-Search query
-Search result
Showing all 36 items for (author: benito & jm)

EMDB-14727: 
3D reconstruction of the cylindrical assembly of DnaJA2 by imposing D5 symmetry

EMDB-14729: 
3D reconstruction of the cylindrical assembly of DnaJA2 without symmetry imposition

EMDB-14736: 
3D reconstruction of the cylindrical assembly of DnaJA2 delta G/F by imposing D5 symmetry

EMDB-14706: 
3D reconstruction of the cylindrical assembly of DnaJA2 delta G/F without symmetry imposition

EMDB-4423: 
Helical part of the influenza A virus ribonucleoprotein. Conformation 3.

EMDB-0175: 
Helical part of the influenza A virus ribonucleoprotein. Conformation 1.

EMDB-4426: 
Helical part of the influenza A virus ribonucleoprotein. Conformation 4.

EMDB-4430: 
Helical part of the influenza A virus ribonucleoprotein. Conformation 5.

EMDB-4412: 
Helical part of the influenza A virus ribonucleoprotein. Conformation 2.

EMDB-0323: 
Structural insights into the ability of nucleoplasmin to assemble and chaperone histone octamers for DNA deposition

EMDB-4293: 
Hsp70:p53-TMGST:CHIP

EMDB-4294: 
Hsp90:p53-TMGST:CHIP

EMDB-3753: 
Negative stain 3D single particle reconstruction of the isolated surface adhesin P140/P110 complex of Mycoplasma genitalium.

EMDB-3756: 
Sub-tomogram average of the surface adhesin (NAP) complex from Mycoplasma genitalium cells by cryo-electron tomography.

EMDB-2447: 
Electron microscopy of the complex formed by chaperones TBCE and TBCB and alpha-tubulin

EMDB-5713: 
Electron microscopy of negatively-stained gp12 tubular protein of T7 bacteriophage

EMDB-5689: 
Cryo-electron microscopy of T7 tail complex

EMDB-5690: 
Cryo-electron microscopy of T7 tail complex formed by gp8, gp11, and gp12 proteins
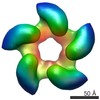
EMDB-2355: 
Negative staining three dimensional reconstruction of bacteriophage T7 large terminase

EMDB-2356: 
Negative staining three-dimensional reconstruction of bacteriophage T7 connector/terminase complex

EMDB-2205: 
Cryo-electron microscopy reconstruction of the helical part of influenza A virus ribonucleoprotein isolated from virions.

EMDB-2206: 
Negative stained electron microscopy reconstruction of the loop of influenza A virus ribonucleoprotein isolated from virions.

EMDB-2207: 
Negative stained electron microscopy reconstruction of the viral polymerase of influenza A virus ribonucleoprotein isolated from virions. Conformation 1.

EMDB-2208: 
Negative stained electron microscopy reconstruction of the viral polymerase of influenza A virus ribonucleoprotein isolated from virions. Conformation 2.

EMDB-5505: 
Hexameric structure of the conjugative VirB4 ATPase TrwK: new insights into a functional and phylogenetic relationship with DNA translocases

EMDB-1774: 
Understanding Ribosome Assembly: The Structure of in vivo Assembled Immature 30S Subunits Revealed by Cryo-Electron Microscopy

EMDB-1775: 
Understanding Ribosome Assembly: the Structure of in vivo Assembled Immature 30S Subunits Revealed by Cryo-electron Microscopy

EMDB-1810: 
Phage T7 empty head.

EMDB-1777: 
Structure of the complex Nucleoplamsin:H2A-H2B histones.

EMDB-1778: 
Structure of the Nuclear Chaperone Nucleoplamsin.

EMDB-1603: 
12 Angstrom resolution cryo-electron microscopy reconstruction of a recombinant active ribonucleoprotein particle of influenza virus (9-fold symmetrized).

EMDB-1231: 
Structure of the connector of bacteriophage T7 at 8A resolution: structural homologies of a basic component of a DNA translocating machinery.

EMDB-1161: 
Maturation of phage T7 involves structural modification of both shell and inner core components.

EMDB-1162: 
Maturation of phage T7 involves structural modification of both shell and inner core components.

EMDB-1163: 
Maturation of phage T7 involves structural modification of both shell and inner core components.

EMDB-1164: 
Maturation of phage T7 involves structural modification of both shell and inner core components.
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
