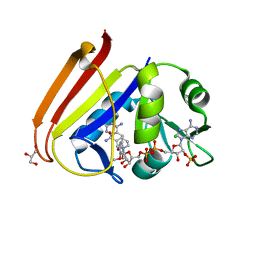 | | X-ray structure of Escherichia coli dihydrofolate reductase L28R mutant in complex with trimethoprim | | Descriptor: | CHLORIDE ION, Dihydrofolate reductase, GLYCEROL, ... | | Authors: | Gaszek, I.K, Manna, M.S, Borek, D, Toprak, E. | | Deposit date: | 2020-06-16 | | Release date: | 2021-03-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A trimethoprim derivative impedes antibiotic resistance evolution.
Nat Commun, 12, 2021
|
|
6XG5
 
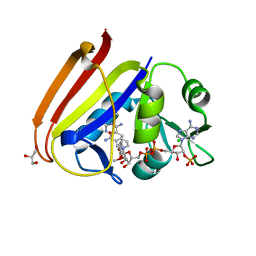 | | X-ray structure of Escherichia coli dihydrofolate reductase in complex with trimethoprim | | Descriptor: | CHLORIDE ION, Dihydrofolate reductase, GLYCEROL, ... | | Authors: | Gaszek, I.K, Manna, M.S, Borek, D, Toprak, E. | | Deposit date: | 2020-06-16 | | Release date: | 2021-03-24 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A trimethoprim derivative impedes antibiotic resistance evolution.
Nat Commun, 12, 2021
|
|
2OGG
 
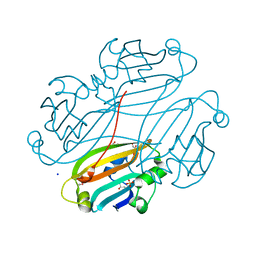 | | Structure of B. subtilis trehalose repressor (TreR) effector binding domain | | Descriptor: | GLYCEROL, SODIUM ION, SULFATE ION, ... | | Authors: | Rezacova, P, Krejcirikova, V, Borek, D, Moy, S.F, Joachimiak, A, Otwinowski, Z, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2007-01-05 | | Release date: | 2007-02-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The crystal structure of the effector-binding domain of the trehalose repressor TreR from Bacillus subtilis 168 reveals a unique quarternary assembly.
Proteins, 69, 2007
|
|
6OTC
 
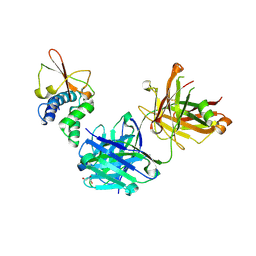 | | Synthetic Fab bound to Marburg virus VP35 interferon inhibitory domain | | Descriptor: | CHLORIDE ION, GLYCEROL, Polymerase cofactor VP35, ... | | Authors: | Amatya, P, Chen, G, Borek, D, Sidhu, S.S, Leung, D.W. | | Deposit date: | 2019-05-02 | | Release date: | 2019-06-05 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Inhibition of Marburg Virus RNA Synthesis by a Synthetic Anti-VP35 Antibody.
Acs Infect Dis., 5, 2019
|
|
1T0T
 
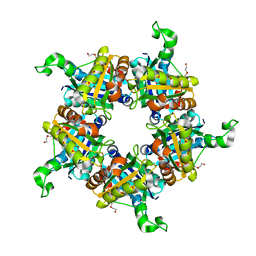 | | Crystallographic structure of a putative chlorite dismutase | | Descriptor: | 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, APC35880, MAGNESIUM ION | | Authors: | Gilski, M, Borek, D, Chen, Y, Collart, F, Joachimiak, A, Otwinowski, Z, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2004-04-12 | | Release date: | 2004-08-24 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal Structure of APC35880 protein from Bacillus Stearothermophilus
To be Published
|
|
7U5U
 
