2PQW
 
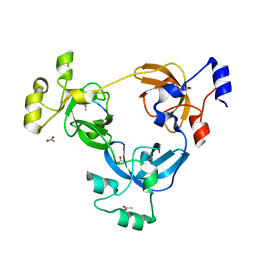 | | Crystal structure of L3MBTL1 in complex with H4K20Me2 (residues 17-25), trigonal form | | Descriptor: | ACETATE ION, Histone H4, Lethal(3)malignant brain tumor-like protein | | Authors: | Allali-Hassani, A, Liu, Y, Herzanych, N, Ouyang, H, Mackenzie, F, Crombet, L, Loppnau, P, Kozieradzki, I, Vedadi, M, Weigelt, J, Sundstrom, M, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Min, J.R, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-05-02 | | Release date: | 2007-07-31 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | L3MBTL1 recognition of mono- and dimethylated histones.
Nat.Struct.Mol.Biol., 14, 2007
|
|
2PQX
 
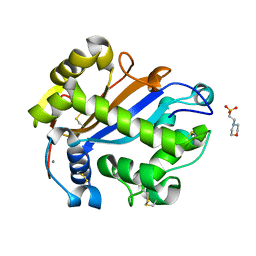 | | E. coli RNase 1 (in vivo folded) | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CALCIUM ION, Ribonuclease I | | Authors: | Messens, J, Loris, R, Wyns, L. | | Deposit date: | 2007-05-03 | | Release date: | 2007-09-18 | | Last modified: | 2017-10-18 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Crystal structure of E. coli ribonuclease 1
To be Published
|
|
2PQY
 
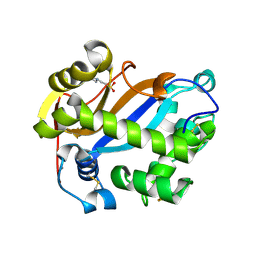 | | E. coli RNase 1 (in vitro refolded with DsbA only) | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CALCIUM ION, Ribonuclease I | | Authors: | Messens, J, Loris, R. | | Deposit date: | 2007-05-03 | | Release date: | 2007-08-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The oxidase DsbA folds a protein with a nonconsecutive disulfide.
J.Biol.Chem., 282, 2007
|
|
2PQZ
 
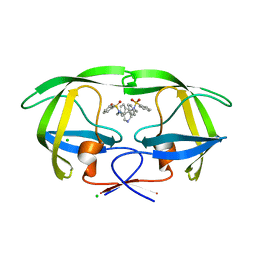 | | HIV-1 Protease in complex with a pyrrolidine-based inhibitor | | Descriptor: | CHLORIDE ION, N,N'-(3S,4S)-PYRROLIDINE-3,4-DIYLBIS(N-BENZYLBENZENESULFONAMIDE), Protease | | Authors: | Boettcher, J, Blum, A, Heine, A, Diederich, W.E, Klebe, G. | | Deposit date: | 2007-05-03 | | Release date: | 2008-04-15 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure-Guided Design of C2-Symmetric HIV-1 Protease Inhibitors Based on a Pyrrolidine Scaffold.
J.Med.Chem., 51, 2008
|
|
2PR0
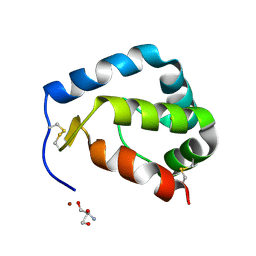 
 | | Crystal structure of Sylvaticin, a new secreted protein from Pythium Sylvaticum | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, NICKEL (II) ION, sylvaticin | | Authors: | Lascombe, M.B, Prange, T, Retailleau, P. | | Deposit date: | 2007-05-03 | | Release date: | 2008-03-18 | | Last modified: | 2017-10-18 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Structure of sylvaticin, a new alpha-elicitin-like protein from Pythium sylvaticum.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2PR1
 
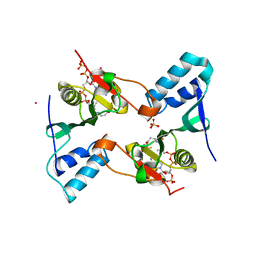 | | Crystal structure of the Bacillus subtilis N-acetyltransferase YlbP protein in complex with Coenzyme-A | | Descriptor: | COBALT (II) ION, COENZYME A, SULFATE ION, ... | | Authors: | Minasov, G, Shuvalova, L, Kiryukhina, O, Vorontsov, I.I, Collart, F.R, Joachimiak, A, Anderson, W.F, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2007-05-03 | | Release date: | 2007-05-15 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal structure of the Bacillus subtilis N-acetyltransferase YlbP protein in complex with Coenzyme-A.
To be Published
|
|
2PR2
 
