8E1A
 
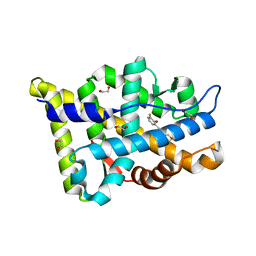 | | Structure-based study to overcome cross-reactivity of novel androgen receptor inhibitors | | Descriptor: | 1,2-ETHANEDIOL, 4-[4-(3-fluoro-2-methoxyphenyl)-1,3-thiazol-2-yl]morpholine, Androgen receptor | | Authors: | Lallous, N, Li, H, Radaeva, M, Dalal, K, Leblanc, E, Ban, F, Ciesielski, F, Chow, B, Morin, M, Singh, K, Rennie, P.S, Cherkasov, A. | | Deposit date: | 2022-08-10 | | Release date: | 2022-09-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Structure-Based Study to Overcome Cross-Reactivity of Novel Androgen Receptor Inhibitors.
Cells, 11, 2022
|
|
4HLW
 
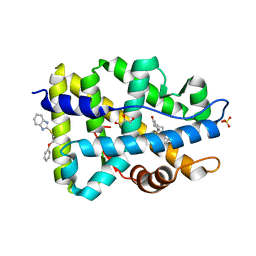 | | Targeting the Binding Function 3 (BF3) Site of the Human Androgen Receptor Through Virtual Screening. 2. Development of 2-((2-phenoxyethyl) thio)-1H-benzoimidazole derivatives. | | Descriptor: | 2-[(2-phenoxyethyl)sulfanyl]-1H-benzimidazole, Androgen receptor, GLYCEROL, ... | | Authors: | Ravi, S.N.M, Leblanc, E, Axerio-Cilies, P, Labriere, C, Frewin, K, Hassona, M.D.H, Lack, N.A, Han, F.Q, Guns, E.S, Young, R, Ban, F, Rennie, P.S, Cherkasov, A. | | Deposit date: | 2012-10-17 | | Release date: | 2013-01-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Targeting the Binding Function 3 (BF3) Site of the Androgen Receptor Through Virtual Screening. 2. Development of 2-((2-phenoxyethyl) thio)-1H-benzimidazole Derivatives.
J.Med.Chem., 56, 2013
|
|
1N57
 
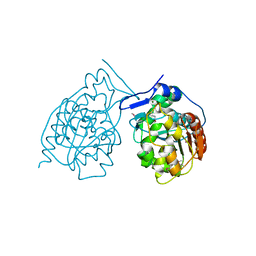 | | Crystal Structure of Chaperone Hsp31 | | Descriptor: | Chaperone Hsp31, MAGNESIUM ION | | Authors: | Quigley, P.M, Korotkov, K, Baneyx, F, Hol, W.G.J. | | Deposit date: | 2002-11-04 | | Release date: | 2003-03-18 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The 1.6A Crystal Structure of the Class of Chaperone Represented by
Escherichia coli Hsp31 Reveals a Putative Catalytic Triad
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1PV2
 
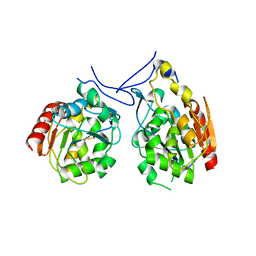 | |
2XM9
 
 | | Structure of a small molecule inhibitor with the kinase domain of Chk2 | | Descriptor: | 4-(1H-pyrazol-5-yl)-2-{4-[(3S)-pyrrolidin-3-ylamino]quinazolin-2-yl}phenol, NITRATE ION, SERINE/THREONINE-PROTEIN KINASE CHK2 | | Authors: | Caldwell, J.J, Welsh, E.J, Matijssen, C, Anderson, V.E, Antoni, L, Boxall, K, Urban, F, Hayes, A, Raynaud, F.I, Rigoreau, L.J, Raynham, T, Aherne, G.W, Pearl, L.H, Oliver, A.W, Garrett, M.D, Collins, I. | | Deposit date: | 2010-07-26 | | Release date: | 2011-01-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure-Based Design of Potent and Selective 2-(Quinazolin-2-Yl)Phenol Inhibitors of Checkpoint Kinase 2.
J.Med.Chem., 54, 2011
|
|
2XM8
 
