3T55
 
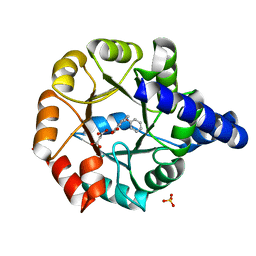 | | Crystal structure of Mycobacterium tuberculosis Indole Glycerol Phosphate Synthase (IGPS) in complex with Phenoxymethyl Benzoic Acid (PMBA) | | Descriptor: | 3-(phenoxymethyl)benzoic acid, Indole-3-glycerol phosphate synthase, MALONATE ION, ... | | Authors: | Reddy, M.C.M, Bruning, J.B, Thurman, C, Sacchettini, J.C, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2011-07-26 | | Release date: | 2012-08-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Structural Insights and Inhibition of Mycobacterium tuberculosis Indole Glycerol Phosphate Synthase (IGPS): an Essential Enzyme for Tryptophan Biosynthesis
To be Published
|
|
1XEQ
 
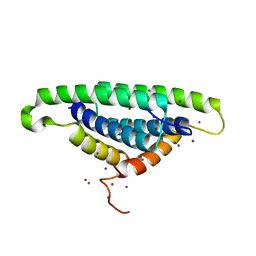 | | Crystal tructure of RNA binding domain of influenza B virus non-structural protein | | Descriptor: | BROMIDE ION, Nonstructural protein NS1 | | Authors: | Khan, J.A, Yin, C, Krug, R.M, Montelione, G.T, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2004-09-11 | | Release date: | 2005-09-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Conserved surface features form the double-stranded RNA binding site of non-structural protein 1 (NS1) from influenza A and B viruses.
J.Biol.Chem., 282, 2007
|
|
2PSS
 
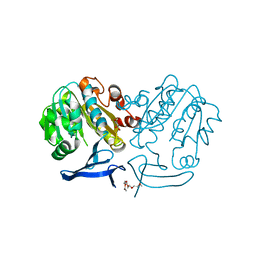 | | The structure of Plasmodium falciparum spermidine synthase in its apo-form | | Descriptor: | 2-(2-{2-[2-(2-METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHANOL, GLYCEROL, SULFATE ION, ... | | Authors: | Dufe, V.T, Qiu, W, Muller, I.B, Hui, R, Walter, R.D, Al-Karadaghi, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-05-07 | | Release date: | 2008-04-01 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of Plasmodium falciparum spermidine synthase in complex with the substrate decarboxylated S-adenosylmethionine and the potent inhibitors 4MCHA and AdoDATO.
J.Mol.Biol., 373, 2007
|
|
3T40
 
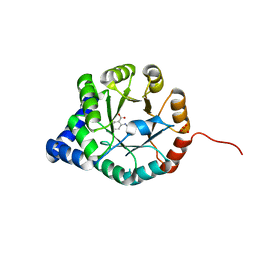 | | Crystal Structure of Mycobacterium tuberculosis Indole Glycerol Phosphate Synthase (IGPS) complex with N-2-Carboxyphenyl Glycine (CPG) | | Descriptor: | 2-[(carboxymethyl)amino]benzoic acid, Indole-3-glycerol phosphate synthase, SULFATE ION | | Authors: | Reddy, M.C.M, Bruning, J.B, Thurman, C, Sacchettini, J.C, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2011-07-25 | | Release date: | 2012-08-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.752 Å) | | Cite: | Structural Insights and Inhibition of Mycobacterium tuberculosis Indole Glycerol Phosphate Synthase (IGPS): an Essential Enzyme for Tryptophan Biosynthesis
To be Published
|
|
1YGY
 
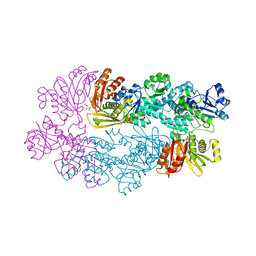 | |
3KUL
 
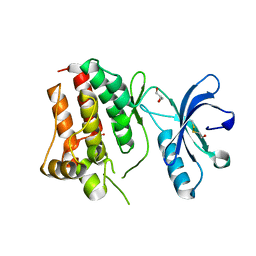 | | Kinase domain of human ephrin type-A receptor 8 (EPHA8) | | Descriptor: | BETA-MERCAPTOETHANOL, Ephrin type-A receptor 8, GLYCEROL, ... | | Authors: | Walker, J.R, Yermekbayeva, L, Kania, J, Bountra, C, Weigelt, J, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2009-11-27 | | Release date: | 2009-12-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Kinase Domain of Human Ephrin Type-A Receptor 8 (Epha8)
To be Published
|
|
4LG9
 
