7UYI
 | |
5KOU
 
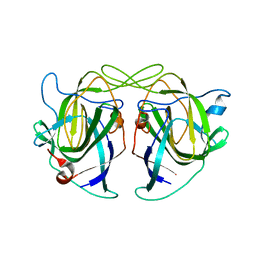 | |
5KOV
 
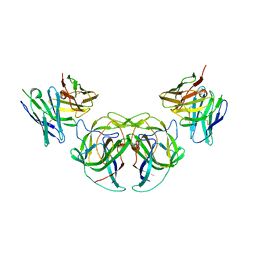 | |
8CT6
 
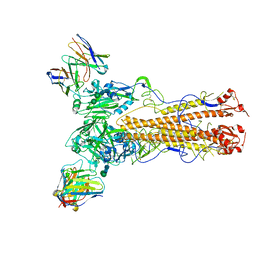 | |
6UVO
 
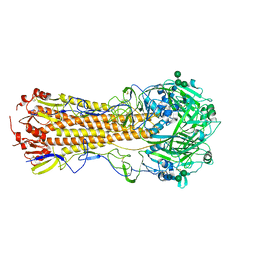
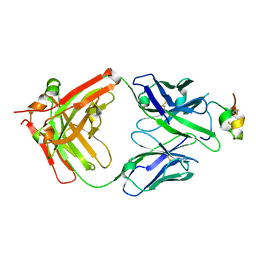 | | Structure of antibody 3G12 bound to the central conserved domain of RSV G | | Descriptor: | 3G12 Fab Heavy chain, 3G12 Fab light chain, Major surface glycoprotein G | | Authors: | Fedechkin, S.O, George, N.L, Nunez Castrejon, A.M, Dillen, J, Kauvar, L.M, DuBois, R.M. | | Deposit date: | 2019-11-03 | | Release date: | 2019-12-18 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Conformational Flexibility in Respiratory Syncytial Virus G Neutralizing Epitopes.
J.Virol., 94, 2020
|
|
7RK1
 
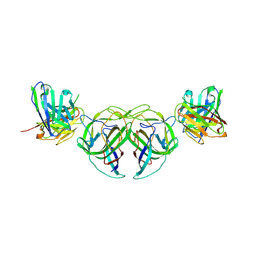 | |
7RK2
 
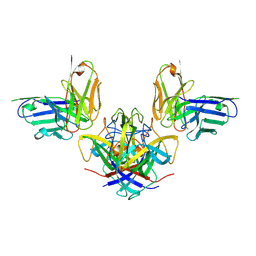 | | Crystal structure of the human astrovirus serotype 8 capsid spike in complex with scFv 2D9, an astrovirus-neutralizing antibody, at 2.65-A resolution | | Descriptor: | Capsid protein VP25, scFv 2D9 | | Authors: | Meyer, L, Cuellar, C, DuBois, R.M. | | Deposit date: | 2021-07-21 | | Release date: | 2021-10-13 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structures of Two Human Astrovirus Capsid/Neutralizing Antibody Complexes Reveal Distinct Epitopes and Inhibition of Virus Attachment to Cells.
J.Virol., 96, 2022
|
|
5EWO
 
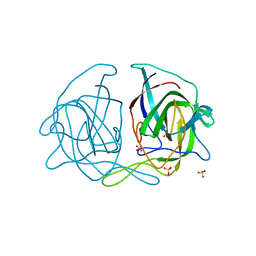 | | Crystal structure of the human astrovirus 1 capsid protein spike domain at 0.95-A resolution | | Descriptor: | SULFATE ION, Structural protein | | Authors: | Bogdanoff, W, York, R.L, Yousefi, P.A, Haile, S, Tripathi, S, DuBois, R.M. | | Deposit date: | 2015-11-20 | | Release date: | 2015-12-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (0.95 Å) | | Cite: | Structural, Mechanistic, and Antigenic Characterization of the Human Astrovirus Capsid.
J.Virol., 90, 2015
|
|
5EWN
 