 | | Co-crystal structure of a small molecule inhibitor bound to the kinase domain of Chk2 | | Descriptor: | 2-{4-[(3S)-PYRROLIDIN-3-YLAMINO]QUINAZOLIN-2-YL}PHENOL, SERINE/THREONINE-PROTEIN KINASE CHK2 | | Authors: | Caldwell, J.J, Welsh, E.J, Matijssen, C, Anderson, V.E, Antoni, L, Boxall, K, Urban, F, Hayes, A, Raynaud, F.I, Rigoreau, L.J, Raynham, T, Aherne, G.W, Pearl, L.H, Oliver, A.W, Garrett, M.D, Collins, I. | | Deposit date: | 2010-07-26 | | Release date: | 2011-01-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Structure-Based Design of Potent and Selective 2-(Quinazolin-2-Yl)Phenol Inhibitors of Checkpoint Kinase 2.
J.Med.Chem., 54, 2011
|
|
2BJH
 
 | | Crystal Structure Of S133A AnFaeA-ferulic acid complex | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, 3-(4-HYDROXY-3-METHOXYPHENYL)-2-PROPENOIC ACID, ... | | Authors: | Faulds, C.B, Molina, R, Gonzalez, R, Husband, F, Juge, N, Sanz-Aparicio, J, Hermoso, J.A. | | Deposit date: | 2005-02-02 | | Release date: | 2005-09-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Probing the Determinants of Substrate Specificity of a Feruloyl Esterase, Anfaea, from Aspergillus Niger
FEBS J., 272, 2005
|
|
6DLG
 
 | | Crystal structure of a SHIP1 surface entropy reduction mutant | | Descriptor: | ISOPROPYL ALCOHOL, Phosphatidylinositol 3,4,5-trisphosphate 5-phosphatase 1 | | Authors: | Gardill, B.R, Cheung, S.T, Mui, A.L, Van Petegem, F. | | Deposit date: | 2018-06-01 | | Release date: | 2019-06-05 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.499 Å) | | Cite: | Interleukin-10 and Small Molecule SHIP1 Allosteric Regulators Trigger Anti-Inflammatory Effects Through SHIP1/STAT3 Complexes
Biorxiv, 2020
|
|
8SXR
 
 | | Crystal structure of SARS-CoV-2 Mpro with C5a | | Descriptor: | 3C-like proteinase nsp5, N-[(4-chlorothiophen-2-yl)methyl]-N-[4-(dimethylamino)phenyl]-2-(5-hydroxyisoquinolin-4-yl)acetamide | | Authors: | Worrall, L.J, Kenward, C, Lee, J, Strynadka, N.C.J. | | Deposit date: | 2023-05-23 | | Release date: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.114 Å) | | Cite: | A novel class of broad-spectrum active-site-directed 3C-like protease inhibitors with nanomolar antiviral activity against highly immune-evasive SARS-CoV-2 Omicron subvariants.
Emerg Microbes Infect, 12, 2023
|
|
7KHP
 
 | | Acyl-enzyme intermediate structure of SARS-CoV-2 Mpro in complex with its C-terminal autoprocessing sequence. | | Descriptor: | 3C-like proteinase, DIMETHYL SULFOXIDE | | Authors: | Lee, J, Worrall, L.J, Paetzel, M, Strynadka, N.C.J. | | Deposit date: | 2020-10-21 | | Release date: | 2020-10-28 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystallographic structure of wild-type SARS-CoV-2 main protease acyl-enzyme intermediate with physiological C-terminal autoprocessing site.
Nat Commun, 11, 2020
|
|
7JP1
 
 | |
7JOY
 
 | |
8CYZ
 
 | | Crystal structure of SARS-CoV-2 Mpro with compound C4 | | Descriptor: | 3C-like proteinase, N-[(4-chlorothiophen-2-yl)methyl]-2-(isoquinolin-4-yl)-N-[4-(methylsulfanyl)phenyl]acetamide | | Authors: | Worrall, L.J, Lee, J, Strynadka, N.C.J. | | Deposit date: | 2022-05-24 | | Release date: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A novel class of broad-spectrum active-site-directed 3C-like protease inhibitors with nanomolar antiviral activity against highly immune-evasive SARS-CoV-2 Omicron subvariants.
Emerg Microbes Infect, 12, 2023
|
|
8CZ7
 