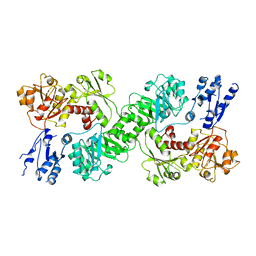 | |
7U5T
 
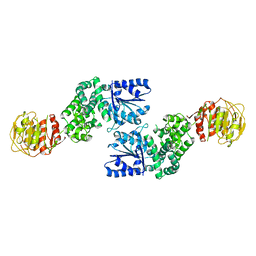 | |
7U5S
 
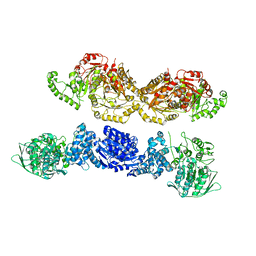 | |
3OAM
 
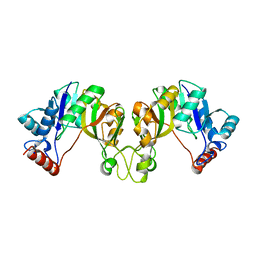 | | Crystal structure of cytidylyltransferase from Vibrio cholerae | | Descriptor: | 3-deoxy-manno-octulosonate cytidylyltransferase, SODIUM ION | | Authors: | Hattne, J, Borek, D, Grimshaw, S, Nakka, C, Rostankowski, R, Otwinowski, Z, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2010-08-05 | | Release date: | 2010-09-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structure of cytidylyltransferase from Vibrio cholerae
TO BE PUBLISHED
|
|
3P8C
 
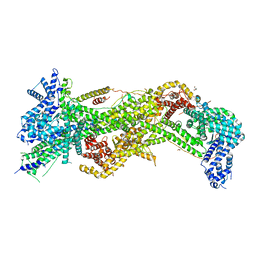 | | Structure and Control of the Actin Regulatory WAVE Complex | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Abl interactor 2, CHLORIDE ION, ... | | Authors: | Chen, Z, Borek, D, Padrick, S.B, Gomez, T.S, Metlagel, Z, Ismail, A.M, Umetani, J, Billadeau, D.D, Otwinowski, Z, Rosen, M.K. | | Deposit date: | 2010-10-13 | | Release date: | 2010-12-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Structure and control of the actin regulatory WAVE complex.
Nature, 468, 2010
|
|
3PS8
 
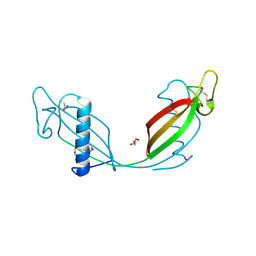 | | Crystal structure of L68V mutant of human cystatin C | | Descriptor: | ACETATE ION, Cystatin-C, DI(HYDROXYETHYL)ETHER | | Authors: | Orlikowska, M, Borek, D, Otwinowski, Z, Skowron, P, Szymanska, A. | | Deposit date: | 2010-12-01 | | Release date: | 2011-12-21 | | Last modified: | 2013-04-03 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Crystal structure of L68V mutant of human cystatin C
To be Published
|
|
3QRD
 
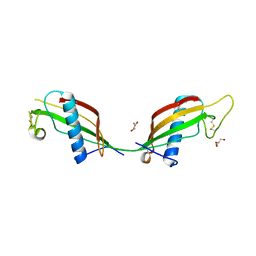 | | Crystal structure of L68V mutant of human cystatin C | | Descriptor: | Cystatin-C, DI(HYDROXYETHYL)ETHER | | Authors: | Orlikowska, M, Borek, D, Otwinowski, Z, Skowron, P, Szymanska, A. | | Deposit date: | 2011-02-17 | | Release date: | 2012-02-29 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Crystal structure of L68V mutant of human cystatin C
To be Published
|
|
2FCJ
 
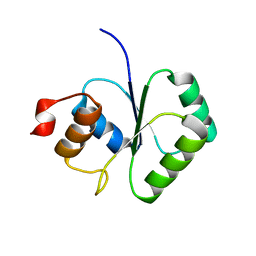 | | Structure of small TOPRIM domain protein from Bacillus stearothermophilus. | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, GLYCEROL, SULFATE ION, ... | | Authors: | Rezacova, P, Chen, Y, Borek, D, Collart, F, Joachimiak, A, Otwinowski, Z, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2005-12-12 | | Release date: | 2006-01-24 | | Last modified: | 2012-10-24 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal structure and putative function of small Toprim domain-containing protein from Bacillus stearothermophilus.
Proteins, 70, 2008
|
|
3RV5
 