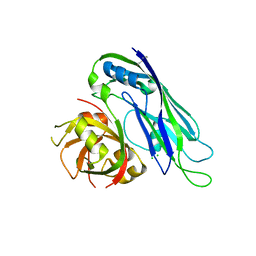 | | Crystal structure of the human astrovirus 1 capsid protein core domain at 2.6 A resolution | | Descriptor: | CHLORIDE ION, Structural protein | | Authors: | York, R.L, Yousefi, P.A, Bogdanoff, W, Haile, S, Tripathi, S, DuBois, R.M. | | Deposit date: | 2015-11-20 | | Release date: | 2015-12-23 | | Last modified: | 2019-12-11 | | Method: | X-RAY DIFFRACTION (2.602 Å) | | Cite: | Structural, Mechanistic, and Antigenic Characterization of the Human Astrovirus Capsid.
J.Virol., 90, 2015
|
|
8SJ9
 
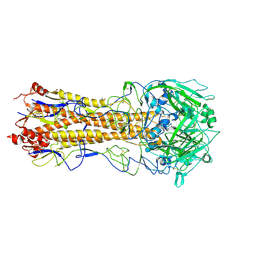 | | Crystal structure of the H1 hemagglutinin COBRA X6 | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Dzimianski, J.V, DuBois, R.M. | | Deposit date: | 2023-04-17 | | Release date: | 2024-05-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Structural basis for the broad antigenicity of the computationally optimized influenza hemagglutinin X6.
Structure, 32, 2024
|
|
8TN8
 
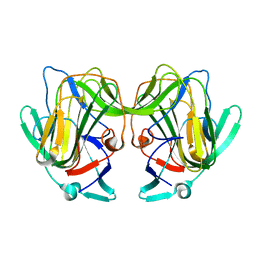 | |
8TUI
 
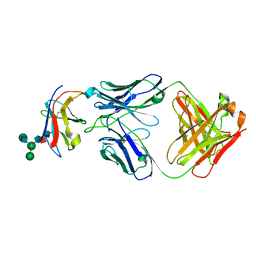 | | Crystal structure of Fab-Lirilumab bound to KIR2DL3 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)-[alpha-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Killer cell immunoglobulin-like receptor 2DL3, Lirilumab Fab Heavy Chain, ... | | Authors: | Lorig-Roach, N, DuBois, R.M. | | Deposit date: | 2023-08-16 | | Release date: | 2023-12-06 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural basis for the activity and specificity of the immune checkpoint inhibitor lirilumab.
Sci Rep, 14, 2024
|
|
5W1N
 
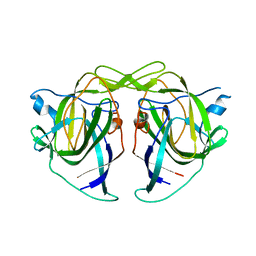 | |
5WN9
 
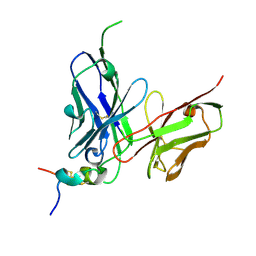 | | Structure of antibody 2D10 bound to the central conserved region of RSV G | | Descriptor: | Major surface glycoprotein G, scFv 2D10 | | Authors: | Fedechkin, S.O, George, N.L, Wolff, J.T, Kauvar, L.M, DuBois, R.M. | | Deposit date: | 2017-07-31 | | Release date: | 2018-03-14 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.551 Å) | | Cite: | Structures of respiratory syncytial virus G antigen bound to broadly neutralizing antibodies.
Sci Immunol, 3, 2018
|
|
5WNA
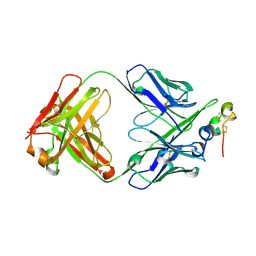 
 | | Structure of antibody 3D3 bound to the central conserved region of RSV G | | Descriptor: | Major surface glycoprotein G, mAb 3D3 Fab heavy chain, mAb 3D3 Fab light chain | | Authors: | Fedechkin, S.O, George, N.L, Wolff, J.T, Kauvar, L.M, DuBois, R.M. | | Deposit date: | 2017-07-31 | | Release date: | 2018-03-14 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structures of respiratory syncytial virus G antigen bound to broadly neutralizing antibodies.
Sci Immunol, 3, 2018
|
|
5WNB
 
