1AKD
 
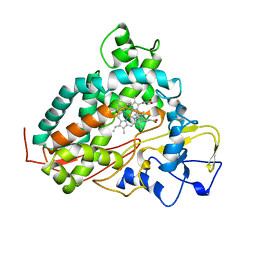 | | CYTOCHROME P450CAM FROM PSEUDOMONAS PUTIDA, COMPLEXED WITH 1S-CAMPHOR | | Descriptor: | CAMPHOR, CYTOCHROME P450CAM, POTASSIUM ION, ... | | Authors: | Schlichting, I, Jung, C, Schulze, H. | | Deposit date: | 1997-05-16 | | Release date: | 1997-11-19 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of cytochrome P-450cam complexed with the (1S)-camphor enantiomer.
FEBS Lett., 415, 1997
|
|
1XX4
 
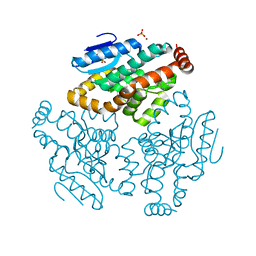 | | Crystal Structure of Rat Mitochondrial 3,2-Enoyl-CoA | | Descriptor: | 3,2-trans-enoyl-CoA isomerase, mitochondrial, BENZAMIDINE, ... | | Authors: | Hubbard, P.A, Yu, W, Schulz, H, Kim, J.-J. | | Deposit date: | 2004-11-03 | | Release date: | 2004-11-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Domain swapping in the low-similarity isomerase/hydratase superfamily: the crystal structure of rat mitochondrial Delta3, Delta2-enoyl-CoA isomerase.
Protein Sci., 14, 2005
|
|
1PS9
 
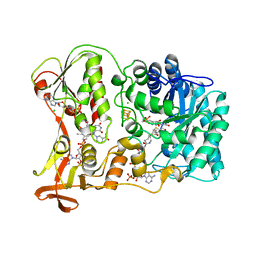 | | The Crystal Structure and Reaction Mechanism of E. coli 2,4-Dienoyl CoA Reductase | | Descriptor: | 2,4-dienoyl-CoA reductase, 5-MERCAPTOETHANOL-2-DECENOYL-COENZYME A, CHLORIDE ION, ... | | Authors: | Hubbard, P.A, Liang, X, Schulz, H, Kim, J.J. | | Deposit date: | 2003-06-20 | | Release date: | 2003-09-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The crystal structure and reaction mechanism of Escherichia coli 2,4-dienoyl-CoA reductase
J.Biol.Chem., 278, 2003
|
|
3DWB
 
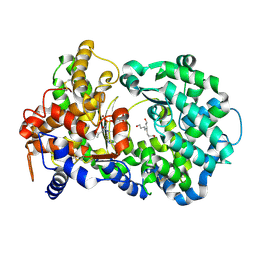 | | structure of human ECE-1 complexed with phosphoramidon | | Descriptor: | 5-(2-hydroxyethyl)nonane-1,9-diol, Endothelin-converting enzyme 1, N-ALPHA-L-RHAMNOPYRANOSYLOXY(HYDROXYPHOSPHINYL)-L-LEUCYL-L-TRYPTOPHAN, ... | | Authors: | Oefner, C. | | Deposit date: | 2008-07-22 | | Release date: | 2008-11-25 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Structure of human endothelin-converting enzyme I complexed with phosphoramidon
J.Mol.Biol., 385, 2009
|
|
6R0X
 
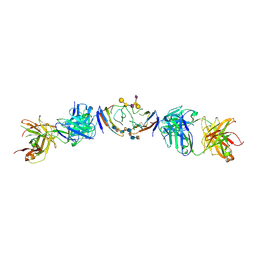 | | The extracellular domain of G6b-B in complex with Fab fragment and DP12 heparin oligosaccharide. | | Descriptor: | 2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose, Megakaryocyte and platelet inhibitory receptor G6b, antibody fab fragment heavy chain, ... | | Authors: | Ogg, D.J, McMiken, H.J, Howard, T.D. | | Deposit date: | 2019-03-13 | | Release date: | 2019-09-04 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (3.13 Å) | | Cite: | Heparan sulfates are critical regulators of the inhibitory megakaryocyte-platelet receptor G6b-B.
Elife, 8, 2019
|
|
1QXZ
 
