9MM2
 
 | | Crystal structure of bacterial pectin methylesterase PmeC5 from B. fibrisolvens D1T | | Descriptor: | 1,2-ETHANEDIOL, AMMONIUM ION, MAGNESIUM ION, ... | | Authors: | Carbone, V, Reilly, K, Sang, C, Schofield, L, Ronimus, R, Attwood, G.T, Palevich, N. | | Deposit date: | 2024-12-19 | | Release date: | 2025-01-15 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | Crystal Structure of the Multidomain Pectin Methylesterase PmeC5 from Butyrivibrio fibrisolvens D1 T.
Biomolecules, 15, 2025
|
|
4E7C
 
 | |
8HDN
 
 | |
8QPE
 
 | | Cryo-EM Structure of Pre-B-like Complex (core part) | | Descriptor: | 116 kDa U5 small nuclear ribonucleoprotein component, INOSITOL HEXAKISPHOSPHATE, Microfibrillar-associated protein 1, ... | | Authors: | Zhang, Z, Kumar, V, Dybkov, O, Will, C.L, Zhong, J, Ludwig, S, Urlaub, H, Kastner, B, Stark, H, Luehrmann, R. | | Deposit date: | 2023-10-01 | | Release date: | 2024-05-22 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structural insights into the cross-exon to cross-intron spliceosome switch.
Nature, 630, 2024
|
|
6NLV
 
 | | Selective inhibition of carbonic anhydrase IX activity, using compound SLC-149, displays limited anticancer effects in breast cancer cell lines | | Descriptor: | 4-[3-(2,4-difluorophenyl)-2-oxo-2,3-dihydro-1H-imidazol-1-yl]benzene-1-sulfonamide, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Singh, S, McKenna, R. | | Deposit date: | 2019-01-09 | | Release date: | 2020-01-15 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.794 Å) | | Cite: | Inhibition of Carbonic Anhydrase Using SLC-149: Support for a Noncatalytic Function of CAIX in Breast Cancer.
J.Med.Chem., 64, 2021
|
|
9CX9
 
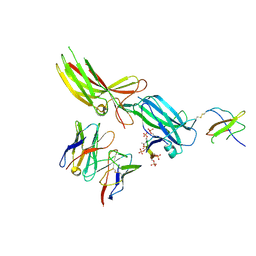 | | Structure of SH3 domain of Src in complex with beta-arrestin 1 | | Descriptor: | Antibody fragment Fab30, heavy chain, light chain, ... | | Authors: | Pakharukova, N, Bansia, H, Lefkowitz, R.J, des Georges, A. | | Deposit date: | 2024-07-31 | | Release date: | 2024-11-13 | | Last modified: | 2024-11-27 | | Method: | ELECTRON MICROSCOPY (3.34 Å) | | Cite: | Beta-arrestin 1 mediated Src activation via Src SH3 domain revealed by cryo-electron microscopy.
Biorxiv, 2024
|
|
7O3E
 
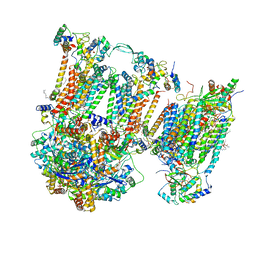 | | Murine supercomplex CIII2CIV in the intermediate locked conformation | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 1,2-Distearoyl-sn-glycerophosphoethanolamine, CARDIOLIPIN, ... | | Authors: | Vercellino, I, Sazanov, L.A. | | Deposit date: | 2021-04-01 | | Release date: | 2021-10-13 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structure and assembly of the mammalian mitochondrial supercomplex CIII 2 CIV.
Nature, 598, 2021
|
|
7O3C
 
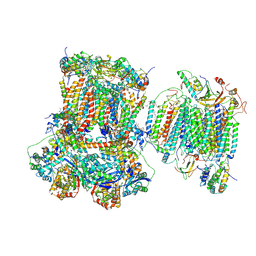 | | Murine supercomplex CIII2CIV in the mature unlocked conformation | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 1,2-Distearoyl-sn-glycerophosphoethanolamine, CARDIOLIPIN, ... | | Authors: | Vercellino, I, Sazanov, L.A. | | Deposit date: | 2021-04-01 | | Release date: | 2021-10-13 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Structure and assembly of the mammalian mitochondrial supercomplex CIII 2 CIV.
Nature, 598, 2021
|
|
7O37
 
