4QZF
 
 | | Mouse Tdt, F401A mutant, in complex with a DSB substrate, C-A base pair | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*TP*TP*TP*TP*GP*A)-3', ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2014-07-27 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Embo J., 34, 2015
|
|
4QZ9
 
 | | Mouse Tdt in complex with a DSB substrate, C-A base pair | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*TP*TP*TP*TP*GP*A)-3', ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2014-07-27 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Embo J., 34, 2015
|
|
4QZE
 
 | | Mouse Tdt, F401A mutant, in complex with a DSB substrate, C-G base pair | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*TP*TP*TP*TP*GP*GP*G)-3', ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2014-07-27 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Embo J., 34, 2015
|
|
4QZI
 
 | | Mouse Tdt, F401A mutant, in complex with a DSB substrate and Zn2+ | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*TP*TP*TP*TP*GP*GP*G)-3', ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2014-07-27 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Embo J., 34, 2015
|
|
4QZ8
 
 | | Mouse Tdt in complex with a DSB substrate, C-G base pair | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*TP*TP*TP*TP*GP*G)-3', ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2014-07-27 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Embo J., 34, 2015
|
|
4QZC
 
 | | Mouse Tdt, F405A mutant, in complex with a DSB substrate, C-G base pair | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*TP*TP*TP*TP*GP*G)-3', ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2014-07-27 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Embo J., 34, 2015
|
|
4QZG
 
 | | Mouse Tdt, F401A mutant, in complex with a DSB substrate, C-C base pair | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*TP*TP*TP*TP*GP*C)-3', ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2014-07-27 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Embo J., 34, 2015
|
|
4QZA
 
 | | Mouse Tdt in complex with a DSB substrate, C-C base pair | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*TP*TP*TP*TP*GP*C)-3', ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2014-07-27 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Embo J., 34, 2015
|
|
4QZD
 
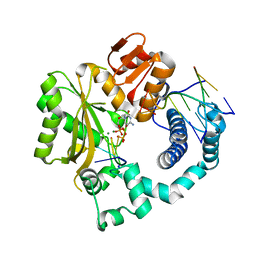 | | Mouse Tdt, F405A mutant, in complex with a DSB substrate, C-C base pair | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*TP*TP*TP*TP*GP*C)-3', ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2014-07-27 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Embo J., 34, 2015
|
|
4QZH
 
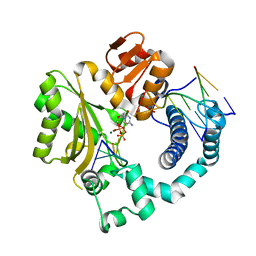 | | Mouse Tdt, F401A mutant, in complex with a DSB substrate, C-T base pair | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*TP*TP*TP*TP*GP*T)-3', ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2014-07-27 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Embo J., 34, 2015
|
|
5MUR
 
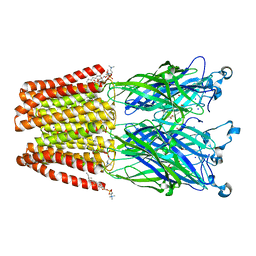 | | X-ray structure of the F14'A mutant of GLIC in complex with propofol | | Descriptor: | 2,6-BIS(1-METHYLETHYL)PHENOL, ACETATE ION, CHLORIDE ION, ... | | Authors: | Sauguet, L, Fourati, Z, Delarue, M. | | Deposit date: | 2017-01-13 | | Release date: | 2018-02-14 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural Basis for a Bimodal Allosteric Mechanism of General Anesthetic Modulation in Pentameric Ligand-Gated Ion Channels.
Cell Rep, 23, 2018
|
|
5MVM
 
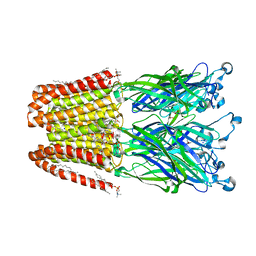 | |
5MVN
 
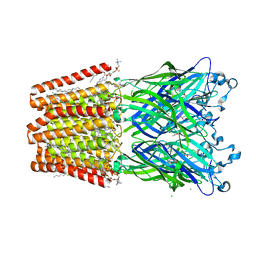 | |
5MUO
 
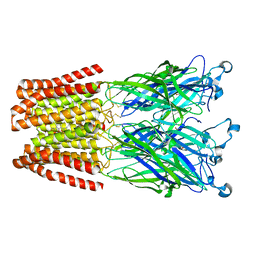 | | X-ray structure of the 2-22' locally-closed mutant of GLIC in complex with propofol | | Descriptor: | 2,6-BIS(1-METHYLETHYL)PHENOL, ACETATE ION, CHLORIDE ION, ... | | Authors: | Fourati, Z, Ruza, R.R, Delarue, M. | | Deposit date: | 2017-01-13 | | Release date: | 2018-02-14 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.19 Å) | | Cite: | Structural Basis for a Bimodal Allosteric Mechanism of General Anesthetic Modulation in Pentameric Ligand-Gated Ion Channels.
Cell Rep, 23, 2018
|
|
5MZQ
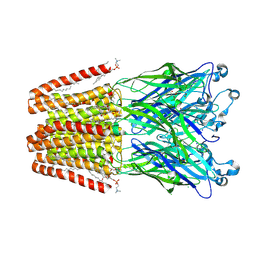 
 | | X-ray structure of the M205W mutant of GLIC in complex with bromoform | | Descriptor: | (3R)-3-(dodecanoyloxy)tetradecanoic acid, 1,2-DIOCTANOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, ACETATE ION, ... | | Authors: | Fourati, Z, Delarue, M. | | Deposit date: | 2017-02-01 | | Release date: | 2018-02-28 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural Basis for a Bimodal Allosteric Mechanism of General Anesthetic Modulation in Pentameric Ligand-Gated Ion Channels.
Cell Rep, 23, 2018
|
|
5NJY
 
