5UFU
 
 | | Structure of AMPK bound to activator | | Descriptor: | 1,4:3,6-dianhydro-2-O-(6-chloro-5-{4-[1-(hydroxymethyl)cyclopropyl]phenyl}-1H-benzimidazol-2-yl)-D-mannitol, 5'-AMP-activated protein kinase catalytic subunit alpha-1, 5'-AMP-activated protein kinase subunit beta-1, ... | | Authors: | Calabrese, M.F, Kurumbail, R.G. | | Deposit date: | 2017-01-05 | | Release date: | 2017-05-17 | | Method: | X-RAY DIFFRACTION (3.45 Å) | | Cite: | Activation of Skeletal Muscle AMPK Promotes Glucose Disposal and Glucose Lowering in Non-human Primates and Mice.
Cell Metab., 25, 2017
|
|
5WEQ
 
 | | The crystal structure of a MR78 mutant | | Descriptor: | MR78 mutant Fab heavy chain, MR78 mutant light chain | | Authors: | Dong, J, Williamson, L.E, Crowe, J.E. | | Deposit date: | 2017-07-10 | | Release date: | 2018-04-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Role of Non-local Interactions between CDR Loops in Binding Affinity of MR78 Antibody to Marburg Virus Glycoprotein.
Structure, 25, 2017
|
|
6DLB
 
 | |
6DLA
 
 | |
6DL8
 
 | |
5V7U
 
 | |
5V7R
 
 | |
6W8E
 
 | |
6H0E
 
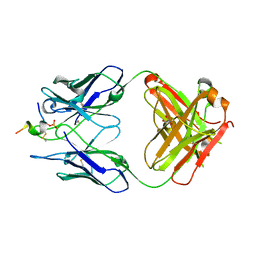 | | FAB dmCBTAU-22.1 IN COMPLEX WITH TAU PEPTIDE V1088-23 | | Descriptor: | GLYCEROL, HUMAN FAB ANTIBODY FRAGMENT OF dmCBTAU-22.1, Microtubule-associated protein tau | | Authors: | Juraszek, J, Steinbacher, S. | | Deposit date: | 2018-07-09 | | Release date: | 2018-07-25 | | Last modified: | 2020-03-11 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Enhancement of therapeutic potential of a naturally occurring human antibody targeting a phosphorylated Ser422containing epitope on pathological tau.
Acta Neuropathol Commun, 6, 2018
|
|
6H06
 
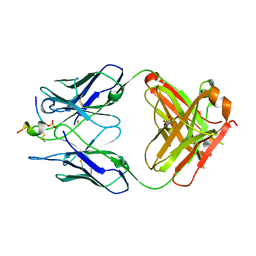 | | FAB CBTAU-22.1 IN COMPLEX WITH TAU PEPTIDE V1088-5 | | Descriptor: | GLYCEROL, HUMAN FAB ANTIBODY FRAGMENT OF CBTAU-22.1, HUMAN FAB ANTIBODY FRAGMENT OF HCBTAU-22.1, ... | | Authors: | Juraszek, J, Steinbacher, S. | | Deposit date: | 2018-07-06 | | Release date: | 2018-07-25 | | Last modified: | 2020-03-11 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Enhancement of therapeutic potential of a naturally occurring human antibody targeting a phosphorylated Ser422containing epitope on pathological tau.
Acta Neuropathol Commun, 6, 2018
|
|
3B9S
 
 | | Macrophage Migration Inhibitory Factor (MIF) complexed with Inhibitor, 4-IPP. | | Descriptor: | 4-phenylpyrimidine, GLYCEROL, Macrophage Migration Inhibitory Factor, ... | | Authors: | Zierow, S, Crichlow, G, Lolis, E. | | Deposit date: | 2007-11-06 | | Release date: | 2008-09-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A novel, macrophage migration inhibitory factor suicide substrate inhibits motility and growth of lung cancer cells.
Cancer Res., 68, 2008
|
|
1BH1
 
 | | STRUCTURAL STUDIES OF D-PRO MELITTIN, NMR, 20 STRUCTURES | | Descriptor: | MELITTIN | | Authors: | Barnham, K.J, Hewish, D, Werkmeister, J, Curtain, C, Kirkpatrick, A, Bartone, N, Liu, S.T, Norton, R, Rivett, D. | | Deposit date: | 1998-06-11 | | Release date: | 1999-01-06 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Structure and activity of D-Pro14 melittin.
J.Protein Chem., 21, 2002
|
|
4DAW
 
