4NIK
 
 | |
3DR1
 
 | | Side-chain fluorine atoms of non-steroidal vitamin D3 analogs stabilize helix 12 of vitamin D receptor | | Descriptor: | (1R,3R)-5-[(2E)-3-{(1S,3R)-2,2,3-trimethyl-3-[6,6,6-trifluoro-5-hydroxy-5-(trifluoromethyl)hex-3-yn-1-yl]cyclopentyl}prop-2-en-1-ylidene]cyclohexane-1,3-diol, MAGNESIUM ION, SRC-1 (LXXLL motif) from Nuclear receptor coactivator 1, ... | | Authors: | Sato, Y, Rochel, N, Moras, D. | | Deposit date: | 2008-07-10 | | Release date: | 2009-06-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Superagonistic fluorinated vitamin D3 analogs stabilize helix 12 of the vitamin D receptor.
Chem.Biol., 15, 2008
|
|
7E62
 
 | | Mouse TAB2 NZF in complex with Lys6-linked diubiquitin | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, TGF-beta-activated kinase 1 and MAP3K7-binding protein 2, Ubiquitin, ... | | Authors: | Sato, Y, Li, Y, Okatsu, K, Fukai, S. | | Deposit date: | 2021-02-21 | | Release date: | 2021-08-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Structural basis for specific recognition of K6-linked polyubiquitin chains by the TAB2 NZF domain.
Biophys.J., 120, 2021
|
|
6KC6
 
 | | HOIP-HOIPIN8 complex | | Descriptor: | 2-[3-[2,6-bis(fluoranyl)-4-(1~{H}-pyrazol-4-yl)phenyl]-3-oxidanylidene-propyl]-4-(1-methylpyrazol-4-yl)benzoic acid, CHLORIDE ION, E3 ubiquitin-protein ligase RNF31, ... | | Authors: | Sato, Y, Fukai, S. | | Deposit date: | 2019-06-27 | | Release date: | 2020-04-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.123 Å) | | Cite: | Molecular bases for HOIPINs-mediated inhibition of LUBAC and innate immune responses.
Commun Biol, 3, 2020
|
|
6KC5
 
 | | HOIP-HOIPIN1 complex | | Descriptor: | 2-[3-(2-methoxyphenyl)-3-oxidanylidene-propyl]benzoic acid, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, E3 ubiquitin-protein ligase RNF31, ... | | Authors: | Sato, Y, Fukai, S. | | Deposit date: | 2019-06-27 | | Release date: | 2020-04-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.543 Å) | | Cite: | Molecular bases for HOIPINs-mediated inhibition of LUBAC and innate immune responses.
Commun Biol, 3, 2020
|
|
6IES
 
 | | Onion lachrymatory factor synthase (LFS) containing (E)-2-propen 1-ol (crotyl alcohol) | | Descriptor: | (2E)-but-2-en-1-ol, Lachrymatory-factor synthase | | Authors: | Sato, Y, Arakawa, T, Takabe, J, Masamura, N, Tsuge, N, Imai, S, Fushinobu, S. | | Deposit date: | 2018-09-17 | | Release date: | 2019-09-18 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Dissecting the Stereocontrolled Conversion of Short-Lived Sulfenic Acid by Lachrymatory Factor Synthase.
Acs Catalysis, 10, 2020
|
|
3WWQ
 
 | |
5YJE
 
 | | Crystal structure of HIRA(644-1017) | | Descriptor: | Protein HIRA, SULFATE ION | | Authors: | Sato, Y, Senda, M, Senda, T. | | Deposit date: | 2017-10-10 | | Release date: | 2018-06-20 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Functional activity of the H3.3 histone chaperone complex HIRA requires trimerization of the HIRA subunit
Nat Commun, 9, 2018
|
|
3A2M
 
