[English] 日本語
 Yorodumi
Yorodumi- EMDB-3756: Sub-tomogram average of the surface adhesin (NAP) complex from My... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3756 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
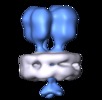
| Title | Sub-tomogram average of the surface adhesin (NAP) complex from Mycoplasma genitalium cells by cryo-electron tomography. | |||||||||
 Map data Map data | Sub-tomogram average of the surface adhesin (NAP) complex from Mycoplasma genitalium cells by cryo-electron tomography. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Mycoplasma genitalium G37 (bacteria) Mycoplasma genitalium G37 (bacteria) | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 26.0 Å | |||||||||
 Authors Authors | Scheffer MP / Gonzalez-Gonzalez L / Frangakis AS | |||||||||
 Citation Citation |  Journal: Mol Microbiol / Year: 2017 Journal: Mol Microbiol / Year: 2017Title: Structural characterization of the NAP; the major adhesion complex of the human pathogen Mycoplasma genitalium. Authors: Margot P Scheffer / Luis Gonzalez-Gonzalez / Anja Seybert / Mercè Ratera / Michael Kunz / José M Valpuesta / Ignacio Fita / Enrique Querol / Jaume Piñol / Jaime Martín-Benito / Achilleas S Frangakis /   Abstract: Mycoplasma genitalium, the causative agent of non-gonococcal urethritis and pelvic inflammatory disease in humans, is a small eubacterium that lacks a peptidoglycan cell wall. On the surface of its ...Mycoplasma genitalium, the causative agent of non-gonococcal urethritis and pelvic inflammatory disease in humans, is a small eubacterium that lacks a peptidoglycan cell wall. On the surface of its plasma membrane is the major surface adhesion complex, known as NAP that is essential for adhesion and gliding motility of the organism. Here, we have performed cryo-electron tomography of intact cells and detergent permeabilized M. genitalium cell aggregates, providing sub-tomogram averages of free and cell-attached NAPs respectively, revealing a tetrameric complex with two-fold rotational (C2) symmetry. Each NAP has two pairs of globular lobes (named α and β lobes), arranged as a dimer of heterodimers with each lobe connected by a stalk to the cell membrane. The β lobes are larger than the α lobes by 20%. Classification of NAPs showed that the complex can tilt with respect to the cell membrane. A protein complex containing exclusively the proteins P140 and P110, was purified from M. genitalium and was structurally characterized by negative-stain single particle EM reconstruction. The close structural similarity found between intact NAPs and the isolated P140/P110 complexes, shows that dimers of P140/P110 heterodimers are the only components of the extracellular region of intact NAPs in M. genitalium. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3756.map.gz emd_3756.map.gz | 78 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3756-v30.xml emd-3756-v30.xml emd-3756.xml emd-3756.xml | 8.6 KB 8.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3756.png emd_3756.png | 86.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3756 http://ftp.pdbj.org/pub/emdb/structures/EMD-3756 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3756 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3756 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3756.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3756.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sub-tomogram average of the surface adhesin (NAP) complex from Mycoplasma genitalium cells by cryo-electron tomography. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 6.7 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Sub-tomogram average of the surface adhesin (NAP) complex from My...
| Entire | Name: Sub-tomogram average of the surface adhesin (NAP) complex from Mycoplasma genitalium cells by cryo-electron tomography. |
|---|---|
| Components |
|
-Supramolecule #1: Sub-tomogram average of the surface adhesin (NAP) complex from My...
| Supramolecule | Name: Sub-tomogram average of the surface adhesin (NAP) complex from Mycoplasma genitalium cells by cryo-electron tomography. type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Mycoplasma genitalium G37 (bacteria) Mycoplasma genitalium G37 (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 26.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Super-sampling SART / Number subtomograms used: 1500 |
|---|---|
| Extraction | Number tomograms: 2 / Number images used: 1500 / Software - Name:  MATLAB MATLAB |
| CTF correction | Software - Name: Super-sampling SART |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