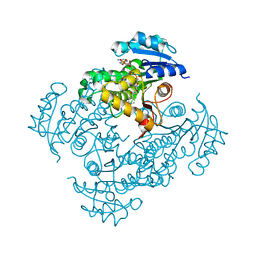 | |
2PR3
 
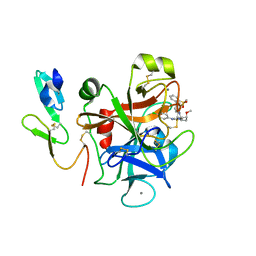 | | Factor XA inhibitor | | Descriptor: | (2R,4R)-N~1~-(4-CHLOROPHENYL)-N~2~-[3-FLUORO-2'-(METHYLSULFONYL)BIPHENYL-4-YL]-4-METHOXYPYRROLIDINE-1,2-DICARBOXAMIDE, CALCIUM ION, COAGULATION FACTOR X, ... | | Authors: | Zhang, E, Kohrt, J.T, Bigge, C.F, Finzel, B.C. | | Deposit date: | 2007-05-03 | | Release date: | 2007-08-14 | | Last modified: | 2017-10-18 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structure-based drug design of pyrrolidine-1, 2-dicarboxamides as a novel series of orally bioavailable factor Xa inhibitors
Chem.Biol.Drug Des., 69, 2007
|
|
2PR4
 
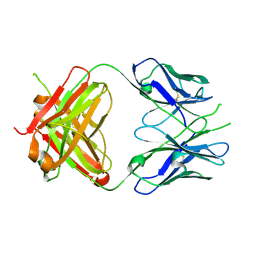 | | Crystal Structure of Fab' from the HIV-1 Neutralizing Antibody 2F5 | | Descriptor: | nmAb 2F5 Fab' Heavy Chain, nmAb 2F5 Fab' light Chain | | Authors: | Bryson, S, Julien, J.-P, Pai, E.F. | | Deposit date: | 2007-05-03 | | Release date: | 2007-05-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural details of HIV-1 recognition by the broadly neutralizing monoclonal antibody 2F5: epitope conformation, antigen-recognition loop mobility, and anion-binding site.
J.Mol.Biol., 384, 2008
|
|
2PR5
 
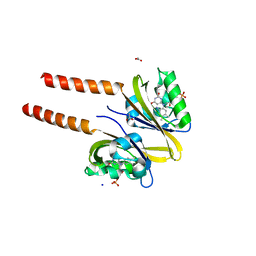 | |
2PR6
 
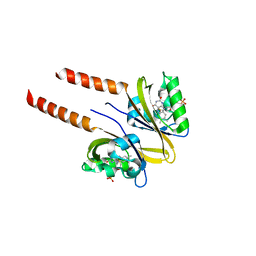 | |
2PR7
 
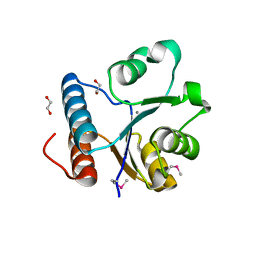 | |
2PR8
 
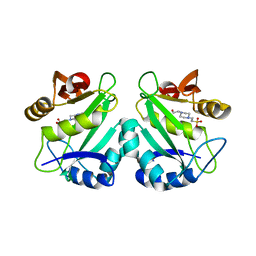 | | crystal structure of aminoglycoside N-acetyltransferase AAC(6')-Ib11 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Aminoglycoside 6-N-acetyltransferase type Ib11 | | Authors: | Maurice, F, Broutin, I, Podglajen, I, Benas, P, Collatz, E, Dardel, F. | | Deposit date: | 2007-05-04 | | Release date: | 2008-04-08 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Enzyme structural plasticity and the emergence of broad-spectrum antibiotic resistance.
Embo Rep., 9, 2008
|
|
2PR9
 