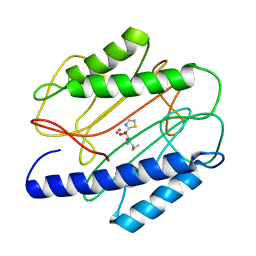 | | Crystal structure of S. aureus methionine aminopeptidase in complex with a ketoheterocycle inhibitor 119 | | Descriptor: | (2S)-2-AMINO-4-(METHYLSULFANYL)-1-(1,3-THIAZOL-2-YL)BUTANE-1,1-DIOL, COBALT (II) ION, methionyl aminopeptidase | | Authors: | Douangamath, A, Dale, G.E, D'Arcy, A, Oefner, C. | | Deposit date: | 2003-09-09 | | Release date: | 2004-03-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Crystal structures of staphylococcusaureus methionine aminopeptidase complexed with keto heterocycle and aminoketone inhibitors reveal the formation of a tetrahedral intermediate.
J.Med.Chem., 47, 2004
|
|
1QXW
 
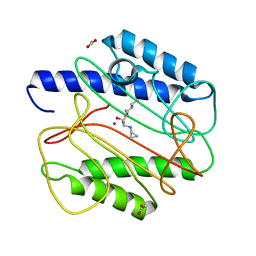 | | Crystal structure of Staphyloccocus aureus in complex with an aminoketone inhibitor 54135. | | Descriptor: | (3S)-3-AMINO-1-(CYCLOPROPYLAMINO)HEPTANE-2,2-DIOL, ACETATE ION, COBALT (II) ION, ... | | Authors: | Douangamath, A, Dale, G.E, D'Arcy, A, Oefner, C. | | Deposit date: | 2003-09-09 | | Release date: | 2004-03-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Crystal structures of staphylococcusaureus methionine aminopeptidase complexed with keto heterocycle and aminoketone inhibitors reveal the formation of a tetrahedral intermediate.
J.Med.Chem., 47, 2004
|
|
1QXY
 
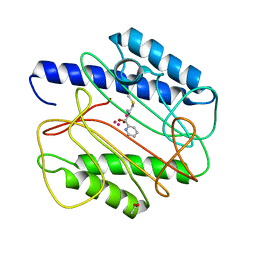 | | Crystal structure of S. aureus methionine aminopeptidase in complex with a ketoheterocycle 618 | | Descriptor: | (2S)-2-AMINO-4-(METHYLSULFANYL)-1-PYRIDIN-2-YLBUTANE-1,1-DIOL, ACETATE ION, COBALT (II) ION, ... | | Authors: | Douangamath, A, Dale, G.E, D'Arcy, A, Oefner, C. | | Deposit date: | 2003-09-09 | | Release date: | 2004-03-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | Crystal structures of staphylococcusaureus methionine aminopeptidase complexed with keto heterocycle and aminoketone inhibitors reveal the formation of a tetrahedral intermediate.
J.Med.Chem., 47, 2004
|
|
3FRB
 
 | | S. aureus F98Y DHFR complexed with TMP | | Descriptor: | Dihydrofolate reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, TRIMETHOPRIM | | Authors: | Oefner, C, Dale-Glenn, E. | | Deposit date: | 2009-01-08 | | Release date: | 2010-01-12 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Increased hydrophobic interactions of iclaprim with Staphylococcus aureus dihydrofolate reductase are responsible for the increase in affinity and antibacterial activity
J.Antimicrob.Chemother., 63, 2009
|
|
3FRF
 
 | | S. aureus DHFR complexed with NADPH and iclaprim | | Descriptor: | 5-[[(2R)-2-cyclopropyl-7,8-dimethoxy-2H-chromen-5-yl]methyl]pyrimidine-2,4-diamine, Dihydrofolate reductase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Oefner, C, Dale-Glenn, E. | | Deposit date: | 2009-01-08 | | Release date: | 2010-01-12 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Increased hydrophobic interactions of iclaprim with Staphylococcus aureus dihydrofolate reductase are responsible for the increase in affinity and antibacterial activity
J.Antimicrob.Chemother., 63, 2009
|
|
3FRE
 
 | | S. aureus DHFR complexed with NADPH and TMP | | Descriptor: | Dihydrofolate reductase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, TRIMETHOPRIM | | Authors: | Oefner, C, Dale-Glenn, E. | | Deposit date: | 2009-01-08 | | Release date: | 2010-01-12 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Increased hydrophobic interactions of iclaprim with Staphylococcus aureus dihydrofolate reductase are responsible for the increase in affinity and antibacterial activity
J.Antimicrob.Chemother., 63, 2009
|
|
3FRD
 