 | | CRYSTAL STRUCTURE OF DBJA (WILD TYPE Type I) | | Descriptor: | CHLORIDE ION, Haloalkane dehalogenase, beta-D-fructofuranose-(2-1)-alpha-D-glucopyranose | | Authors: | Sato, Y, Senda, T. | | Deposit date: | 2009-05-23 | | Release date: | 2010-05-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Enantioselectivity of haloalkane dehalogenases and its modulation by surface loop engineering
Angew.Chem.Int.Ed.Engl., 49, 2010
|
|
3A2L
 
 | | Crystal structure of DBJA (mutant dbja delta) | | Descriptor: | CHLORIDE ION, GLYCEROL, Haloalkane dehalogenase, ... | | Authors: | Sato, Y, Senda, T. | | Deposit date: | 2009-05-23 | | Release date: | 2010-05-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Enantioselectivity of haloalkane dehalogenases and its modulation by surface loop engineering
Angew.Chem.Int.Ed.Engl., 49, 2010
|
|
3AFI
 
 | | Crystal structure of DBJA (HIS-DBJA) | | Descriptor: | CHLORIDE ION, Haloalkane dehalogenase | | Authors: | Sato, Y, Senda, T. | | Deposit date: | 2010-03-02 | | Release date: | 2011-01-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Enantioselectivity of haloalkane dehalogenases and its modulation by surface loop engineering
Angew.Chem.Int.Ed.Engl., 49, 2010
|
|
3A2N
 
 | | Crystal structure of DBJA (Wild Type Type II P21) | | Descriptor: | CHLORIDE ION, Haloalkane dehalogenase | | Authors: | Sato, Y, Senda, T. | | Deposit date: | 2009-05-23 | | Release date: | 2010-05-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Enantioselectivity of haloalkane dehalogenases and its modulation by surface loop engineering
Angew.Chem.Int.Ed.Engl., 49, 2010
|
|
2E7S
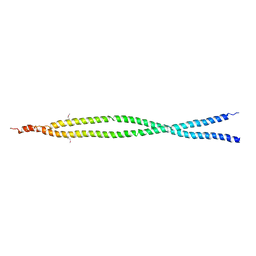 
 | |
2ZXZ
 
 | | Crystal structure of the human RXR alpha ligand binding domain bound to a synthetic agonist compound and a coactivator peptide | | Descriptor: | 4-[2-(1,1,3,3-tetramethyl-2,3-dihydro-1H-inden-5-yl)-1,3-dioxolan-2-yl]benzoic acid, GRIP1 from Nuclear receptor coactivator 2, Retinoic acid receptor RXR-alpha | | Authors: | Sato, Y, Antony, P, Rochel, N, Moras, D, Structural Genomics Consortium for Research on Gene Expression (SGCGES) | | Deposit date: | 2009-01-09 | | Release date: | 2009-08-11 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Silicon analogues of the RXR-selective retinoid agonist SR11237 (BMS649): chemistry and biology
Chemmedchem, 4, 2009
|
|
2ZY0
 
 | | Crystal structure of the human RXR alpha ligand binding domain bound to a synthetic agonist compound and a coactivator peptide | | Descriptor: | 4-[2-(1,1,3,3-tetramethyl-2,3-dihydro-1H-1,3-benzodisilol-5-yl)-1,3-dioxolan-2-yl]benzoic acid, GRIP1 from Nuclear receptor coactivator 2, Retinoic acid receptor RXR-alpha | | Authors: | Sato, Y, Antony, P, Rochel, N, Moras, D, Structural Genomics Consortium for Research on Gene Expression (SGCGES) | | Deposit date: | 2009-01-09 | | Release date: | 2009-08-11 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Silicon analogues of the RXR-selective retinoid agonist SR11237 (BMS649): chemistry and biology
Chemmedchem, 4, 2009
|
|
3A3Z
 