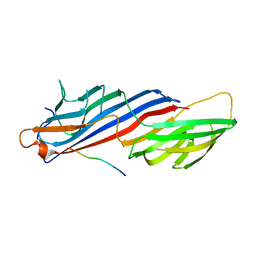 | | Mu2 adaptin subunit (AP50) of AP2 adaptor (second domain), complexed with GABAA receptor-gamma2 subunit-derived internalization peptide DEEYGYECL | | Descriptor: | AP-2 complex subunit mu-1, GABA(A) receptor subunit gamma-2 peptide | | Authors: | Vahedi-Faridi, A, Haucke, V, Kittler, J.T, Kukhtina, V, Moss, S.J, Saenger, W, Chen, G.-J, Tretter, V, Smith, K, Yan, Z, McAinsh, K, Arancibia-Carcamo, L. | | Deposit date: | 2007-05-04 | | Release date: | 2008-03-18 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Regulation of synaptic inhibition by phospho-dependent binding of the AP2 complex to a YECL motif in the GABAA receptor gamma2 subunit.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
2PRB
 
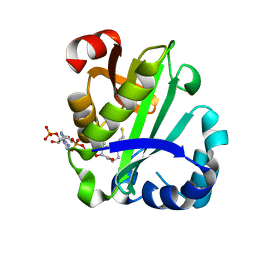 | | crystal structure of aminoglycoside acetyltransferase AAC(6')-Ib in complex whith coenzyme A | | Descriptor: | Aminoglycoside 6-N-acetyltransferase type Ib11, COENZYME A | | Authors: | Maurice, F, Broutin, I, Podglajen, I, Benas, P, Collatz, E, Dardel, F. | | Deposit date: | 2007-05-04 | | Release date: | 2008-04-08 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Enzyme structural plasticity and the emergence of broad-spectrum antibiotic resistance.
Embo Rep., 9, 2008
|
|
2PRC
 
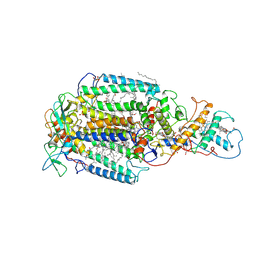 | | PHOTOSYNTHETIC REACTION CENTER FROM RHODOPSEUDOMONAS VIRIDIS (UBIQUINONE-2 COMPLEX) | | Descriptor: | 15-cis-1,2-dihydroneurosporene, BACTERIOCHLOROPHYLL B, BACTERIOPHEOPHYTIN B, ... | | Authors: | Lancaster, C.R.D, Michel, H. | | Deposit date: | 1997-07-29 | | Release date: | 1998-11-11 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | The coupling of light-induced electron transfer and proton uptake as derived from crystal structures of reaction centres from Rhodopseudomonas viridis modified at the binding site of the secondary quinone, QB.
Structure, 5, 1997
|
|
2PRD
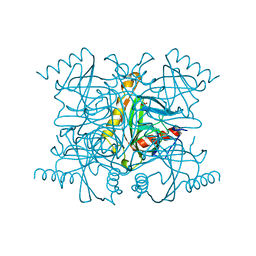 
 | |
2PRE
 
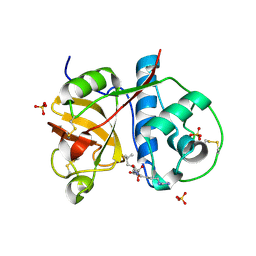 | | Crystal structure of plant cysteine protease Ervatamin-C complexed with irreversible inhibitor E-64 at 2.7 A resolution | | Descriptor: | Ervatamin-C, N-[N-[1-HYDROXYCARBOXYETHYL-CARBONYL]LEUCYLAMINO-BUTYL]-GUANIDINE, SULFATE ION | | Authors: | Ghosh, R, Chakrabarti, C, Dattagupta, J.K, Biswas, S. | | Deposit date: | 2007-05-04 | | Release date: | 2008-01-22 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural insights into the substrate specificity and activity of ervatamins, the papain-like cysteine proteases from a tropical plant, Ervatamia coronaria.
Febs J., 275, 2008
|
|
2PRF
 
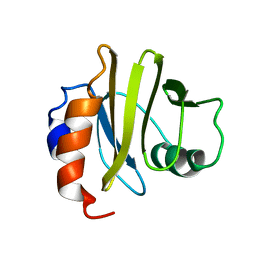 | | THREE DIMENSIONAL SOLUTION STRUCTURE OF ACANTHAMOEBA PROFILIN I | | Descriptor: | PROFILIN IA | | Authors: | Archer, S.J, Vinson, V.K, Pollard, T.D, Lattman, E.E, Torchia, D.A. | | Deposit date: | 1994-01-12 | | Release date: | 1994-05-31 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional solution structure of Acanthamoeba profilin-I.
J.Cell Biol., 122, 1993
|
|
2PRG
 
