4XX8
 
 | |
8SPN
 
 | | Crystal structure of macrophage migration inhibitory factor in complex with T614 | | Descriptor: | GLYCEROL, ISOPROPYL ALCOHOL, Iguratimod, ... | | Authors: | Pantouris, G, Lolis, E. | | Deposit date: | 2023-05-03 | | Release date: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Iguratimod, an allosteric inhibitor of macrophage migration inhibitory factor (MIF), prevents mortality and oxidative stress in a murine model of acetaminophen overdose.
Mol Med, 30, 2024
|
|
3DJI
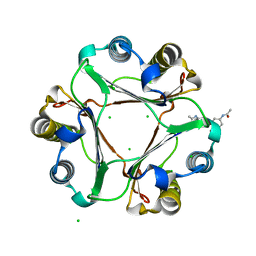 
 | | Crystal Structure of Macrophage Migration Inhibitory Factor Bound to an Acetaminophen Dimer Derived from NAPQI | | Descriptor: | CHLORIDE ION, Macrophage migration inhibitory factor, N,N'-(6,6'-dihydroxybiphenyl-3,3'-diyl)diacetamide | | Authors: | Crichlow, G.V, Lolis, E. | | Deposit date: | 2008-06-23 | | Release date: | 2008-12-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural and kinetic analyses of macrophage migration inhibitory factor active site interactions.
Biochemistry, 48, 2009
|
|
3GV3
 | |
6BG7
 
 | |
6BG6
 
 | |
8TT9
 
 | | X-ray structure of Macrophage Migration Inhibitory Factor (MIF) Covalently Bound to 4-hydroxyphenylpyruvate (HPP) | | Descriptor: | 3-(4-HYDROXY-PHENYL)PYRUVIC ACID, ISOPROPYL ALCOHOL, Macrophage migration inhibitory factor | | Authors: | Schroder, G.C, Meilleur, F, Nix, J.C, Crichlow, G.V, Lolis, E.J. | | Deposit date: | 2023-08-13 | | Release date: | 2024-08-28 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | X-ray structure of Macrophage Migration Inhibitory Factor (MIF) Covalently Bound to 4-hydroxyphenylpyruvate (HPP)
To Be Published
|
|
6OYB
 
 | |
6OYE
 
 | |
6OYG
 
 | |
6OY8
 | |
3KER
 
 | |
3KAN
 
 | |
3HP3
 
 | | Crystal structure of CXCL12 | | Descriptor: | CXCL12 protein, GOLD ION | | Authors: | Murphy, J.W, Lolis, E, Xiong, Y, Yuan, H, Crichlow, G. | | Deposit date: | 2009-06-03 | | Release date: | 2010-01-26 | | Last modified: | 2018-05-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Heterologous quaternary structure of CXCL12 and its relationship to the CC chemokine family
Proteins, 78, 2009
|
|
5UZY
 
 | |
5UMJ
 | |
5UMK
 
 | |
3CE4
 
 | |
4P0H
 
 | |
5BSC
 | |
5BSI
 
 | |
5BS9
 
 | |
7MRU
 
 | | Crystal structure of S62A MIF2 mutant | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, D-dopachrome decarboxylase | | Authors: | Murphy, E.L, Manjula, R, Murphy, J.W, Lolis, E. | | Deposit date: | 2021-05-09 | | Release date: | 2021-08-18 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | A structurally preserved allosteric site in the MIF superfamily affects enzymatic activity and CD74 activation in D-dopachrome tautomerase.
J.Biol.Chem., 297, 2021
|
|
7MSE
 
 | | High-resolution crystal structure of hMIF2 with tartrate at the active site | | Descriptor: | D-dopachrome decarboxylase, L(+)-TARTARIC ACID | | Authors: | Murphy, E.L, Manjula, R, Murphy, J.W, Lolis, E. | | Deposit date: | 2021-05-11 | | Release date: | 2021-08-18 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.27 Å) | | Cite: | A structurally preserved allosteric site in the MIF superfamily affects enzymatic activity and CD74 activation in D-dopachrome tautomerase.
J.Biol.Chem., 297, 2021
|
|
7MRV
 
 | | F100A mutant structure of MIF2 (D-DT) | | Descriptor: | D-dopachrome decarboxylase, SULFATE ION | | Authors: | Murphy, E.L, Manjula, R, Murphy, J.W, Lolis, E. | | Deposit date: | 2021-05-09 | | Release date: | 2021-08-18 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | A structurally preserved allosteric site in the MIF superfamily affects enzymatic activity and CD74 activation in D-dopachrome tautomerase.
J.Biol.Chem., 297, 2021
|
|
