[English] 日本語
 Yorodumi
Yorodumi- EMDB-14727: 3D reconstruction of the cylindrical assembly of DnaJA2 by imposi... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 3D reconstruction of the cylindrical assembly of DnaJA2 by imposing D5 symmetry | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | oligomer / filaments / helix / Hsp40 / DnaJA2 / holdase / foldase / Hsp70 co-chaperone / CHAPERONE | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
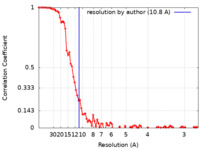
| Method | single particle reconstruction / cryo EM / Resolution: 10.8 Å | |||||||||
 Authors Authors | Cuellar J / Velasco-Carneros L / Santiago C / Martin-Benito J / Valpuesta JM / Muga A | |||||||||
| Funding support |  Spain, 2 items Spain, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: The self-association equilibrium of DNAJA2 regulates its interaction with unfolded substrate proteins and with Hsc70. Authors: Lorea Velasco-Carneros / Jorge Cuéllar / Leire Dublang / César Santiago / Jean-Didier Maréchal / Jaime Martín-Benito / Moisés Maestro / José Ángel Fernández-Higuero / Natalia Orozco ...Authors: Lorea Velasco-Carneros / Jorge Cuéllar / Leire Dublang / César Santiago / Jean-Didier Maréchal / Jaime Martín-Benito / Moisés Maestro / José Ángel Fernández-Higuero / Natalia Orozco / Fernando Moro / José María Valpuesta / Arturo Muga /  Abstract: J-domain proteins tune the specificity of Hsp70s, engaging them in precise functions. Despite their essential role, the structure and function of many J-domain proteins remain largely unknown. We ...J-domain proteins tune the specificity of Hsp70s, engaging them in precise functions. Despite their essential role, the structure and function of many J-domain proteins remain largely unknown. We explore human DNAJA2, finding that it reversibly forms highly-ordered, tubular structures that can be dissociated by Hsc70, the constitutively expressed Hsp70 isoform. Cryoelectron microscopy and mutational studies reveal that different domains are involved in self-association. Oligomer dissociation into dimers potentiates its interaction with unfolded client proteins. The J-domains are accessible to Hsc70 within the tubular structure. They allow binding of closely spaced Hsc70 molecules that could be transferred to the unfolded substrate for its cooperative remodelling, explaining the efficient recovery of DNAJA2-bound clients. The disordered C-terminal domain, comprising the last 52 residues, regulates its holding activity and productive interaction with Hsc70. These in vitro findings suggest that the association equilibrium of DNAJA2 could regulate its interaction with client proteins and Hsc70. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14727.map.gz emd_14727.map.gz | 41.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14727-v30.xml emd-14727-v30.xml emd-14727.xml emd-14727.xml | 17 KB 17 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14727_fsc.xml emd_14727_fsc.xml | 11.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_14727.png emd_14727.png | 31.2 KB | ||
| Filedesc metadata |  emd-14727.cif.gz emd-14727.cif.gz | 5.4 KB | ||
| Others |  emd_14727_half_map_1.map.gz emd_14727_half_map_1.map.gz emd_14727_half_map_2.map.gz emd_14727_half_map_2.map.gz | 41.3 MB 41.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14727 http://ftp.pdbj.org/pub/emdb/structures/EMD-14727 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14727 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14727 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14727.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14727.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.37 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
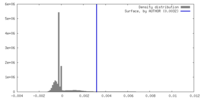
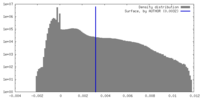
-Half map: #2
| File | emd_14727_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
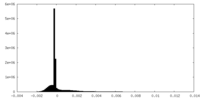
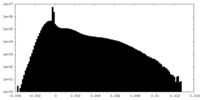
| Density Histograms |
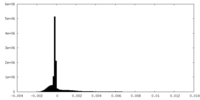
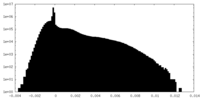
-Half map: #1
| File | emd_14727_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Oligomeric structure of DnaJA2
| Entire | Name: Oligomeric structure of DnaJA2 |
|---|---|
| Components |
|
-Supramolecule #1: Oligomeric structure of DnaJA2
| Supramolecule | Name: Oligomeric structure of DnaJA2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: DnaJA2
| Macromolecule | Name: DnaJA2 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MANVADTKLY DILGVPPGAS ENELKKAYRK LAKEYHPDKN PNAGDKFKEI SFAYEVLSNP EKRELYDRY GEQGLREGSG GGGGMDDIFS HIFGGGLFGF MGNQSRSRNG RRRGEDMMHP L KVSLEDLY NGKTTKLQLS KNVLCSACSG QGGKSGAVQK CSACRGRGVR ...String: MANVADTKLY DILGVPPGAS ENELKKAYRK LAKEYHPDKN PNAGDKFKEI SFAYEVLSNP EKRELYDRY GEQGLREGSG GGGGMDDIFS HIFGGGLFGF MGNQSRSRNG RRRGEDMMHP L KVSLEDLY NGKTTKLQLS KNVLCSACSG QGGKSGAVQK CSACRGRGVR IMIRQLAPGM VQ QMQSVCS DCNGEGEVIN EKDRCKKCEG KKVIKEVKIL EVHVDKGMKH GQRITFTGEA DQA PGVEPG DIVLLLQEKE HEVFQRDGND LHMTYKIGLV EALCGFQFTF KHLDGRQIVV KYPP GKVIE PGCVRVVRGE GMPQYRNPFE KGDLYIKFDV QFPENNWINP DKLSELEDLL PSRPE VPNI IGETEEVELQ EFDSTRGSGG GQRREAYNDS SDEESSSHHG PGVQCAHQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER/RHODIUM / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS TALOS |
|---|---|
| Details | Preliminary grid screening was performed manually |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 1482 / Average exposure time: 1.0 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.2 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)