 | | S. aureus DHFR complexed with NADPH and folate | | Descriptor: | DIHYDROFOLIC ACID, Dihydrofolate reductase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Oefner, C, Dale-Glenn, E. | | Deposit date: | 2009-01-08 | | Release date: | 2010-01-12 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Increased hydrophobic interactions of iclaprim with Staphylococcus aureus dihydrofolate reductase are responsible for the increase in affinity and antibacterial activity
J.Antimicrob.Chemother., 63, 2009
|
|
3FRA
 
 | | Staphylococcus aureus F98Y DHFR complexed with iclaprim | | Descriptor: | 5-{[(2S)-2-cyclopropyl-7,8-dimethoxy-2H-chromen-5-yl]methyl}pyrimidine-2,4-diamine, Dihydrofolate reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Oefner, C, Dale-Glenn, E. | | Deposit date: | 2009-01-08 | | Release date: | 2010-01-12 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Increased hydrophobic interactions of iclaprim with Staphylococcus aureus dihydrofolate reductase are responsible for the increase in affinity and antibacterial activity
J.Antimicrob.Chemother., 63, 2009
|
|
5AA6
 | | Homohexameric Structure of the second Vanadate-Dependent Bromoperoxidase (AnII) from Ascophyllum nodosum | | Descriptor: | VANADATE ION, VANADIUM-DEPENDENT BROMOPEROXIDASE 2 | | Authors: | Radlow, M, Jeudy, A, Dabin, J, Delage, L, Leblanc, C, Hartung, J, Czjzek, M. | | Deposit date: | 2015-07-23 | | Release date: | 2016-08-03 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Homohexameric Structure of the Second Vanadate Dependant Bromoperoxidase from Ascophyllum Nodosum
To be Published
|
|
3FYV
 
 | | Staph. aureus DHFR complexed with NADPH and AR-102 | | Descriptor: | 5-[[(2R)-2-cyclopropyl-7,8-dimethoxy-2H-chromen-5-yl]methyl]pyrimidine-2,4-diamine, Dihydrofolate reductase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Oefner, C. | | Deposit date: | 2009-01-23 | | Release date: | 2009-08-04 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Inhibitory properties and X-ray crystallographic study of the binding of AR-101, AR-102 and iclaprim in ternary complexes with NADPH and dihydrofolate reductase from Staphylococcus aureus
Acta Crystallogr.,Sect.D, 65, 2009
|
|
3FY8
 
 | | Crystal Structure of Staph. aureus DHFR complexed with NADPH and AR-101 | | Descriptor: | 5-[[(2R)-2-cyclopropyl-7,8-dimethoxy-2H-chromen-5-yl]methyl]pyrimidine-2,4-diamine, Dihydrofolate reductase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Oefner, C, Dale, G.E. | | Deposit date: | 2009-01-22 | | Release date: | 2009-08-04 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Inhibitory properties and X-ray crystallographic study of the binding of AR-101, AR-102 and iclaprim in ternary complexes with NADPH and dihydrofolate reductase from Staphylococcus aureus
Acta Crystallogr.,Sect.D, 65, 2009
|
|
3FY9
 
 | | Staph. aureus DHFR F98Y complexed with AR-102 | | Descriptor: | 5-[[(2R)-2-cyclopropyl-7,8-dimethoxy-2H-chromen-5-yl]methyl]pyrimidine-2,4-diamine, Dihydrofolate reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Oefner, C. | | Deposit date: | 2009-01-22 | | Release date: | 2009-08-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Inhibitory properties and X-ray crystallographic study of the binding of AR-101, AR-102 and iclaprim in ternary complexes with NADPH and dihydrofolate reductase from Staphylococcus aureus
Acta Crystallogr.,Sect.D, 65, 2009
|
|
3FYW
 
 | | Staph. aureus DHFR complexed with NADPH and AR-101 | | Descriptor: | 5-[[(2R)-2-cyclopropyl-7,8-dimethoxy-2H-chromen-5-yl]methyl]pyrimidine-2,4-diamine, Dihydrofolate reductase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Oefner, C. | | Deposit date: | 2009-01-23 | | Release date: | 2009-08-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Inhibitory properties and X-ray crystallographic study of the binding of AR-101, AR-102 and iclaprim in ternary complexes with NADPH and dihydrofolate reductase from Staphylococcus aureus
Acta Crystallogr.,Sect.D, 65, 2009
|
|
2G2Z
 