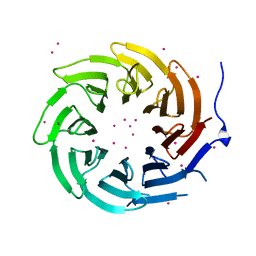 | | Crystal structure of TBL1XR1 WD40 repeats | | Descriptor: | F-box-like/WD repeat-containing protein TBL1XR1, UNKNOWN ATOM OR ION | | Authors: | Xu, C, Tempel, W, He, H, Wu, X, Seitova, A, Bountra, C, Arrowsmith, C.H, Edwards, A.M, Min, J, Structural Genomics Consortium (SGC) | | Deposit date: | 2013-06-27 | | Release date: | 2014-04-16 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Crystal structure of TBL1XR1 WD40 repeats
TO BE PUBLISHED
|
|
2OB4
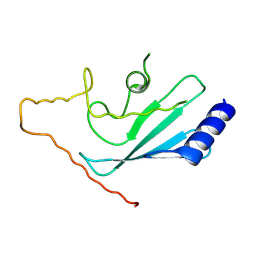 
 | | Human Ubiquitin-Conjugating Enzyme CDC34 | | Descriptor: | Ubiquitin-conjugating enzyme E2-32 kDa complementing | | Authors: | Neculai, D, Avvakumov, G.V, Xue, S, Walker, J.R, Mackenzie, F, Weigelt, J, Sundstrom, M, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Sicheri, F, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2006-12-18 | | Release date: | 2006-12-26 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | A human ubiquitin conjugating enzyme (E2)-HECT E3 ligase structure-function screen.
Mol Cell Proteomics, 11, 2012
|
|
4LDF
 
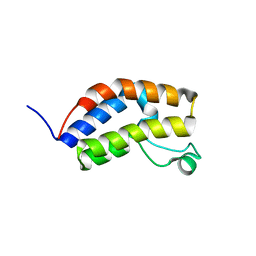 | | Crystal Structure of CpBRD2 from cryptosporidium, cgd3_3190 | | Descriptor: | GCN5 like acetylase + bromodomain, GLYCEROL, UNKNOWN ATOM OR ION | | Authors: | Wernimont, A.K, Loppnau, P, Fonseca, M, Knapp, S, Tempel, W, Bountra, C, Arrowsmith, C.H, Edwards, A.M, Hui, R, Mottaghi, K, Structural Genomics Consortium (SGC) | | Deposit date: | 2013-06-24 | | Release date: | 2013-07-24 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structure of CpBRD2 from cryptosporidium, cgd3_3190
TO BE PUBLISHED
|
|
3G15
 
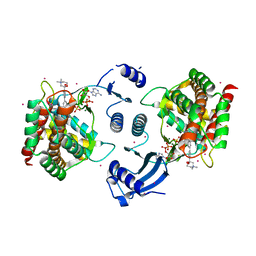 | | Crystal structure of human choline kinase alpha in complex with hemicholinium-3 and ADP | | Descriptor: | (2S,2'S)-2,2'-biphenyl-4,4'-diylbis(2-hydroxy-4,4-dimethylmorpholin-4-ium), ADENOSINE-5'-DIPHOSPHATE, Choline kinase alpha, ... | | Authors: | Hong, B.S, Tempel, W, Rabeh, W.M, MacKenzie, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Weigelt, J, Bochkarev, A, Park, H.W, Structural Genomics Consortium (SGC) | | Deposit date: | 2009-01-29 | | Release date: | 2009-02-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structures of human choline kinase isoforms in complex with hemicholinium-3: single amino acid near the active site influences inhibitor sensitivity.
J.Biol.Chem., 285, 2010
|
|
4LG7
 