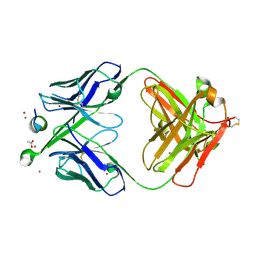 | | Structure of antibody 3D3 bound to the linear epitope of RSV G | | Descriptor: | ACETIC ACID, Major surface glycoprotein G, ZINC ION, ... | | Authors: | Fedechkin, S.O, George, N.L, Wolff, J.T, Kauvar, L.M, DuBois, R.M. | | Deposit date: | 2017-07-31 | | Release date: | 2018-03-14 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structures of respiratory syncytial virus G antigen bound to broadly neutralizing antibodies.
Sci Immunol, 3, 2018
|
|
2XQY
 
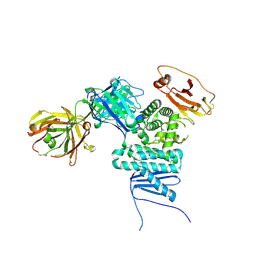 | | CRYSTAL STRUCTURE OF PSEUDORABIES CORE FRAGMENT OF GLYCOPROTEIN H IN COMPLEX WITH FAB D6.3 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, A13-D6.3 MONOCLONAL ANTIBODY, ... | | Authors: | Backovic, M, Dubois, R, Cockburn, J, Sharff, A, Vaney, M, Granzow, H, Klupp, B, Bricogne, G, Mettenleiter, T, Rey, F. | | Deposit date: | 2010-09-08 | | Release date: | 2010-12-22 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure of a Core Fragment of Glycoprotein H from Pseudorabies Virus in Complex with Antibody.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
7UZT
 