 | | Crystal structure of the human VDR ligand binding domain bound to the synthetic agonist compound 2alpha-methyl-AMCR277A(C23S) | | Descriptor: | (1S,2S,3R,5Z,7E,14beta,17alpha)-17-[(2S,4S)-4-(2-hydroxy-2-methylpropyl)-2-methyltetrahydrofuran-2-yl]-2-methyl-9,10-secoandrosta-5,7,10-triene-1,3-diol, SULFATE ION, Vitamin D3 receptor | | Authors: | Sato, Y, Antony, P, Huet, T, Sigueiro, R, Rochel, N, Moras, D, Structural Proteomics in Europe 2 (SPINE-2) | | Deposit date: | 2009-06-25 | | Release date: | 2010-02-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Structure-function relationships and crystal structures of the vitamin D receptor bound 2 alpha-methyl-(20S,23S)- and 2 alpha-methyl-(20S,23R)-epoxymethano-1 alpha,25-dihydroxyvitamin D3
J.Med.Chem., 53, 2010
|
|
3A78
 
 | | Crystal structure of the human VDR ligand binding domain bound to the natural metabolite 1alpha,25-dihydroxy-3-epi-vitamin D3 | | Descriptor: | (1S,3S,5Z,7E,14beta,17alpha)-9,10-secocholesta-5,7,10-triene-1,3,25-triol, SULFATE ION, Vitamin D3 receptor | | Authors: | Sato, Y, Sigueiro, R, Antony, P, Rochel, N, Moras, D. | | Deposit date: | 2009-09-18 | | Release date: | 2010-10-13 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural basis of the activation of the human Vitamin D receptor by the natural metabolite 1alpha,25-dihydroxy-3-epi-vitamin D3
To be Published
|
|
3A40
 
 | | Crystal structure of the human VDR ligand binding domain bound to the synthetic agonist compound 2alpha-methyl-AMCR277B(C23R) | | Descriptor: | (1S,2S,3R,5Z,7E,14beta,17alpha,23R)-23-(2-hydroxy-2-methylpropyl)-2-methyl-20,24-epoxy-9,10-secochola-5,7,10-triene-1,3-diol, SULFATE ION, Vitamin D3 receptor | | Authors: | Sato, Y, Antony, P, Huet, T, Sigueiro, R, Rochel, N, Moras, D, Structural Proteomics in Europe 2 (SPINE-2) | | Deposit date: | 2009-06-25 | | Release date: | 2010-02-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structure-function relationships and crystal structures of the vitamin D receptor bound 2 alpha-methyl-(20S,23S)- and 2 alpha-methyl-(20S,23R)-epoxymethano-1 alpha,25-dihydroxyvitamin D3
J.Med.Chem., 53, 2010
|
|
3JSV
 
 | | Crystal structure of mouse NEMO CoZi in complex with Lys63-linked di-ubiquitin | | Descriptor: | NF-kappa-B essential modulator, Ubiquitin | | Authors: | Yoshikawa, A, Sato, Y, Mimura, H, Yamashita, M, Yamagata, A, Fukai, S. | | Deposit date: | 2009-09-11 | | Release date: | 2009-10-27 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the NEMO ubiquitin-binding domain in complex with Lys 63-linked di-ubiquitin
Febs Lett., 583, 2009
|
|
9JBQ
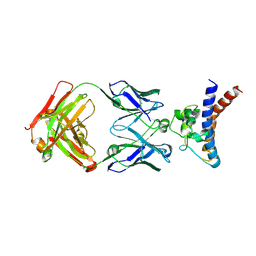 
 | | Structure of the complex between h1F3 Fab and PcrV fragment | | Descriptor: | Heavy chain, PcrV, light chain | | Authors: | Numata, S, Hara, T, Izawa, M, Okuno, Y, Sato, Y, Yamane, S, Maki, H, Sato, T, Yamano, Y. | | Deposit date: | 2024-08-27 | | Release date: | 2024-09-11 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Novel humanized anti-PcrV monoclonal antibody COT-143 protects mice from lethal Pseudomonas aeruginosa infection via inhibition of toxin translocation by the type III secretion system.
Antimicrob.Agents Chemother., 68, 2024
|
|
3P8X
 