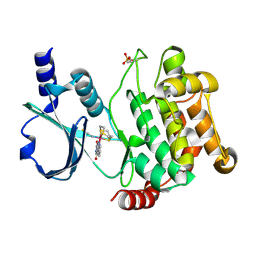 | | Crystal structure of PAK1 kinase domain with the ruthenium phthalimide complex | | Descriptor: | Serine/threonine-protein kinase PAK 1, [1,3-dioxo-6-(pyridin-2-yl-kappaN)-2,3-dihydro-1H-isoindol-5-yl-kappaC~5~][(thioxomethylidene)azanido-kappaN](1,4,7-trithionane-kappa~3~S~1~,S~4~,S~7~)ruthenium | | Authors: | Maksimoska, J, Marmorstein, R. | | Deposit date: | 2012-01-13 | | Release date: | 2012-03-14 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The art of filling protein pockets efficiently with octahedral metal complexes.
Angew.Chem.Int.Ed.Engl., 51, 2012
|
|
5ERB
 
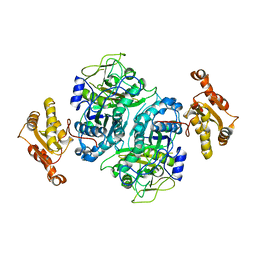 | |
3TDM
 
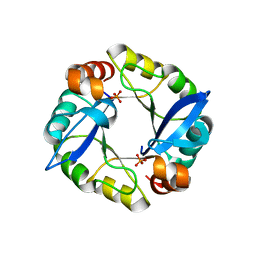 | | Computationally designed TIM-barrel protein, HalfFLR | | Descriptor: | Computationally designed two-fold symmetric TIM-barrel protein, FLR (half molecule), PHOSPHATE ION | | Authors: | Harp, J.M, Fortenberry, C, Bowman, E, Profitt, W, Dorr, B, Mizoue, L. | | Deposit date: | 2011-08-11 | | Release date: | 2011-11-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Exploring symmetry as an avenue to the computational design of large protein domains.
J.Am.Chem.Soc., 133, 2011
|
|
6W7P
 
 | |
6X1T
 
 | | Structure of pHis Fab (SC50-3) in complex with pHis mimetic peptide | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Kalagiri, R, Stanfield, R.L, Wilson, I.A, Hunter, T. | | Deposit date: | 2020-05-19 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Structural basis for differential recognition of phosphohistidine-containing peptides by 1-pHis and 3-pHis monoclonal antibodies.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
6X1W
 
 | | Structure of pHis Fab (SC56-2) in complex with pHis mimetic peptide | | Descriptor: | ACLYana-3-pTza peptide, SC56-2 Heavy chain, SC56-2 Light chain | | Authors: | Kalagiri, R, Stanfield, R, Wilson, I.A, Hunter, T. | | Deposit date: | 2020-05-19 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural basis for differential recognition of phosphohistidine-containing peptides by 1-pHis and 3-pHis monoclonal antibodies.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
4YAQ
 
 | | Crystal structure of a computationally optimized PG9 mutant | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, PG9_N100FY Fab heavy chain, ... | | Authors: | Murrell, S, Julien, J.P, Wilson, I.A. | | Deposit date: | 2015-02-17 | | Release date: | 2015-06-03 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Redesigned HIV antibodies exhibit enhanced neutralizing potency and breadth.
J.Clin.Invest., 125, 2015
|
|
6X1V
 
 | | Structure of pHis Fab (SC44-8) in complex with pHis mimetic peptide | | Descriptor: | 1,2-ETHANEDIOL, ACLYana-3-pTza peptide, SC44-8 Heavy chain, ... | | Authors: | Kalagiri, R, Stanfield, R, Wilson, I.A, Hunter, T. | | Deposit date: | 2020-05-19 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Structural basis for differential recognition of phosphohistidine-containing peptides by 1-pHis and 3-pHis monoclonal antibodies.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
6X1U
 
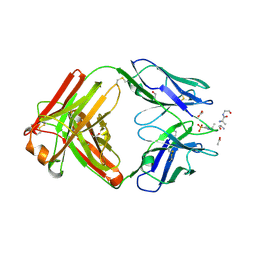 | | Structure of pHis Fab (SC39-4) in complex with pHis mimetic peptide | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, ACLYana-3-pTza peptide, ... | | Authors: | Kalagiri, R, Stanfield, R, Wilson, I.A, Hunter, T. | | Deposit date: | 2020-05-19 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.641 Å) | | Cite: | Structural basis for differential recognition of phosphohistidine-containing peptides by 1-pHis and 3-pHis monoclonal antibodies.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
6X1S
 
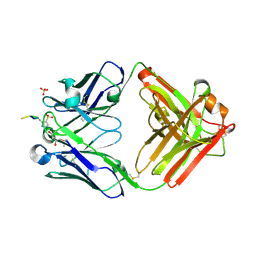 | | Structure of pHis Fab (SC1-1) in complex with pHis mimetic peptide | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, NM23-1-pTza peptide, SC1-1 Heavy chain, ... | | Authors: | Kalagiri, R, Stanfield, R.L, Wilson, I.A, Hunter, T. | | Deposit date: | 2020-05-19 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural basis for differential recognition of phosphohistidine-containing peptides by 1-pHis and 3-pHis monoclonal antibodies.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
6WC6
 
 | |
8DBZ
 
 | |
5J3D
 | |