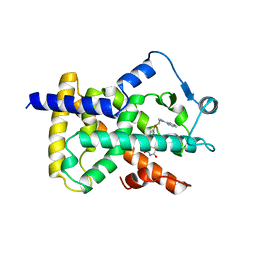 | | LIGAND-BINDING DOMAIN OF THE HUMAN PEROXISOME PROLIFERATOR ACTIVATED RECEPTOR GAMMA | | Descriptor: | 2,4-THIAZOLIDIINEDIONE, 5-[[4-[2-(METHYL-2-PYRIDINYLAMINO)ETHOXY]PHENYL]METHYL]-(9CL), NUCLEAR RECEPTOR COACTIVATOR SRC-1, ... | | Authors: | Nolte, R.T, Wisely, G.B, Milburn, M.V. | | Deposit date: | 1998-08-14 | | Release date: | 1999-07-19 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Ligand binding and co-activator assembly of the peroxisome proliferator-activated receptor-gamma.
Nature, 395, 1998
|
|
2PRH
 
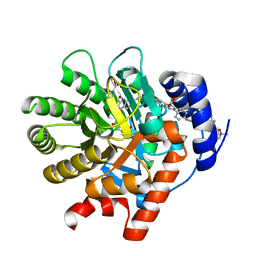 | | The structures of apo- and inhibitor bound human dihydroorotate dehydrogenase reveal conformational flexibility within the inhibitor binding site | | Descriptor: | 6-CHLORO-2-(2'-FLUOROBIPHENYL-4-YL)-3-METHYLQUINOLINE-4-CARBOXYLIC ACID, DECYLAMINE-N,N-DIMETHYL-N-OXIDE, Dihydroorotate dehydrogenase, ... | | Authors: | Walse, B, Dufe, V.T, Al-Karadaghi, S. | | Deposit date: | 2007-05-04 | | Release date: | 2008-05-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The structures of human dihydroorotate dehydrogenase with and without inhibitor reveal conformational flexibility in the inhibitor and substrate binding sites
Biochemistry, 47, 2008
|
|
2PRI
 
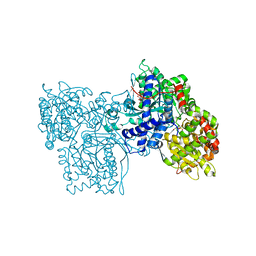 | | BINDING OF 2-DEOXY-GLUCOSE-6-PHOSPHATE TO GLYCOGEN PHOSPHORYLASE B | | Descriptor: | 2-deoxy-6-O-phosphono-alpha-D-glucopyranose, GLYCOGEN PHOSPHORYLASE B, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Oikonomakos, N.G, Zographos, S.E, Johnson, L.N, Papageorgiou, A.C, Acharya, K.R. | | Deposit date: | 1998-12-11 | | Release date: | 1998-12-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The binding of 2-deoxy-D-glucose 6-phosphate to glycogen phosphorylase b: kinetic and crystallographic studies.
J.Mol.Biol., 254, 1995
|
|
2PRJ
 
 | | Binding of N-acetyl-beta-D-glucopyranosylamine to Glycogen Phosphorylase B | | Descriptor: | GLYCOGEN PHOSPHORYLASE, INOSINIC ACID, N-acetyl-beta-D-glucopyranosylamine, ... | | Authors: | Oikonomakos, N.G, Kontou, M, Zographos, S.E, Watson, K.A, Johnson, L.N, Bichard, C.J.F, Fleet, G.W.J, Acharya, K.R. | | Deposit date: | 1998-12-11 | | Release date: | 1998-12-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | N-acetyl-beta-D-glucopyranosylamine: a potent T-state inhibitor of glycogen phosphorylase. A comparison with alpha-D-glucose.
Protein Sci., 4, 1995
|
|
2PRK
 
 | |
2PRL
 
 | | The structures of apo- and inhibitor bound human dihydroorotate dehydrogenase reveal conformational flexibility within the inhibitor binding site | | Descriptor: | 5-METHOXY-2-[(4-PHENOXYPHENYL)AMINO]BENZOIC ACID, ACETATE ION, DECYLAMINE-N,N-DIMETHYL-N-OXIDE, ... | | Authors: | Walse, B, Dufe, V.T, Al-Karadaghi, S. | | Deposit date: | 2007-05-04 | | Release date: | 2008-05-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The structures of human dihydroorotate dehydrogenase with and without inhibitor reveal conformational flexibility in the inhibitor and substrate binding sites
Biochemistry, 47, 2008
|
|