 | | Crystal structure of SARS-CoV-2 Mpro with compound C2 | | Descriptor: | 3C-like proteinase, N-[(4-chlorothiophen-2-yl)methyl]-2-(isoquinolin-4-yl)-N-(4-methoxyphenyl)acetamide | | Authors: | Worrall, L.J, Lee, J, Strynadka, N.C.J. | | Deposit date: | 2022-05-24 | | Release date: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A novel class of broad-spectrum active-site-directed 3C-like protease inhibitors with nanomolar antiviral activity against highly immune-evasive SARS-CoV-2 Omicron subvariants.
Emerg Microbes Infect, 12, 2023
|
|
8CYU
 
 | | Crystal structure of SARS-CoV-2 Mpro with compound C5 | | Descriptor: | 3C-like proteinase, N-[(4-chlorothiophen-2-yl)methyl]-N-[4-(dimethylamino)phenyl]-2-(isoquinolin-4-yl)acetamide | | Authors: | Worrall, L.J, Lee, J, Strynadka, N.C.J. | | Deposit date: | 2022-05-24 | | Release date: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A novel class of broad-spectrum active-site-directed 3C-like protease inhibitors with nanomolar antiviral activity against highly immune-evasive SARS-CoV-2 Omicron subvariants.
Emerg Microbes Infect, 12, 2023
|
|
8CZ4
 
 | | Crystal structure of SARS-CoV-2 Mpro with compound C3 | | Descriptor: | 3C-like proteinase, N-(4-tert-butylphenyl)-N-[(4-chlorothiophen-2-yl)methyl]-2-(isoquinolin-4-yl)acetamide | | Authors: | Worrall, L.J, Lee, J, Strynadka, N.C.J. | | Deposit date: | 2022-05-24 | | Release date: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A novel class of broad-spectrum active-site-directed 3C-like protease inhibitors with nanomolar antiviral activity against highly immune-evasive SARS-CoV-2 Omicron subvariants.
Emerg Microbes Infect, 12, 2023
|
|
6DV5
 
 | |
5I8O
 
 | | HMM5 Fab in complex with disaccharide | | Descriptor: | HMM5 antibody heavy chain, HMM5 antibody light chain, alpha-L-fucopyranose-(1-3)-methyl 2-acetamido-2-deoxy-alpha-D-glucopyranoside | | Authors: | Andersen, G.R, Tjerrild, L.M, Spillner, E. | | Deposit date: | 2016-02-19 | | Release date: | 2016-03-23 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of HMM5 Fab with a bound disaccharide
To be published
|
|
8C89
 
 | | SARS-CoV-2 spike in complex with the 17T2 neutralizing antibody Fab fragment (local refinement of RBD and Fab) | | Descriptor: | 17T2 Fab heavy chain, 17T2 Fab light chain, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Modrego, A, Carlero, D, Bueno-Carrasco, M.T, Santiago, C, Carolis, C, Arranz, R, Blanco, J, Magri, G. | | Deposit date: | 2023-01-19 | | Release date: | 2024-01-10 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (4.41 Å) | | Cite: | A monoclonal antibody targeting a large surface of the receptor binding motif shows pan-neutralizing SARS-CoV-2 activity.
Nat Commun, 15, 2024
|
|
7U0O
 
 | |
7U0M
 
 | |
2B5Z
 
 | | Hen lysozyme chemically glycosylated | | Descriptor: | (1S)-1,5-anhydro-1-(ethylsulfonyl)-D-glucitol, AZIDE ION, GLYCEROL, ... | | Authors: | Lopez-Jaramillo, F.J, Perez-Balderas, F, Hernandez-Mateo, F, Santoyo-Gonzalez, F. | | Deposit date: | 2005-09-29 | | Release date: | 2006-10-03 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Production, crystallization and X-ray characterization of chemically glycosylated hen egg-white lysozyme.
Acta Crystallogr.,Sect.F, 61, 2005
|
|
7KLI
 
 | |
7L6C
 
 | |
7K73
 
 | |