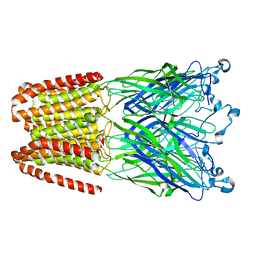 | |
5NKJ
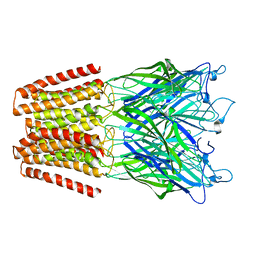 
 | | X-ray structure of the N239C mutant of GLIC | | Descriptor: | Proton-gated ion channel | | Authors: | Hu, H, Fourati, Z, Delarue, M. | | Deposit date: | 2017-03-31 | | Release date: | 2018-06-20 | | Last modified: | 2019-05-08 | | Method: | X-RAY DIFFRACTION (3.74 Å) | | Cite: | Structural Basis for a Bimodal Allosteric Mechanism of General Anesthetic Modulation in Pentameric Ligand-Gated Ion Channels.
Cell Rep, 23, 2018
|
|
5MZR
 
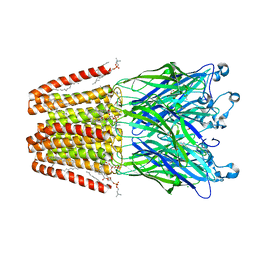 | |
5MZT
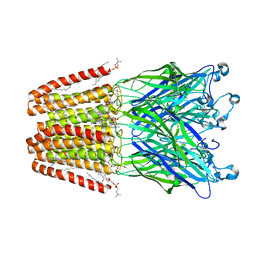 
 | |
6HMF
 
 | | D-family DNA polymerase - DP1 subunit (3'-5' proof-reading exonuclease) H451 proof-reading deficient variant | | Descriptor: | ACETATE ION, CACODYLATE ION, CALCIUM ION, ... | | Authors: | Raia, P, Delarue, M, Sauguet, L. | | Deposit date: | 2018-09-12 | | Release date: | 2019-01-30 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of the DP1-DP2 PolD complex bound with DNA and its implications for the evolutionary history of DNA and RNA polymerases.
PLoS Biol., 17, 2019
|
|
6GO3
 
 | | TdT chimera (Loop1 of pol mu) - apoenzyme | | Descriptor: | DNA nucleotidylexotransferase,DNA-directed DNA/RNA polymerase mu,DNA nucleotidylexotransferase | | Authors: | Loc'h, J, Gerodimos, C.A, Rosario, S, Lieber, M.R, Delarue, M. | | Deposit date: | 2018-06-01 | | Release date: | 2019-06-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural evidence for an intransbase selection mechanism involving Loop1 in polymerase mu at an NHEJ double-strand break junction.
J.Biol.Chem., 294, 2019
|
|
6GO4
 
 | | TdT chimera (Loop1 of pol mu) - binary complex with ddCTP | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, DNA nucleotidylexotransferase,DNA-directed DNA/RNA polymerase mu,DNA nucleotidylexotransferase, MAGNESIUM ION, ... | | Authors: | Loc'h, J, Gerodimos, C.A, Rosario, S, Lieber, M.R, Delarue, M. | | Deposit date: | 2018-06-01 | | Release date: | 2019-06-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Structural evidence for an intransbase selection mechanism involving Loop1 in polymerase mu at an NHEJ double-strand break junction.
J.Biol.Chem., 294, 2019
|
|
6GO7
 
 | | TdT chimera (Loop1 of pol mu) - full DNA synapsis complex | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, DNA (5'-D(*AP*AP*AP*AP*AP*C)-3'), DNA (5'-D(*TP*TP*TP*TP*TP*GP*C)-3'), ... | | Authors: | Loc'h, J, Gerodimos, C.A, Rosario, S, Lieber, M.R, Delarue, M. | | Deposit date: | 2018-06-01 | | Release date: | 2019-06-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structural evidence for an intransbase selection mechanism involving Loop1 in polymerase mu at an NHEJ double-strand break junction.
J.Biol.Chem., 294, 2019
|
|
6GO5
 
 | | TdT chimera (Loop1 of pol mu) - Ternary complex with 1-nt gapped DNA substrate | | Descriptor: | 2'-deoxy-5'-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]methyl}phosphoryl]cytidine, DNA (5'-D(*AP*CP*AP*GP*CP*G)-3'), DNA (5'-D(*CP*GP*CP*TP*GP*GP*CP*AP*AP*AP*CP*A)-3'), ... | | Authors: | Loc'h, J, Gerodimos, C.A, Rosario, S, Lieber, M.R, Delarue, M. | | Deposit date: | 2018-06-01 | | Release date: | 2019-06-05 | | Last modified: | 2019-12-25 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural evidence for an intransbase selection mechanism involving Loop1 in polymerase mu at an NHEJ double-strand break junction.
J.Biol.Chem., 294, 2019
|
|
6GO6
 
 | | TdT chimera (Loop1 of pol mu) - ternary complex with downstream dsDNA | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, DNA (5'-D(*AP*AP*AP*AP*AP*C)-3'), DNA (5'-D(*TP*TP*TP*TP*TP*GP*GP*C)-3'), ... | | Authors: | Loc'h, J, Gerodimos, C.A, Rosario, S, Lieber, M.R, Delarue, M. | | Deposit date: | 2018-06-01 | | Release date: | 2019-06-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Structural evidence for an intransbase selection mechanism involving Loop1 in polymerase mu at an NHEJ double-strand break junction.
J.Biol.Chem., 294, 2019
|
|