 | | Synthesis, Structure, and Biological Activity of des-Side Chain Analogues of 1alpha,25-Dihydroxyvitamin D3 with Substituents at C-18 | | Descriptor: | (1R,3S,5Z)-5-{(2E)-2-[(3aR,7aS)-7a-(7-hydroxy-7-methyloctyl)octahydro-4H-inden-4-ylidene]ethylidene}-4-methylidenecyclohexane-1,3-diol, SULFATE ION, Vitamin D3 receptor | | Authors: | Rochel, N, Sato, Y, Moras, D. | | Deposit date: | 2010-10-15 | | Release date: | 2011-08-31 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Synthesis, Structure, and Biological Activity of des-Side Chain Analogues of 1 ,25-Dihydroxyvitamin D3 with Substituents at C-18
Chemmedchem, 6, 2011
|
|
4Z5T
 
 | | The nucleosome containing human H3.5 | | Descriptor: | DNA (146-MER), Histone H2A type 1-B/E, Histone H2B type 1-J, ... | | Authors: | Urahama, T, Harada, A, Maehara, K, Horikoshi, N, Sato, K, Sato, Y, Shiraishi, K, Sugino, N, Osakabe, A, Tachiwana, H, Kagawa, W, Kimura, H, Ohkawa, Y, Kurumizaka, H. | | Deposit date: | 2015-04-03 | | Release date: | 2016-02-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Histone H3.5 forms an unstable nucleosome and accumulates around transcription start sites in human testis.
Epigenetics Chromatin, 9, 2016
|
|
4K2A
 
 | | Crystal structure of haloalkane dehalogenase DbeA from Bradyrhizobium elkani USDA94 | | Descriptor: | ACETATE ION, CHLORIDE ION, Haloalkane dehalogenase | | Authors: | Prudnikova, T, Chaloupkova, R, Rezacova, P, Mozga, T, Koudelakova, T, Sato, Y, Kuty, M, Nagata, Y, Damborsky, J, Kuta Smatanova, I, Structure 2 Function Project (S2F) | | Deposit date: | 2013-04-08 | | Release date: | 2014-06-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and functional analysis of a novel haloalkane dehalogenase with two halide-binding sites.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
1KEG
 
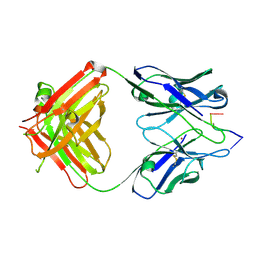 | | Antibody 64M-2 Fab complexed with dTT(6-4)TT | | Descriptor: | 5'-D(*TP*(64T)P*TP*T)-3', Anti-(6-4) photoproduct antibody 64M-2 Fab (heavy chain), Anti-(6-4) photoproduct antibody 64M-2 Fab (light chain), ... | | Authors: | Yokoyama, H, Mizutani, R, Satow, Y, Sato, K, Komatsu, Y, Ohtsuka, E, Nikaido, O. | | Deposit date: | 2001-11-15 | | Release date: | 2002-11-15 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of the DNA (6-4) photoproduct dTT(6-4)TT in complex with the 64M-2 antibody Fab fragment implies increased antibody-binding affinity by the flanking nucleotides.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
2V9Z
 
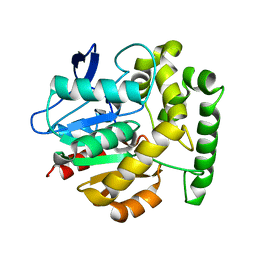 | | Structure of the Rhodococcus haloalkane dehalogenase mutant with enhanced enantioselectivity | | Descriptor: | HALOALKANE DEHALOGENASE | | Authors: | Koudelakova, T, Prokop, Z, Sato, Y, Lapkouski, M, Chovancova, E, Monincova, M, Jesenska, A, Emmer, J, Senda, T, Nagata, Y, Kuta Smatanova, I, Damborsky, J. | | Deposit date: | 2007-08-28 | | Release date: | 2008-09-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Rational Engineering of Rhodococcus Haloalkane Dehalogenase with Enhanced Enantioselectivity
To be Published
|
|
