2WPL
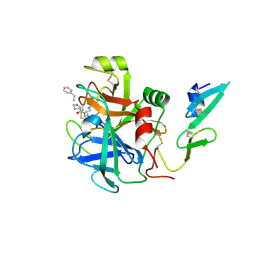 
 | | factor IXa superactive triple mutant, EDTA-soaked | | Descriptor: | COAGULATION FACTOR IXA HEAVY CHAIN, COAGULATION FACTOR IXA LIGHT CHAIN, D-PHE-PRO-ARG-CHLOROMETHYL KETONE | | Authors: | Zogg, T, Brandstetter, H. | | Deposit date: | 2009-08-06 | | Release date: | 2009-12-22 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Structural Basis of the Cofactor- and Substrate-Assisted Activation of Human Coagulation Factor Ixa
Structure, 17, 2009
|
|
4BTZ
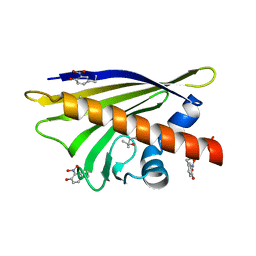 
 | |
6YSA
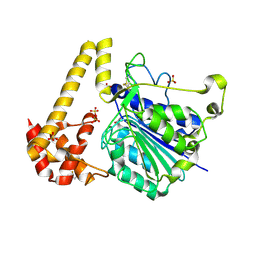 
 | | Crystal structure of Arabidopsis thaliana legumain isoform beta in zymogen state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CITRIC ACID, SULFATE ION, ... | | Authors: | Dall, E, Zauner, F.B, Brandstetter, H. | | Deposit date: | 2020-04-21 | | Release date: | 2020-07-29 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Structural and functional studies ofArabidopsis thalianalegumain beta reveal isoform specific mechanisms of activation and substrate recognition.
J.Biol.Chem., 295, 2020
|
|
4FGU
 
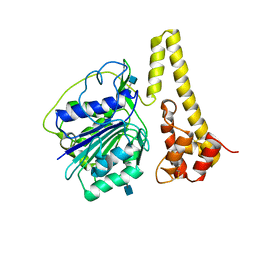 | | Crystal structure of prolegumain | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Legumain | | Authors: | Dall, E, Brandstetter, H. | | Deposit date: | 2012-06-05 | | Release date: | 2013-07-03 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3.9 Å) | | Cite: | Mechanistic and structural studies on legumain explain its zymogenicity, distinct activation pathways, and regulation.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4N6L
 
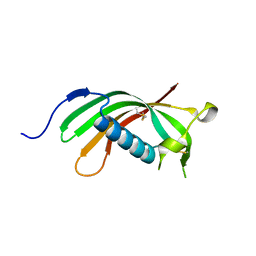 | | Crystal structure of human cystatin E/M | | Descriptor: | Cystatin-M | | Authors: | Dall, E, Brandstetter, H. | | Deposit date: | 2013-10-14 | | Release date: | 2015-02-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.952 Å) | | Cite: | Structure and mechanism of an aspartimide-dependent Peptide ligase in human legumain.
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
4N1J
 
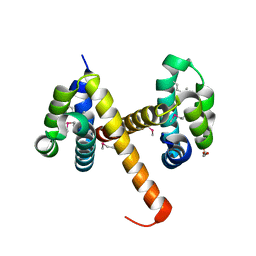 | | Crystal structures of NLRP14 pyrin domain reveal a conformational switch mechanism, regulating its molecular interactions | | Descriptor: | GLYCEROL, NACHT, LRR and PYD domains-containing protein 14 | | Authors: | Eibl, C, Hessenberger, M, Wenger, J, Brandstetter, H. | | Deposit date: | 2013-10-04 | | Release date: | 2014-07-16 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structures of the NLRP14 pyrin domain reveal a conformational switch mechanism regulating its molecular interactions.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
5NIJ
 
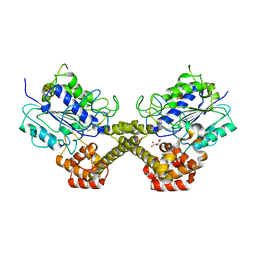 | |
5O7E
 
 | |
5NZC
 
 | |
5NZB
 
 | |
5OBT
 
 | |
5LUA
 
 | | Crystal structure of human legumain (AEP) in complex with compound 11b | | Descriptor: | 2,4-di(morpholin-4-yl)aniline, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Dall, E, Ye, K, Brandstetter, H. | | Deposit date: | 2016-09-08 | | Release date: | 2017-03-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Inhibition of delta-secretase improves cognitive functions in mouse models of Alzheimer's disease.
Nat Commun, 8, 2017
|
|
5LUB
 
 | | Crystal structure of human legumain (AEP) in complex with compound 11 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 7-(morpholin-4-yl)-2,1,3-benzoxadiazol-4-amine, ... | | Authors: | Dall, E, Ye, K, Brandstetter, H. | | Deposit date: | 2016-09-08 | | Release date: | 2017-03-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Inhibition of delta-secretase improves cognitive functions in mouse models of Alzheimer's disease.
Nat Commun, 8, 2017
|
|
5LU8
 
 | | CRYSTAL STRUCTURE OF YVAD-CMK BOUND HUMAN LEGUMAIN (AEP) IN COMPLEX WITH COMPOUND 11B | | Descriptor: | 2,4-di(morpholin-4-yl)aniline, 2-acetamido-2-deoxy-beta-D-glucopyranose, AC-TYR-VAL-ALA-ASP-CHLOROMETHYLKETONE, ... | | Authors: | Dall, E, Ye, K, Brandstetter, H. | | Deposit date: | 2016-09-08 | | Release date: | 2017-03-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Inhibition of delta-secretase improves cognitive functions in mouse models of Alzheimer's disease.
Nat Commun, 8, 2017
|
|
5LU9
 
 | | Crystal structure of YVAD-cmk bound human legumain (AEP) in complex with compound 11 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 7-(morpholin-4-yl)-2,1,3-benzoxadiazol-4-amine, AC-TYR-VAL-ALA-ASP-CHLOROMETHYLKETONE, ... | | Authors: | Dall, E, Ye, K, Brandstetter, H. | | Deposit date: | 2016-09-08 | | Release date: | 2017-03-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Inhibition of delta-secretase improves cognitive functions in mouse models of Alzheimer's disease.
Nat Commun, 8, 2017
|
|
2Y72
 
 | |
5JBB
 
 | |
5JB8
 
 | |
5JBC
 
 | |
5JBA
 
 | |
7O1U
 
 | |
7O20
 
 | |
7O22
 
 | |
7O1W
 
 | |
7O1Y
 
 | |
