[English] 日本語
 Yorodumi
Yorodumi- EMDB-4412: Helical part of the influenza A virus ribonucleoprotein. Conforma... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4412 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Helical part of the influenza A virus ribonucleoprotein. Conformation 2. | |||||||||
 Map data Map data | Influenza A nucleoprotein docked into 3D helical structure of the wild type ribonucleoprotein complex obtained using cryoEM. Conformation s. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Influenza A virus Ribonucleoprotein RNA binding protein / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative stranded viral RNA replication / helical viral capsid / viral penetration into host nucleus / host cell / viral nucleocapsid / ribonucleoprotein complex / symbiont entry into host cell / host cell nucleus / structural molecule activity / RNA binding / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Influenza A virus (A/Wilson-Smith/1933(H1N1)) Influenza A virus (A/Wilson-Smith/1933(H1N1)) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 10.0 Å | |||||||||
 Authors Authors | Coloma R / Arranz R | |||||||||
| Funding support |  Spain, 2 items Spain, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2020 Journal: Nat Microbiol / Year: 2020Title: Structural insights into influenza A virus ribonucleoproteins reveal a processive helical track as transcription mechanism. Authors: Rocío Coloma / Rocío Arranz / José M de la Rosa-Trevín / Carlos O S Sorzano / Sandie Munier / Diego Carlero / Nadia Naffakh / Juan Ortín / Jaime Martín-Benito /    Abstract: The influenza virus genome consists of eight viral ribonucleoproteins (vRNPs), each consisting of a copy of the polymerase, one of the genomic RNA segments and multiple copies of the nucleoprotein ...The influenza virus genome consists of eight viral ribonucleoproteins (vRNPs), each consisting of a copy of the polymerase, one of the genomic RNA segments and multiple copies of the nucleoprotein arranged in a double helical conformation. vRNPs are macromolecular machines responsible for messenger RNA synthesis and genome replication, that is, the formation of progeny vRNPs. Here, we describe the structural basis of the transcription process. The mechanism, which we call the 'processive helical track', is based on the extreme flexibility of the helical part of the vRNP that permits a sliding movement between both antiparallel nucleoprotein-RNA strands, thereby allowing the polymerase to move over the genome while bound to both RNA ends. Accordingly, we demonstrate that blocking this movement leads to inhibition of vRNP transcriptional activity. This mechanism also reveals a critical role of the nucleoprotein in maintaining the double helical structure throughout the copying process to make the RNA template accessible to the polymerase. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4412.map.gz emd_4412.map.gz | 6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4412-v30.xml emd-4412-v30.xml emd-4412.xml emd-4412.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4412.png emd_4412.png | 158.1 KB | ||
| Filedesc metadata |  emd-4412.cif.gz emd-4412.cif.gz | 6.3 KB | ||
| Others |  emd_4412_additional.map.gz emd_4412_additional.map.gz | 2.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4412 http://ftp.pdbj.org/pub/emdb/structures/EMD-4412 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4412 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4412 | HTTPS FTP |
-Related structure data
| Related structure data |  6i54MC  0175C  4423C  4426C  4430C  6h9gC  6i7bC  6i7mC  6i85C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4412.map.gz / Format: CCP4 / Size: 6.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4412.map.gz / Format: CCP4 / Size: 6.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
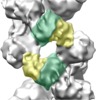
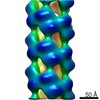
| Annotation | Influenza A nucleoprotein docked into 3D helical structure of the wild type ribonucleoprotein complex obtained using cryoEM. Conformation s. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.26 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Resolution map.
| File | emd_4412_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Resolution map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Influenza A virus
| Entire | Name:   Influenza A virus Influenza A virus |
|---|---|
| Components |
|
-Supramolecule #1: Influenza A virus
| Supramolecule | Name: Influenza A virus / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Influenza A virus (A/Wilson-Smith/1933(H1N1)) Influenza A virus (A/Wilson-Smith/1933(H1N1)) |
-Macromolecule #1: Nucleoprotein
| Macromolecule | Name: Nucleoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Influenza A virus (A/Wilson-Smith/1933(H1N1)) Influenza A virus (A/Wilson-Smith/1933(H1N1)) |
| Molecular weight | Theoretical: 43.079227 KDa |
| Sequence | String: NATEIRASVG KMIDGIGRFY IQMCTELKLS DYEGRLIQNS LTIERMVLSA FDTGGPIYRR VDGKWRRELI LYDKEEIRRI WRQANNGDD ATAGLTHMMI WHSNLNDATY QRTRALVRTG MDPRMCSLMQ GSTLPRRSGA AGAAVKGVGT MVMELIRMIK R GINRRTRI ...String: NATEIRASVG KMIDGIGRFY IQMCTELKLS DYEGRLIQNS LTIERMVLSA FDTGGPIYRR VDGKWRRELI LYDKEEIRRI WRQANNGDD ATAGLTHMMI WHSNLNDATY QRTRALVRTG MDPRMCSLMQ GSTLPRRSGA AGAAVKGVGT MVMELIRMIK R GINRRTRI AYERMCNILK GKFQTAAQRT MVDQVRESRN PGNAEFEDLI FLARSALILR GSVAHKSCLP ACVYGSAVAS GY DFEREGY SLVGIDPFRL LQNSQVYSLI RPNENPAHKS QLVWMACHSA AFEDLRVSSF IRGTKVVPRG KLSTRGVQIA SNE NMETME SSTLELRSRY WAIRTRSGGN TDVSFQGRGV FELSDEKATS PIVPSFDMSN EGSYFF UniProtKB: Nucleoprotein |
-Macromolecule #2: Influenza virus nucleoprotein
| Macromolecule | Name: Influenza virus nucleoprotein / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Influenza A virus (A/Wilson-Smith/1933(H1N1)) Influenza A virus (A/Wilson-Smith/1933(H1N1)) |
| Molecular weight | Theoretical: 2.107346 KDa |
| Sequence | String: SSGQISIQPT FSVQRNLPF |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Component - Concentration: 50.0 mM / Component - Formula: Tris-HCl / Component - Name: TN buffer / Details: TN buffer (50 mM Tris-HCl, 150 mM KCl) |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER/RHODIUM / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA EM CPC |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Frames/image: 3-68 / Number grids imaged: 1 / Number real images: 420 / Average electron dose: 2.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Number classes used: 1 Applied symmetry - Helical parameters - Δz: 34.47 Å Applied symmetry - Helical parameters - Δ&Phi: -68.33 ° Applied symmetry - Helical parameters - Axial symmetry: D1 (2x1 fold dihedral) Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 10.0 Å / Resolution method: OTHER / Software - Name: RELION (ver. 2.0) Details: Local resolution calculated using MonoRes software. Vilas et al. Structure. 2018 Feb 6;26(2):337-344.e4. doi: 10.1016/j.str Number images used: 4806 |
|---|---|
| Segment selection | Number selected: 137461 / Software - Name: Xmipp (ver. 3.0) / Details: Manual picking |
| Startup model | Type of model: OTHER Details: Initial model created using Iterative Helical Real Space Reconstruction |
| Final angle assignment | Type: NOT APPLICABLE / Software - Name: RELION (ver. 2.0) |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
| Output model |  PDB-6i54: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)