[English] 日本語
 Yorodumi
Yorodumi- EMDB-4804: Cryo-EM informed directed evolution of Nitrilase 4 leads to a cha... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4804 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM informed directed evolution of Nitrilase 4 leads to a change in quaternary structure. | |||||||||
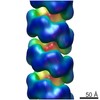
 Map data Map data | AtNIT4 R95T nitrilase filament | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | helical reconstruction / negative staining / Resolution: 20.0 Å | |||||||||
 Authors Authors | Mulelu AE / Woodward JD | |||||||||
| Funding support |  South Africa, 1 items South Africa, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2019 Journal: Commun Biol / Year: 2019Title: Cryo-EM and directed evolution reveal how nitrilase specificity is influenced by its quaternary structure. Authors: Andani E Mulelu / Angela M Kirykowicz / Jeremy D Woodward /  Abstract: Nitrilases are helical enzymes that convert nitriles to acids and/or amides. All plants have a nitrilase 4 homolog specific for ß-cyanoalanine, while in some plants neofunctionalization has produced ...Nitrilases are helical enzymes that convert nitriles to acids and/or amides. All plants have a nitrilase 4 homolog specific for ß-cyanoalanine, while in some plants neofunctionalization has produced nitrilases with altered specificity. Plant nitrilase substrate size and specificity correlate with helical twist, but molecular details of this relationship are lacking. Here we determine, to our knowledge, the first close-to-atomic resolution (3.4 Å) cryo-EM structure of an active helical nitrilase, the nitrilase 4 from . We apply site-saturation mutagenesis directed evolution to three residues (R95, S224, and L169) and generate a mutant with an altered helical twist that accepts substrates not catalyzed by known plant nitrilases. We reveal that a loop between α2 and α3 limits the length of the binding pocket and propose that it shifts position as a function of helical twist. These insights will allow us to start designing nitrilases for chemoenzymatic synthesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4804.map.gz emd_4804.map.gz | 334.3 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4804-v30.xml emd-4804-v30.xml emd-4804.xml emd-4804.xml | 12.9 KB 12.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4804.png emd_4804.png | 98.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4804 http://ftp.pdbj.org/pub/emdb/structures/EMD-4804 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4804 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4804 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4804.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4804.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AtNIT4 R95T nitrilase filament | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.22 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Arabidopsis thaliana NITRILASE 4 filament
| Entire | Name: Arabidopsis thaliana NITRILASE 4 filament |
|---|---|
| Components |
|
-Supramolecule #1: Arabidopsis thaliana NITRILASE 4 filament
| Supramolecule | Name: Arabidopsis thaliana NITRILASE 4 filament / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Wild-type Arabidopsis thaliana NIT4 modified by site directed mutagenesis at position 95. |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 49 kDa/nm |
-Macromolecule #1: Arabidopsis thaliana Nitrilase 4 R95T
| Macromolecule | Name: Arabidopsis thaliana Nitrilase 4 R95T / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO / EC number: nitrilase |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MSMQQETSHM TAAPQTNGHQ IFPEIDMSAG DSSSIVRATV VQASTVFYDT PATLDKAERL LSEAAENGSQ LVVFPEAFIG GYPRGSTFEL AIGSTTAKGR DDFRKYHASA IDVPGPEVER LALMAKKYKV YLVMGVIERE GYTLYCTVLF FDSQGLFLGK HRKLMPTALE ...String: MSMQQETSHM TAAPQTNGHQ IFPEIDMSAG DSSSIVRATV VQASTVFYDT PATLDKAERL LSEAAENGSQ LVVFPEAFIG GYPRGSTFEL AIGSTTAKGR DDFRKYHASA IDVPGPEVER LALMAKKYKV YLVMGVIERE GYTLYCTVLF FDSQGLFLGK HRKLMPTALE RCIWGFGDGS TIPVFDTPIG KIGAAICWEN RMPSLRTAMY AKGIEIYCAP TADSRETWLA SMTHIALEGG CFVLSANQFC RRKDYPSPPE YMFSGSEESL TPDSVVCAGG SSIISPLGIV LAGPNYRGEA LITADLDLGD IARAKFDFDV VGHYSRPEVF SLNIREHPRK AVSFKTSKVM EDESV |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Staining | Type: NEGATIVE / Material: Uranyl Acetate Details: The protein was allowed to adhere for 30 s, blotted, washed three-times with distilled water and stained with uranyl acetate, blotted again and allowed to dry at room temperature. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 30 / Average exposure time: 1.0 sec. / Average electron dose: 20.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated defocus max: 0.5 µm / Calibrated defocus min: 0.3 µm / Calibrated magnification: 50200 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 1.2 mm / Nominal defocus max: 0.5 µm / Nominal defocus min: 0.3 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder model: PHILIPS ROTATION HOLDER |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 16.0 Å Applied symmetry - Helical parameters - Δ&Phi: -73 ° Applied symmetry - Helical parameters - Axial symmetry: D2 (2x2 fold dihedral) Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 20.0 Å / Resolution method: FSC 1/2 BIT CUT-OFF / Software - Name: SPIDER (ver. 11) / Software - details: IHRSR / Number images used: 3167 |
|---|---|
| Segment selection | Number selected: 1015 / Software - Name: EMAN / Software - details: Boxer / Details: Picked using Eman Boxer in helix mode |
| Startup model | Type of model: INSILICO MODEL In silico model: Featureless cylinder approximating the diameter of the filament. |
| Final angle assignment | Type: NOT APPLICABLE / Software - Name: SPIDER (ver. 11) / Software - details: IHRSR |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















