-Search query
-Search result
Showing 1 - 50 of 2,062 items for (author: you & z)

EMDB entry, No image
EMDB-38418: 
A neutralizing nanobody VHH60 against wt SARS-CoV-2

PDB-8xki: 
A neutralizing nanobody VHH60 against wt SARS-CoV-2

EMDB entry, No image
EMDB-60384: 
Cryo-EM structure of the GPR15L(C11)-bound GPR15 complex

PDB-8zqe: 
Cryo-EM structure of the GPR15L(C11)-bound GPR15 complex

EMDB-40971: 
Atomic model of the mammalian mouse Mediator complex with CKM module

EMDB-16759: 
Cryo-EM structure of retinal-free proteoopsin bound to decanoate

EMDB-16795: 
Cryo-EM structure of pentameric proteorhodopsin A18L mutant

EMDB-16796: 
Cryo-EM structure of hexameric proteorhodopsin A18L mutant

EMDB-37133: 
Cryo-EM structure of an intermediate-state complex during the process of photosystem II repair

EMDB-37265: 
Overall cryo-EM map of an intermediate-state complex during the process of photosystem II repair

EMDB-37288: 
A focused cryo-EM map of an intermediate-state complex during the process of photosystem II repair (Part1)

EMDB-37289: 
A focused cryo-EM map of an intermediate-state complex during the process of photosystem II repair (Part 2)

EMDB-60026: 
Cryo-EM structure of an intermediate-state PSII-PRF2' complex during the process of photosystem II repair

PDB-8kde: 
Cryo-EM structure of an intermediate-state complex during the process of photosystem II repair

PDB-8zee: 
Cryo-EM structure of an intermediate-state PSII-PRF2' complex during the process of photosystem II repair

EMDB-37944: 
Structure of 26RFa-pyroglutamylated RFamide peptide receptor complex

PDB-8wz2: 
Structure of 26RFa-pyroglutamylated RFamide peptide receptor complex

EMDB-41569: 
Cryo-EM structure of HmAb64 scFv in complex with CNE40 SOSIP trimer

PDB-8tr3: 
Cryo-EM structure of HmAb64 scFv in complex with CNE40 SOSIP trimer
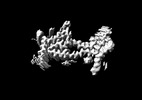
EMDB-37445: 
Cryo-EM structure of human disease-associated P301L Tau amyloid fibril from mouse brain
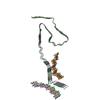
PDB-8wcp: 
Cryo-EM structure of human disease-associated P301L Tau amyloid fibril from mouse brain

EMDB-40968: 
Atomic model of the mammalian Mediator complex with MED26 subunit
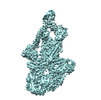
EMDB-40972: 
CryoEM map of TR-TRAP

EMDB-38148: 
Cryo-EM structure of the cortistatin 17-bound Somatostatin receptor 5-Gi protein complex

PDB-8x8l: 
Cryo-EM structure of the cortistatin 17-bound Somatostatin receptor 5-Gi protein complex

EMDB-40975: 
CryoEM map of mouse mediator complex with alternate conformation CKM module

EMDB-38150: 
Cryo-EM structure of the octreotide-bound Somatostatin receptor 5-Gi protein complex

PDB-8x8n: 
Cryo-EM structure of the octreotide-bound Somatostatin receptor 5-Gi protein complex

EMDB-37858: 
SpCas9-MMLV RT-pegRNA-target DNA complex (termination)

EMDB-37859: 
SpCas9-MMLV RT-pegRNA-target DNA complex (initiation)

EMDB-37860: 
SpCas9-pegRNA-target DNA complex (pre-initiation)

EMDB-37861: 
SpCas9-MMLV RT-pegRNA-target DNA complex (elongation 16-nt)

EMDB-39253: 
SpCas9-MMLV RT-pegRNA-target DNA complex (elongation 28-nt)

PDB-8wus: 
SpCas9-MMLV RT-pegRNA-target DNA complex (termination)

PDB-8wut: 
SpCas9-MMLV RT-pegRNA-target DNA complex (initiation)

PDB-8wuu: 
SpCas9-pegRNA-target DNA complex (pre-initiation)

PDB-8wuv: 
SpCas9-MMLV RT-pegRNA-target DNA complex (elongation 16-nt)

PDB-8ygj: 
SpCas9-MMLV RT-pegRNA-target DNA complex (elongation 28-nt)

EMDB-39582: 
Cryo-EM structure of the amthamine-bound H2R-Gs complex

EMDB-39583: 
Cryo-EM structure of the histamine-bound H3R-Gi complex

EMDB-39584: 
Cryo-EM structure of the immepip-bound H3R-Gi complex

PDB-8yut: 
Cryo-EM structure of the amthamine-bound H2R-Gs complex

PDB-8yuu: 
Cryo-EM structure of the histamine-bound H3R-Gi complex

PDB-8yuv: 
Cryo-EM structure of the immepip-bound H3R-Gi complex

EMDB-36987: 
Structure of CUL3-RBX1-KLHL22 complex without CUL3 NA motif

PDB-8k9i: 
Structure of CUL3-RBX1-KLHL22 complex without CUL3 NA motif

EMDB-36961: 
Structure of CUL3-RBX1-KLHL22 complex

EMDB-39719: 
Focused map of CUL3-RBX1-KLHL22 dimerization region

EMDB-39720: 
Consensus map of CUL3-RBX1-KLHL22 complex

EMDB-39725: 
Cryo-EM structure of CUL3-RBX1-KLHL22 complex --C1 Symmetry
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
