3COD
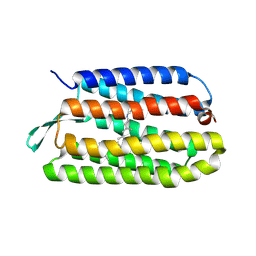 
 | | Crystal Structure of T90A/D115A mutant of Bacteriorhodopsin | | Descriptor: | Bacteriorhodopsin, RETINAL | | Authors: | Joh, N.H, Min, A, Faham, S, Bowie, J.U. | | Deposit date: | 2008-03-27 | | Release date: | 2008-04-08 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Modest stabilization by most hydrogen-bonded side-chain interactions in membrane proteins.
Nature, 453, 2008
|
|
3CPT
 
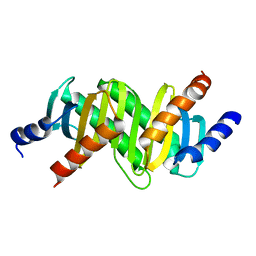 | | MP1-p14 Scaffolding complex | | Descriptor: | Mitogen-activated protein kinase kinase 1-interacting protein 1, Mitogen-activated protein-binding protein-interacting protein | | Authors: | Schrag, J.D, Cygler, M, Munger, C, Magloire, A. | | Deposit date: | 2008-04-01 | | Release date: | 2008-07-01 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular dynamics-solvated interaction energy studies of protein-protein interactions: the MP1-p14 scaffolding complex.
J.Mol.Biol., 379, 2008
|
|
3CWY
 
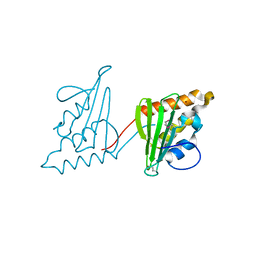 | | Structure of CagD from H. pylori pathogenicity island crystallized in the presence of Cu(II) ions | | Descriptor: | COPPER (II) ION, protein CagD | | Authors: | Cendron, L, Zanotti, G, Angelini, A, Barison, N, Couturier, M, Stein, M. | | Deposit date: | 2008-04-23 | | Release date: | 2008-12-30 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | The Helicobacter pylori CagD (HP0545, Cag24) protein is essential for CagA translocation and maximal induction of interleukin-8 secretion.
J.Mol.Biol., 386, 2009
|
|
8I0C
 
 | | Crystal structure of Aldo-keto reductase 1C3 complexed with compound S0703 | | Descriptor: | 1-[4-[3,5-bis(chloranyl)phenyl]-3-fluoranyl-phenyl]cyclopropane-1-carboxylic acid, Aldo-keto reductase family 1 member C3, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Jiang, J, He, S, Liu, Y, Fang, P, Sun, H. | | Deposit date: | 2023-01-10 | | Release date: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Development of Biaryl-Containing Aldo-Keto Reductase 1C3 (AKR1C3) Inhibitors for Reversing AKR1C3-Mediated Drug Resistance in Cancer Treatment.
J.Med.Chem., 66, 2023
|
|
3D2B
 