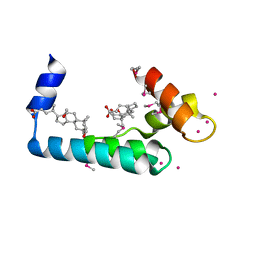 | | Crystal structure of human cardiac troponin C regulatory domain in complex with cadmium and deoxycholic acid | | Descriptor: | (3ALPHA,5BETA,12ALPHA)-3,12-DIHYDROXYCHOLAN-24-OIC ACID, CADMIUM ION, CALCIUM ION, ... | | Authors: | Li, A.Y, Lee, J, Borek, D, Otwinowski, Z, Tibbits, G, Paetzel, M. | | Deposit date: | 2011-05-06 | | Release date: | 2011-08-31 | | Last modified: | 2011-11-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of cardiac troponin C regulatory domain in complex with cadmium and deoxycholic Acid reveals novel conformation.
J.Mol.Biol., 413, 2011
|
|
3S67
 
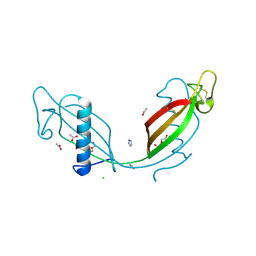 | | Crystal structure of V57P mutant of human cystatin C | | Descriptor: | ACETATE ION, CHLORIDE ION, Cystatin-C, ... | | Authors: | Orlikowska, M, Szymanska, A, Borek, D, Otwinowski, Z, Skowron, P, Jankowska, E. | | Deposit date: | 2011-05-25 | | Release date: | 2012-05-30 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Structural characterization of V57D and V57P mutants of human cystatin C, an amyloidogenic protein.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
3SVA
 
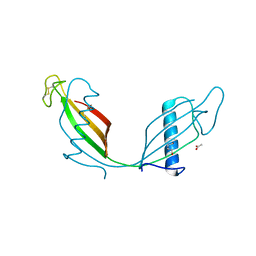 | | Crystal structure of V57D mutant of human cystatin C | | Descriptor: | ACETATE ION, Cystatin-C, DI(HYDROXYETHYL)ETHER | | Authors: | Orlikowska, M, Szymanska, A, Borek, D, Otwinowski, Z, Skowron, P, Jankowska, E. | | Deposit date: | 2011-07-12 | | Release date: | 2012-08-01 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.02 Å) | | Cite: | Structural characterization of V57D and V57P mutants of human cystatin C, an amyloidogenic protein.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
3TU3
 
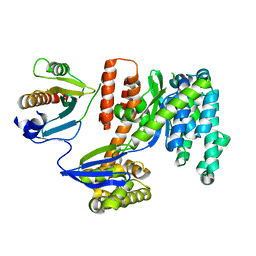 | | 1.92 Angstrom resolution crystal structure of the full-length SpcU in complex with full-length ExoU from the type III secretion system of Pseudomonas aeruginosa | | Descriptor: | ExoU, ExoU chaperone | | Authors: | Halavaty, A.S, Borek, D, Otwinowski, Z, Minasov, G, Veesenmeyer, J.L, Tyson, G, Shuvalova, L, Hauser, A.R, Anderson, W.F, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2011-09-15 | | Release date: | 2012-05-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Structure of the Type III Secretion Effector Protein ExoU in Complex with Its Chaperone SpcU.
Plos One, 7, 2012
|
|
7KOX
 
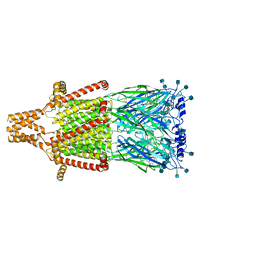 | | Alpha-7 nicotinic acetylcholine receptor bound to epibatidine and PNU-120596 in the activated state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Noviello, C.M, Hibbs, R.E, Gharpure, A, Mukhtasimova, N, Baxter, L, Cabuco, R, Borek, D, Sine, S. | | Deposit date: | 2020-11-10 | | Release date: | 2021-03-17 | | Last modified: | 2021-05-12 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Structure and gating mechanism of the alpha 7 nicotinic acetylcholine receptor.
Cell, 184, 2021
|
|
7KOO
 
