-Search query
-Search result
Showing 1 - 50 of 13,769 items for (author: au & s)

EMDB-19397: 
Composite map of the C. elegans Intron Lariat Spliceosome primed for disassembly (ILS')

EMDB-19398: 
Structure of the C. elegans Intron Lariat Spliceosome double-primed for disassembly (ILS'')

PDB-8ro0: 
Structure of the C. elegans Intron Lariat Spliceosome primed for disassembly (ILS')

PDB-8ro1: 
Structure of the C. elegans Intron Lariat Spliceosome double-primed for disassembly (ILS'')

EMDB-43128: 
Structure of a membrane transport protein

PDB-8vby: 
Structure of a membrane transport protein

EMDB-19822: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+bromosterol (DOPC, DOPE, DOPS, bromo-ergosterol, PI(4,5)P2 35:20:20:15:10)

EMDB-18661: 
Structure of the Bacteriophage PhiKZ non-virion RNA Polymerase bound to DNA and RNA

PDB-8que: 
Structure of the Bacteriophage PhiKZ non-virion RNA Polymerase bound to DNA and RNA

EMDB-42400: 
RORC mRNA 3'UTR riboswitch A97G/G98A mutant class C

EMDB-42401: 
RORC mRNA 3'UTR riboswitch 77-GA mutant class A

EMDB-42403: 
RORC mRNA 3'UTR riboswitch 117-AC mutant class C

EMDB-42404: 
RORC mRNA 3'UTR riboswitch 117-AC mutant class B

EMDB-41501: 
mGluR3 in the presence of the agonist LY379268 and PAM VU6023326

EMDB-41567: 
Metabotropic glutamate receptor 3 class 3 bound to antagonist LY 341495

EMDB-41568: 
mGluR3 in the presence of the agonist LY379268

EMDB-41577: 
mGluR3 in the presence of the antagonist LY 341495 and positive allosteric modulator VU6023326

EMDB-44861: 
metabotropic glutamate receptor subtype three bound to the antagonist LY 341495, class two

PDB-8tqb: 
mGluR3 in the presence of the agonist LY379268 and PAM VU6023326

PDB-8tr0: 
Metabotropic glutamate receptor 3 class 3 bound to antagonist LY 341495

PDB-8tr2: 
mGluR3 in the presence of the agonist LY379268

PDB-8trc: 
mGluR3 in the presence of the antagonist LY 341495 and positive allosteric modulator VU6023326

EMDB-15127: 
Structure of mammalian Pol II-DSIF-SPT6-PAF1-TFIIS-hexasome elongation complex

PDB-8a3y: 
Structure of mammalian Pol II-DSIF-SPT6-PAF1-TFIIS-hexasome elongation complex

EMDB-43139: 
SARS-CoV-2 Spike S2 bound to Fab 54043-5

EMDB-41578: 
mGluR3 class 1 in the presence of the antagonist LY 341495

EMDB-45242: 
mGluR3 in the presence of the antagonist LY 341495 and positive allosteric modulator VU6023326 class 2

PDB-8trd: 
mGluR3 class 1 in the presence of the antagonist LY 341495
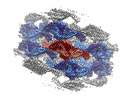

EMDB-18307: 
Native eisosome lattice bound to plasma membrane microdomain

EMDB-18308: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture -PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol 30:20:20:30)

EMDB-18309: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/-sterol (DOPC, DOPE, DOPS, PI(4,5)P2 50:20:20:10)

EMDB-18310: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol, PI(4,5)P2 35:20:20:15:10)

EMDB-18311: 
Compact state - Native eisosome lattice bound to plasma membrane microdomain

EMDB-18312: 
Stretched state - Native eisosome lattice bound to plasma membrane microdomain

PDB-8qb7: 
Pil1 in native eisosome lattice bound to plasma membrane microdomain

PDB-8qb8: 
Lsp1 in native eisosome lattice bound to plasma membrane microdomain

PDB-8qb9: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture -PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol 30:20:20:30)

PDB-8qbb: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/-sterol (DOPC, DOPE, DOPS, PI(4,5)P2 50:20:20:10)

PDB-8qbd: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol, PI(4,5)P2 35:20:20:15:10)

PDB-8qbe: 
Compact state - Pil1 in native eisosome lattice bound to plasma membrane microdomain

PDB-8qbf: 
Compact state - Pil1 dimer with lipid headgroups fitted in native eisosome lattice bound to plasma membrane microdomain

PDB-8qbg: 
Stretched state - Pil1 in native eisosome lattice bound to plasma membrane microdomain
EMDB-19978: 
Outward-open structure of Drosophila dopamine transporter bound to an atypical non-competitive inhibitor
EMDB-19979: 
Inhibitor-free outward-open structure of Drosophila dopamine transporter

PDB-9euo: 
Outward-open structure of Drosophila dopamine transporter bound to an atypical non-competitive inhibitor

PDB-9eup: 
Inhibitor-free outward-open structure of Drosophila dopamine transporter

EMDB-18779: 
Structure of the non-mitochondrial citrate synthase from Ananas comosus

PDB-8qzp: 
Structure of the non-mitochondrial citrate synthase from Ananas comosus

EMDB-43551: 
CCHFV GP38 bound with ADI-46143 and ADI-46158 Fabs

EMDB-43552: 
CCHFV GP38 bound with ADI-58062 and ADI-63530 Fabs
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