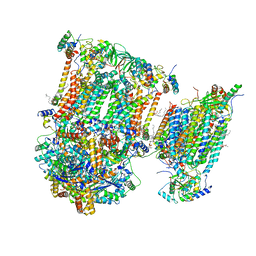 | | Murine supercomplex CIII2CIV in the assembled locked conformation | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 1,2-Distearoyl-sn-glycerophosphoethanolamine, CARDIOLIPIN, ... | | Authors: | Vercellino, I, Sazanov, L.A. | | Deposit date: | 2021-04-01 | | Release date: | 2021-10-13 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structure and assembly of the mammalian mitochondrial supercomplex CIII 2 CIV.
Nature, 598, 2021
|
|
7W61
 
 | | Crystal structure of farnesol dehydrogenase from Helicoverpa armigera | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Farnesol dehydrogenase, ... | | Authors: | Kumar, R, Das, J, Mahto, J.K, Sharma, M, Kumar, P, Sharma, A.K. | | Deposit date: | 2021-12-01 | | Release date: | 2022-07-27 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure and molecular characterization of NADP + -farnesol dehydrogenase from cotton bollworm, Helicoverpaarmigera.
Insect Biochem.Mol.Biol., 147, 2022
|
|
6ZWI
 
 | | Human butyrylcholinesterase in complex with ((6-((2E,4E)-5-(benzo[d][1,3]dioxol-5-yl)penta-2,4-dienamido)hexyl)triphenylphosphonium bromide) | | Descriptor: | (2~{E},4~{E})-5-(1,3-benzodioxol-5-yl)-~{N}-[6-(triphenyl-$l^{5}-phosphanyl)hexyl]penta-2,4-dienamide, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Da Silva, O, Nachon, F, Dias, J, Brazzolotto, X. | | Deposit date: | 2020-07-28 | | Release date: | 2021-06-09 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Fine-Tuning the Biological Profile of Multitarget Mitochondriotropic Antioxidants for Neurodegenerative Diseases.
Antioxidants (Basel), 10, 2021
|
|
6NM0
 
 | | Selective inhibition of carbonic anhydrase IX activity, using compound SLC-149, displays limited anticancer effects in breast cancer cell lines | | Descriptor: | 4-[3-(2,4-difluorophenyl)-2-oxo-2,3-dihydro-1H-imidazol-1-yl]benzene-1-sulfonamide, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Singh, S, McKenna, R. | | Deposit date: | 2019-01-10 | | Release date: | 2020-01-15 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | Inhibition of Carbonic Anhydrase Using SLC-149: Support for a Noncatalytic Function of CAIX in Breast Cancer.
J.Med.Chem., 64, 2021
|
|
4MWS
 
 | |
3U2U
 
 | | Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and maltotetraose | | Descriptor: | GLYCEROL, Glycogenin-1, MANGANESE (II) ION, ... | | Authors: | Chaikuad, A, Froese, D.S, Krysztofinska, E, von Delft, F, Weigelt, J, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Oppermann, U, Yue, W.W, Structural Genomics Consortium (SGC) | | Deposit date: | 2011-10-04 | | Release date: | 2011-11-02 | | Last modified: | 2024-11-27 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Conformational plasticity of glycogenin and its maltosaccharide substrate during glycogen biogenesis.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
8R15
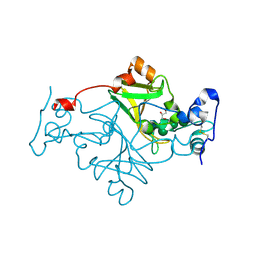 
 | | Crystal structure of Fusarium oxysporum NADase I | | Descriptor: | GLYCEROL, TNT domain-containing protein, alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Kallio, J.P, Ferrario, E, Stromland, O, Ziegler, M. | | Deposit date: | 2023-11-01 | | Release date: | 2024-07-03 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Evolution of fungal tuberculosis necrotizing toxin (TNT) domain-containing enzymes reveals divergent adaptations to enhance NAD cleavage.
Protein Sci., 33, 2024
|
|
2N29
 
 | | Solution-state NMR structure of Vpu cytoplasmic domain | | Descriptor: | Protein Vpu | | Authors: | Zhang, H, Lin, E.C, Tian, Y, Das, B.B, Opella, S.J. | | Deposit date: | 2015-05-01 | | Release date: | 2015-09-30 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural determination of virus protein U from HIV-1 by NMR in membrane environments.
Biochim.Biophys.Acta, 1848, 2015
|
|
4Y4T
 
 | | Endothiapepsin in complex with fragment 114 | | Descriptor: | 1,2-ETHANEDIOL, 3-amino-5-(pyrrolidin-1-yl)-1H-pyrazole-4-carbonitrile, ACETATE ION, ... | | Authors: | Schiebel, J, Heine, A, Klebe, G. | | Deposit date: | 2015-02-11 | | Release date: | 2016-03-02 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.301 Å) | | Cite: | Crystallographic Fragment Screening of an Entire Library
To Be Published
|
|
6NFZ
 
 | | Crystal structure of diphosphorylated HPK1 kinase domain in complex with sunitinib in the active state. | | Descriptor: | Mitogen-activated protein kinase kinase kinase kinase 1, N-[2-(diethylamino)ethyl]-5-[(Z)-(5-fluoro-2-oxo-1,2-dihydro-3H-indol-3-ylidene)methyl]-2,4-dimethyl-1H-pyrrole-3-carbo xamide | | Authors: | Johnson, E, McTigue, M, Cronin, C.N. | | Deposit date: | 2018-12-21 | | Release date: | 2019-05-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.966 Å) | | Cite: | Multiple conformational states of the HPK1 kinase domain in complex with sunitinib reveal the structural changes accompanying HPK1 trans-regulation.
J.Biol.Chem., 294, 2019
|
|
9LGL
 