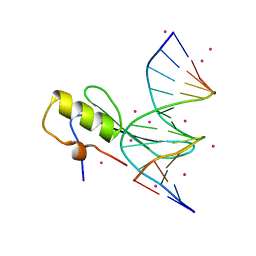 | | Crystal structure MBD4 MBD domain in complex with methylated CpG DNA | | Descriptor: | DNA (5'-D(*GP*CP*CP*AP*AP*(5CM)P*GP*TP*TP*GP*GP*C)-3'), Methyl-CpG-binding domain protein 4, UNKNOWN ATOM OR ION | | Authors: | Xu, C, Tempel, W, Wernimont, A.K, Bountra, C, Arrowsmith, C.H, Edwards, A.M, Min, J, Structural Genomics Consortium (SGC) | | Deposit date: | 2013-06-27 | | Release date: | 2013-07-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure MBD4 MBD domain in complex with methylated CpG DNA
To be Published
|
|
4M0R
 
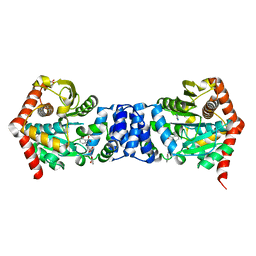 | | Trianthranilate-like analogue bound to anthranilate phosphoribosyltransferase (AnPRT; TrpD). | | Descriptor: | 2,6-bis[(2-carboxyphenyl)amino]benzoic acid, Anthranilate phosphoribosyltransferase, DIMETHYL SULFOXIDE, ... | | Authors: | Evans, G.L, Baker, E.N, Lott, J.S, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2013-08-02 | | Release date: | 2014-05-21 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Repurposing the Chemical Scaffold of the Anti-Arthritic Drug Lobenzarit to Target Tryptophan Biosynthesis in Mycobacterium tuberculosis.
Chembiochem, 15, 2014
|
|
2GRG
 
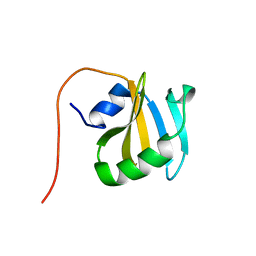 | | Solution NMR Structure of Protein YNR034W-A from Saccharomyces cerevisiae. Northeast Structural Genomics Consortium Target YT727; Ontario Center for Structural Proteomics Target yst6499. | | Descriptor: | hypothetical protein; Ynr034w-ap | | Authors: | Wu, B, Yee, A, Ramelot, T, Lemak, A, Semesi, A, Kennedy, M, Edward, A, Arrowsmith, C.H, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2006-04-24 | | Release date: | 2006-05-23 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of hypothetical protein yst6499 from Saccharomyces cerevisiae/ Northeast Structural Genomics Consortium Target YT727/ Ontario Center for Structural Proteomics Target yst6499
To be Published
|
|
2V40
 
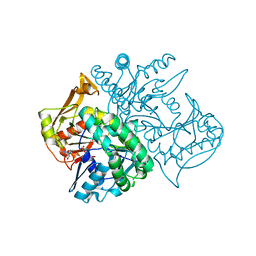 | | Human Adenylosuccinate synthetase isozyme 2 in complex with GDP | | Descriptor: | ADENYLOSUCCINATE SYNTHETASE ISOZYME 2, GUANOSINE-5'-DIPHOSPHATE | | Authors: | Welin, M, Moche, M, Arrowsmith, C, Berglund, H, Busam, R, Collins, R, Dahlgren, L.G, Edwards, A, Flodin, S, Flores, A, Graslund, S, Hammarstrom, M, Hallberg, B.M, Holmberg-Schiavone, L, Johansson, I, Kallas, A, Karlberg, T, Kotenyova, T, Lehtio, L, Nyman, T, Ogg, D, Persson, C, Sagemark, J, Stenmark, P, Sundstrom, M, Thorsell, A.G, Tresaugues, L, van den Berg, S, Weigelt, J, Nordlund, P, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-06-27 | | Release date: | 2007-07-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Human Adenylosuccinate Synthetase Isozyme 2 in Complex with Gdp
To be Published
|
|
2KCZ
 