 | | Structure of E.coli FabD complexed with malonyl-CoA | | Descriptor: | COENZYME A, MALONIC ACID, Malonyl CoA-acyl carrier protein transacylase | | Authors: | Oefner, Christian | | Deposit date: | 2006-02-17 | | Release date: | 2006-06-27 | | Last modified: | 2015-04-29 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Mapping the active site of Escherichia coli malonyl-CoA-acyl carrier protein transacylase (FabD) by protein crystallography.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2G2O
 
 | | Structure of E.coli FabD complexed with sulfate | | Descriptor: | Malonyl CoA-acyl carrier protein transacylase, SULFATE ION | | Authors: | Oefner, C. | | Deposit date: | 2006-02-16 | | Release date: | 2006-05-30 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Mapping the active site of Escherichia coli malonyl-CoA-acyl carrier protein transacylase (FabD) by protein crystallography.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2G2Y
 
 | | Structure of E.coli FabD complexed with malonate | | Descriptor: | MALONATE ION, Malonyl CoA-acyl carrier protein transacylase | | Authors: | Oefner, C. | | Deposit date: | 2006-02-17 | | Release date: | 2006-05-30 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Mapping the active site of Escherichia coli malonyl-CoA-acyl carrier protein transacylase (FabD) by protein crystallography.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2G1H
 
 | | Structure of E.coli FabD complexed with glycerol | | Descriptor: | GLYCEROL, Malonyl CoA-acyl carrier protein transacylase | | Authors: | Oefner, C. | | Deposit date: | 2006-02-14 | | Release date: | 2006-05-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Mapping the active site of Escherichia coli malonyl-CoA-acyl carrier protein transacylase (FabD) by protein crystallography.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2QPJ
 
 | | Human NEP complexed with a bifunctional NEP/DPP IV inhibitor | | Descriptor: | (2S)-2-({(2S)-3-[(R)-[(1R)-1-({(4S)-4-amino-5-[(2S)-2-cyanopyrrolidin-1-yl]-5-oxopentanoyl}amino)ethyl](hydroxy)phosphoryl]-2-benzylpropanoyl}amino)propanoic acid, 2-acetamido-2-deoxy-beta-D-glucopyranose, Neprilysin, ... | | Authors: | Oefner, C, Dale, G.E. | | Deposit date: | 2007-07-24 | | Release date: | 2007-11-27 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural studies of a bifunctional inhibitor of neprilysin and DPP-IV.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
1Q0Q
 
 | | Crystal structure of DXR in complex with the substrate 1-deoxy-D-xylulose-5-phosphate | | Descriptor: | 1-DEOXY-D-XYLULOSE-5-PHOSPHATE, 1-deoxy-D-xylulose 5-phosphate reductoisomerase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Mac Sweeney, A, Lange, R, D'Arcy, A, Douangamath, A, Surivet, J.-P, Oefner, C. | | Deposit date: | 2003-07-17 | | Release date: | 2004-07-20 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The crystal structure of E.coli 1-deoxy-D-xylulose-5-phosphate reductoisomerase in a ternary complex with the antimalarial compound fosmidomycin and NADPH reveals a tight-binding closed enzyme conformation.
J.Mol.Biol., 345, 2005
|
|
1Q0H
 
 | | Crystal structure of selenomethionine-labelled DXR in complex with fosmidomycin | | Descriptor: | 1-deoxy-D-xylulose 5-phosphate reductoisomerase, 3-[FORMYL(HYDROXY)AMINO]PROPYLPHOSPHONIC ACID, CITRIC ACID, ... | | Authors: | Mac Sweeney, A, Lange, R, D'Arcy, A, Douangamath, A, Surivet, J.-P, Oefner, C. | | Deposit date: | 2003-07-16 | | Release date: | 2004-07-20 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The crystal structure of E.coli 1-deoxy-D-xylulose-5-phosphate reductoisomerase in a ternary complex with the antimalarial compound fosmidomycin and NADPH reveals a tight-binding closed enzyme conformation.
J.Mol.Biol., 345, 2005
|
|