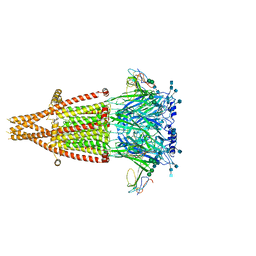 | | Alpha-7 nicotinic acetylcholine receptor bound to alpha-bungarotoxin in a resting state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Alpha-bungarotoxin isoform V31, ... | | Authors: | Noviello, C.M, Hibbs, R.E, Gharpure, A, Mukhtasimova, N, Cabuco, R, Baxter, L, Borek, D, Sine, S. | | Deposit date: | 2020-11-09 | | Release date: | 2021-03-17 | | Last modified: | 2021-05-12 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structure and gating mechanism of the alpha 7 nicotinic acetylcholine receptor.
Cell, 184, 2021
|
|
7KOQ
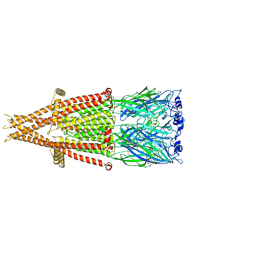 
 | | Alpha-7 nicotinic acetylcholine receptor bound to epibatidine in a desensitized state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Noviello, C.M, Hibbs, R.E, Gharpure, A, Mukhtasimova, N, Cabuco, R, Baxter, L, Borek, D, Sine, S. | | Deposit date: | 2020-11-09 | | Release date: | 2021-03-17 | | Last modified: | 2021-05-12 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structure and gating mechanism of the alpha 7 nicotinic acetylcholine receptor.
Cell, 184, 2021
|
|
2I5R
 
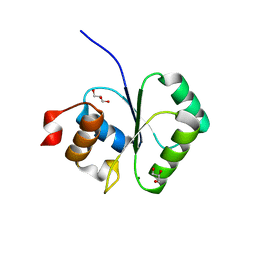 | | Structure of small Toprim domain-containing protein from B. stearothermophilus in complex with Mg2+ | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Rezacova, P, Borek, D, Otwinowski, Z, Joachimiak, A, Moy, S.F, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2006-08-25 | | Release date: | 2007-07-03 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structure and putative function of small Toprim domain-containing protein from Bacillus stearothermophilus.
Proteins, 70, 2007
|
|
7LDK
 
 | | Structure of human respiratory syncytial virus nonstructural protein 2 (NS2) | | Descriptor: | CHLORIDE ION, D(-)-TARTARIC ACID, Non-structural protein 2 | | Authors: | Chatterjee, S, Borek, D, Otwinowski, Z, Amarasinghe, G.K, Leung, D.W. | | Deposit date: | 2021-01-13 | | Release date: | 2021-03-17 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.82 Å) | | Cite: | Structural basis for IFN antagonism by human respiratory syncytial virus nonstructural protein 2.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
2IW5
 
 | | Structural Basis for CoREST-Dependent Demethylation of Nucleosomes by the Human LSD1 Histone Demethylase | | Descriptor: | AMMONIUM ION, CHLORIDE ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Yang, M, Gocke, C.B, Luo, X, Borek, D, Tomchick, D.R, Machius, M, Otwinowski, Z, Yu, H. | | Deposit date: | 2006-06-26 | | Release date: | 2006-08-09 | | Last modified: | 2019-02-06 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | Structural Basis for Corest-Dependent Demethylation of Nucleosomes by the Human Lsd1 Histone Demethylase
Mol.Cell, 23, 2006
|
|
6E59
 
 | | Crystal structure of the human NK1 tachykinin receptor | | Descriptor: | 1-(4-{[(2R,3S)-2-{(1R)-1-[3,5-bis(trifluoromethyl)phenyl]ethoxy}-3-(4-fluorophenyl)morpholin-4-yl]methyl}-1H-1,2,3-triazol-5-yl)-N,N-dimethylmethanamine, Substance-P receptor, GlgA glycogen synthase, ... | | Authors: | Yin, J, Clark, L, Chapman, K, Shao, Z, Borek, D, Xu, Q, Wang, J, Rosenbaum, D.M. | | Deposit date: | 2018-07-19 | | Release date: | 2018-12-12 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Crystal structure of the human NK1tachykinin receptor.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
3KRV
 
 | | The Structure Of Potential Metal-Dependent Hydrolase With Cyclase Activity | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, GLYCEROL, ... | | Authors: | Rakonjac, N, Rezacova, P, Borek, D, Collart, F, Joachimiak, A, Otwinowski, Z, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-11-19 | | Release date: | 2010-01-05 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | The Structure Of Potential Metal-Dependent Hydrolase With Cyclase Activity
To be Published
|
|