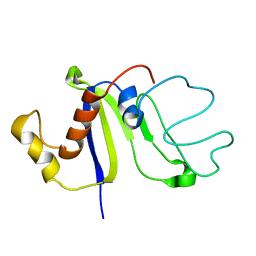 | | Solution NMR structure of the C-terminal domain of protein DR_A0006 from Deinococcus radiodurans. Northeast Structural Genomics Consortium Target DrR147D | | Descriptor: | Uncharacterized protein DR_A0006 | | Authors: | Mills, J.L, Ghosh, A, Garcia, E, Wang, H, Ciccosanti, C, Xiao, R, Nair, R, Everett, J.K, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-12-31 | | Release date: | 2009-01-20 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of the C-Terminal Domain of Protein DR_A0006 from Deinococcus radiodurans. Northeast Structural Genomics Consortium Target DrR147D.
To be Published
|
|
2KD1
 
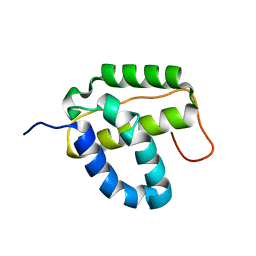 | | Solution NMR structure of the integrase-like domain from Bacillus cereus ordered locus BC_1272. Northeast Structural Genomics Consortium Target BcR268F | | Descriptor: | DNA integration/recombination/invertion protein | | Authors: | Rossi, P, Lee, H, Maglaqui, M, Foote, E.L, Buchwald, W.A, Jiang, M, Swapna, G.V.T, Nair, R, Xiao, R, Acton, T.B, Rost, B, Prestegard, J.H, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-12-31 | | Release date: | 2009-01-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of the Integrase-Like Domain from Bacillus cereus Ordered Locus BC_1272. Northeast Structural Genomics Consortium Target BcR268F.
To be Published
|
|
2KJ5
 
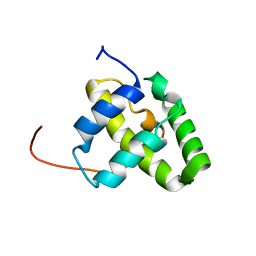 | | Solution NMR structure of a domain from a putative phage integrase protein Nmul_A0064 from Nitrosospira multiformis, Northeast Structural Genomics Consortium Target NmR46C | | Descriptor: | Phage integrase | | Authors: | Mills, J.L, Eletsky, A, Ghosh, A, Wang, D, Lee, H, Ciccosanti, C, Jiang, M, Nair, R, Rost, B, Swapna, G.V.T, Acton, T.B, Xiao, R, Everett, J.K, Prestegard, J.H, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-05-21 | | Release date: | 2009-06-02 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of a domain from a putative phage integrase protein Nmul_A0064 from Nitrosospira multiformis, Northeast Structural Genomics Consortium Target NmR46C.
To be Published
|
|
4NR6
 
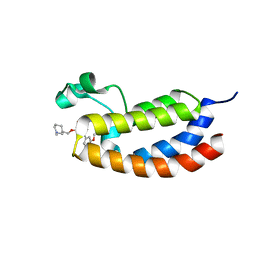 | | Crystal structure of the bromodomain of human CREBBP in complex with an oxazepin ligand | | Descriptor: | 1-[7-(3,4-dimethoxyphenyl)-9-{[(3R)-1-methylpiperidin-3-yl]methoxy}-2,3-dihydro-1,4-benzoxazepin-4(5H)-yl]propan-1-one, CREB-binding protein | | Authors: | Filippakopoulos, P, Picaud, S, Felletar, I, Hay, D, Fedorov, O, Martin, S, Chaikuad, A, von Delft, F, Brennan, P, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2013-11-26 | | Release date: | 2013-12-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Crystal structure of the bromodomain of human CREBBP in complex with an oxazepin ligand
To be Published
|
|
4NRB
 
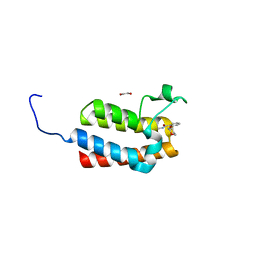 | | Crystal Structure of the bromodomain of human BAZ2B in complex with compound-1 N01197 | | Descriptor: | 1,2-ETHANEDIOL, Bromodomain adjacent to zinc finger domain protein 2B, N-methyl-2-(tetrahydro-2H-pyran-4-yloxy)benzamide | | Authors: | Muniz, J.R.C, Felletar, I, Chaikuad, A, Filippakopoulos, P, Ferguson, F.M, Fedorov, O, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Ciulli, A, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2013-11-26 | | Release date: | 2013-12-25 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Targeting low-druggability bromodomains: fragment based screening and inhibitor design against the BAZ2B bromodomain.
J.Med.Chem., 56, 2013
|
|
2K5N
 