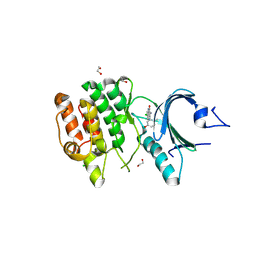 | | Crystal structure of human PKMYT1 protein kinase domain with Naphthyridinone Inhibitor compound 6 | | Descriptor: | 1,2-ETHANEDIOL, 3-azanyl-7-chloranyl-4-(2-methyl-5-oxidanyl-phenyl)-1H-1,5-naphthyridin-2-one, CHLORIDE ION, ... | | Authors: | Xu, Z.H, Chen, S. | | Deposit date: | 2025-01-10 | | Release date: | 2025-05-14 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Discovery of Naphthyridinone Derivatives as Selective and Potent PKMYT1 Inhibitors with Antitumor Efficacy.
J.Med.Chem., 68, 2025
|
|
9KP9
 
 | | PfDXR - Mn2+ - NADPH - TAKK443 quaternary complex | | Descriptor: | 1-deoxy-D-xylulose 5-phosphate reductoisomerase, apicoplastic, MANGANESE (II) ION, ... | | Authors: | Takada, S, Sakamoto, Y, Tanaka, N. | | Deposit date: | 2024-11-22 | | Release date: | 2025-05-21 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | Expanding the Chemical Space of Reverse Fosmidomycin Analogs.
Acs Med.Chem.Lett., 16, 2025
|
|
4EWT
 
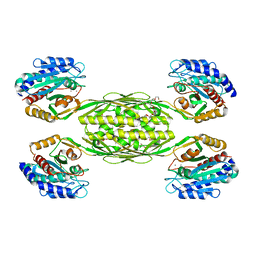 | | The crystal structure of a putative aminohydrolase from methicillin resistant Staphylococcus aureus | | Descriptor: | 1-DEOXY-1-THIO-HEPTAETHYLENE GLYCOL, DI(HYDROXYETHYL)ETHER, MANGANESE (II) ION, ... | | Authors: | Girish, T.S, Vivek, B, Colaco, M, Misquith, S, Gopal, B. | | Deposit date: | 2012-04-27 | | Release date: | 2013-02-20 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of an amidohydrolase, SACOL0085, from methicillin-resistant Staphylococcus aureus COL
Acta Crystallogr.,Sect.F, 69, 2013
|
|
8I1Z
 
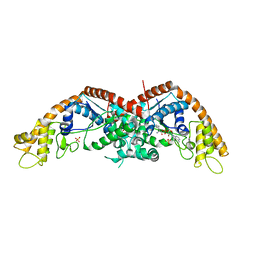 | | E. coli tryptophanyl-tRNA synthetase bound with a chemical fragment | | Descriptor: | 1-(2,3-dihydro-1-benzofuran-5-yl)ethanone, SULFATE ION, TRYPTOPHANYL-5'AMP, ... | | Authors: | Xiang, M, Zhou, H. | | Deposit date: | 2023-01-13 | | Release date: | 2023-04-12 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | An asymmetric structure of bacterial TrpRS supports the half-of-the-sites catalytic mechanism and facilitates antimicrobial screening.
Nucleic Acids Res., 51, 2023
|
|
7O8C
 
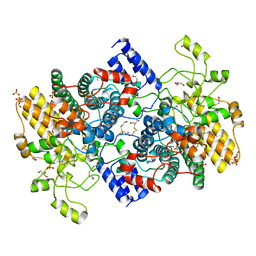 | | Structure of SGBP BO2743 from Bacteroides ovatus | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, AZIDE ION, BETA-MERCAPTOETHANOL, ... | | Authors: | Correia, V.C, Trovao, F, Pinheiro, B.A, Palma, A.S, Carvalho, A.L. | | Deposit date: | 2021-04-15 | | Release date: | 2021-12-08 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Mapping Molecular Recognition of beta 1,3-1,4-Glucans by a Surface Glycan-Binding Protein from the Human Gut Symbiont Bacteroides ovatus.
Microbiol Spectr, 9, 2021
|
|
4EUP
 
 | |
2MPM
 
 | | Structural Basis of Receptor Sulfotyrosine Recognition by a CC Chemokine: the N-terminal Region of CCR3 Bound to CCL11/Eotaxin-1 | | Descriptor: | CCR3, Eotaxin | | Authors: | Millard, C.J, Ludeman, J.P, Canals, M, Bridgford, J.L, Hinds, M.G, Clayton, D.J, Christopoulos, A, Payne, R.J, Stone, M.J. | | Deposit date: | 2014-05-26 | | Release date: | 2014-12-10 | | Last modified: | 2024-11-06 | | Method: | SOLUTION NMR | | Cite: | Structural Basis of Receptor Sulfotyrosine Recognition by a CC Chemokine: The N-Terminal Region of CCR3 Bound to CCL11/Eotaxin-1.
Structure, 22, 2014
|
|