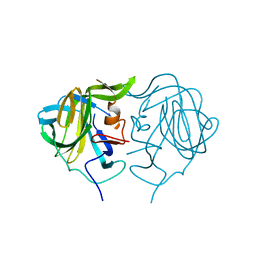 | |
8GHK
 
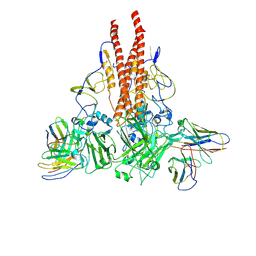 | |
8DTW
 
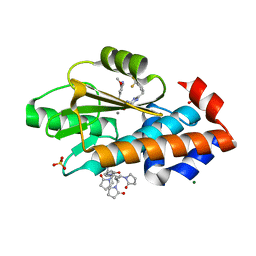 | | The crystal structure of I38T mutant PA endonuclease (2009/H1N1/CALIFORNIA) in complex with compound SJ001023036 | | Descriptor: | 5-hydroxy-N-[2-(2-methoxypyridin-4-yl)ethyl]-6-oxo-2-{[2-(trifluoromethyl)phenyl]methyl}-1,6-dihydropyrimidine-4-carboxamide, Hexa Vinylpyrrolidone K15, MAGNESIUM ION, ... | | Authors: | Cuypers, M.G, Slavish, J.P, Rankovic, Z, White, S.W. | | Deposit date: | 2022-07-26 | | Release date: | 2022-09-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Chemical scaffold recycling: Structure-guided conversion of an HIV integrase inhibitor into a potent influenza virus RNA-dependent RNA polymerase inhibitor designed to minimize resistance potential.
Eur.J.Med.Chem., 247, 2023
|
|
8E4S
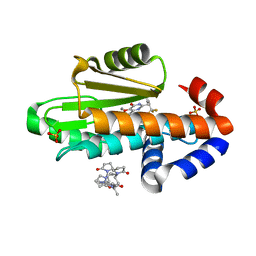 
 | | The crystal structure of I38T mutant PA endonuclease (2009/H1N1/CALIFORNIA) in complex with compound SJ001023034 | | Descriptor: | 5-hydroxy-6-oxo-N-[2-(pyridin-4-yl)ethyl]-2-{[2-(trifluoromethyl)phenyl]methyl}-3,6-dihydropyrimidine-4-carboxamide, Hexa Vinylpyrrolidone K15, MANGANESE (II) ION, ... | | Authors: | Cuypers, M.G, Slavish, J.P, Rankovic, Z, White, S.W. | | Deposit date: | 2022-08-18 | | Release date: | 2022-09-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Chemical scaffold recycling: Structure-guided conversion of an HIV integrase inhibitor into a potent influenza virus RNA-dependent RNA polymerase inhibitor designed to minimize resistance potential.
Eur.J.Med.Chem., 247, 2023
|
|
7RKP
 
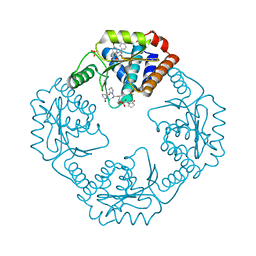 | | The crystal structure of I38T mutant PA endonuclease (2009/H1N1/CALIFORNIA) in complex with cyclic compound SJ001034733 | | Descriptor: | (5R)-5-hydroxy-16,21-dioxa-3,8,28-triazatetracyclo[20.3.1.1~2,6~.1~11,15~]octacosa-1(26),2,6(28),11(27),12,14,22,24-octaene-4,7-dione, Hexa Vinylpyrrolidone K15, MANGANESE (II) ION, ... | | Authors: | Cuypers, M.G, Slavish, J.P, Rankovic, Z, White, S.W. | | Deposit date: | 2021-07-22 | | Release date: | 2022-07-27 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | Chemical scaffold recycling: Structure-guided conversion of an HIV integrase inhibitor into a potent influenza virus RNA-dependent RNA polymerase inhibitor designed to minimize resistance potential.
Eur.J.Med.Chem., 247, 2023
|
|
8F38
 
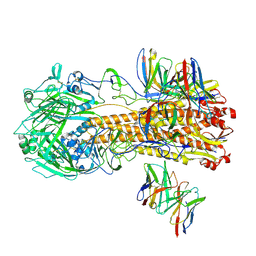 | |
8V7O
 
 | |
7UMR
 
 | | The crystal structure of wild type PA endonuclease (2009/H1N1/CALIFORNIA) in complex with compound SJ001034732-1 (trans-form) | | Descriptor: | (1P,18Z)-5-hydroxy-16,21-dioxa-3,8,28-triazatetracyclo[20.3.1.1~2,6~.1~11,15~]octacosa-1(26),2(28),5,11(27),12,14,18,22,24-nonaene-4,7-dione, Hexa Vinylpyrrolidone K15, MANGANESE (II) ION, ... | | Authors: | Cuypers, M.G, Slavish, J.P, Rankovic, Z, White, S.W. | | Deposit date: | 2022-04-07 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Chemical scaffold recycling: Structure-guided conversion of an HIV integrase inhibitor into a potent influenza virus RNA-dependent RNA polymerase inhibitor designed to minimize resistance potential.
Eur.J.Med.Chem., 247, 2023
|
|