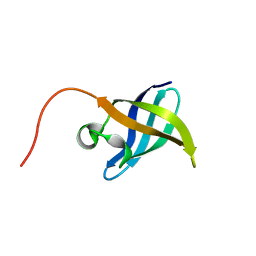 | | Solution NMR Structure of the N-Terminal Domain of Protein ECA1580 from Erwinia carotovora, Northeast Structural Genomics Consortium Target EwR156A | | Descriptor: | Putative cold-shock protein | | Authors: | Mills, J.L, Eletsky, A, Zhang, Q, Lee, D, Jiang, M, Ciccosanti, C, Xiao, R, Lui, J, Everett, J.K, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-30 | | Release date: | 2008-08-19 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of the Putative Cold Shock Protein from Erwinia carotovora: Northeast Structural Genomics Consortium Target EwR156a
To be Published
|
|
3DKM
 
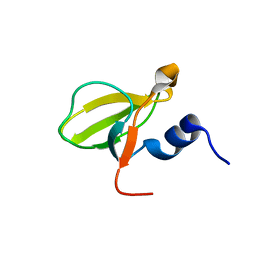 | | Crystal structure of the HECTD1 CPH domain | | Descriptor: | E3 ubiquitin-protein ligase HECTD1 | | Authors: | Walker, J.R, Qiu, L, Li, Y, Bountra, C, Wolkstrom, M, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2008-06-25 | | Release date: | 2008-08-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of the CPH domain of the E3 ubiquitin-protein ligase HECTD1.
To be Published
|
|
4NQN
 
 | | Crystal Structure of the bromodomain of human BRD9 in complex with a triazolo-phthalazine ligand | | Descriptor: | 2-chloro-N-[5-(3-methyl[1,2,4]triazolo[3,4-a]phthalazin-6-yl)-2-(morpholin-4-yl)phenyl]benzenesulfonamide, Bromodomain-containing protein 9 | | Authors: | Filippakopoulos, P, Picaud, S, Felletar, I, Martin, S, Fedorov, O, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2013-11-25 | | Release date: | 2013-12-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Crystal Structure of the bromodomain of human BRD9 in complex with a triazolo-phthalazine ligand
To be Published
|
|
4MR4
 
 | | Crystal Structure of the first bromodomain of human BRD4 in complex with a quinazolinone ligand (RVX-208) | | Descriptor: | 1,2-ETHANEDIOL, 2-[4-(2-hydroxyethoxy)-3,5-dimethylphenyl]-5,7-dimethoxyquinazolin-4(3H)-one, Bromodomain-containing protein 4 | | Authors: | Filippakopoulos, P, Picaud, S, Felletar, I, Martin, S, Fedorov, O, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Weigelt, J, Bountra, C, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2013-09-17 | | Release date: | 2013-11-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | RVX-208, an inhibitor of BET transcriptional regulators with selectivity for the second bromodomain.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
2KON
 
 | | NMR solution structure of CV_2116 from Chromobacterium violaceum. Northeast Structural Genomics Consortium Target CvT4(1-82) | | Descriptor: | Uncharacterized protein | | Authors: | Yang, Y, Ramelot, T.A, Cort, J.R, Garcia, M, Yee, A, Arrowsmith, C.H, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-09-24 | | Release date: | 2009-10-13 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of CV_2116 from Chromobacterium violaceum.Northeast Structural Genomics Consortium Target CvT4(1-82)
To be Published
|
|
4NR4
 
 | | Crystal structure of the bromodomain of human CREBBP in complex with an isoxazolyl-benzimidazole ligand | | Descriptor: | 1,2-ETHANEDIOL, 1-(4-chlorobenzyl)-5-(3,5-dimethyl-1,2-oxazol-4-yl)-1H-benzimidazole, CREB-binding protein, ... | | Authors: | Filippakopoulos, P, Picaud, S, Felletar, I, Hay, D, Fedorov, O, Martin, S, von Delft, F, Brennan, P, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2013-11-26 | | Release date: | 2013-12-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Crystal structure of the bromodomain of human CREBBP in complex with an isoxazolyl-benzimidazole ligand
TO BE PUBLISHED
|
|