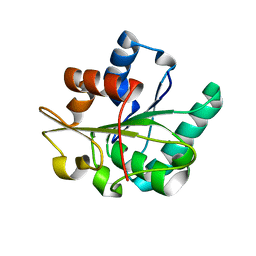 | |
3D5V
 
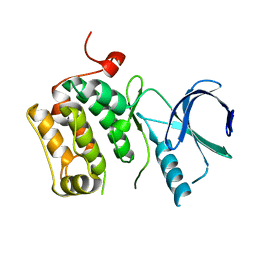 | |
2J6R
 
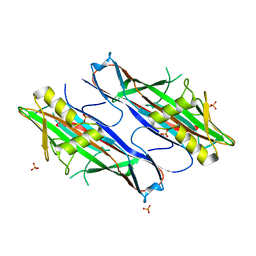 | | FaeG from F4ac ETEC strain GIS26, produced in tobacco plant chloroplast | | Descriptor: | K88 FIMBRIAL PROTEIN, PHOSPHATE ION | | Authors: | Van Molle, I, Joensuu, J.J, Buts, L, Panjikar, S, Kotiaho, M, Bouckaert, J, Wyns, L, Niklander-Teeri, V, De Greve, H. | | Deposit date: | 2006-10-03 | | Release date: | 2007-04-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Chloroplasts Assemble the Major Subunit Faeg of Escherichia Coli F4 (K88) Fimbriae Into Strand-Swapped Dimers
J.Mol.Biol., 368, 2007
|
|
7ZPT
 
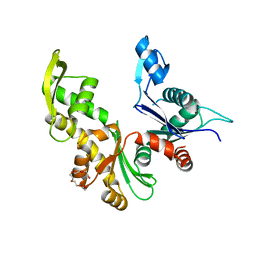 | |
7ZPU
 
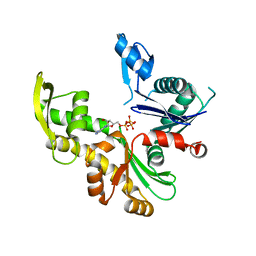 | |
2JH5
 
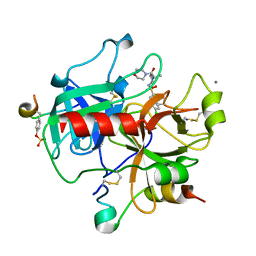 | | Human Thrombin Hirugen Inhibitor complex | | Descriptor: | 2-(5-CHLORO-2-THIENYL)-N-{(3S)-1-[(1S)-1-METHYL-2-MORPHOLIN-4-YL-2-OXOETHYL]-2-OXOPYRROLIDIN-3-YL}ETHENESULFONAMIDE, CALCIUM ION, HIRUDIN IIIA, ... | | Authors: | Senger, S, Chan, C, Convery, M.A, Hubbard, J.A, Young, R.J, Shah, G.P, Watson, N.S. | | Deposit date: | 2007-02-20 | | Release date: | 2007-05-08 | | Last modified: | 2017-06-28 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Sulfonamide-Related Conformational Effects and Their Importance in Structure-Based Design.
Bioorg.Med.Chem.Lett., 17, 2007
|
|
2JH0
 
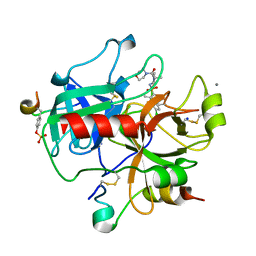 | | Human Thrombin Hirugen Inhibitor complex | | Descriptor: | (2R)-2-(5-CHLORO-2-THIENYL)-N-{(3S)-1-[(1S)-1-METHYL-2-MORPHOLIN-4-YL-2-OXOETHYL]-2-OXOPYRROLIDIN-3-YL}PROPENE-1-SULFONAMIDE, CALCIUM ION, HIRUDIN IIIA, ... | | Authors: | Senger, S, Chan, C, Convery, M.A, Hubbard, J.A, Young, R.J, Shah, G.P, Watson, N.S. | | Deposit date: | 2007-02-19 | | Release date: | 2007-05-08 | | Last modified: | 2013-08-07 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Sulfonamide-Related Conformational Effects and Their Importance in Structure-Based Design.
Bioorg.Med.Chem.Lett., 17, 2007
|
|
3DAT
 
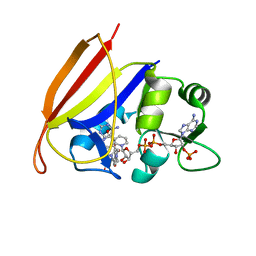 | |
2J5A
 
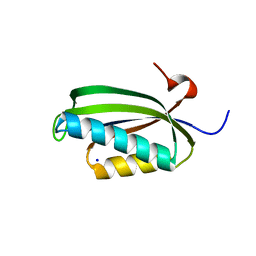 | | Folding of S6 structures with divergent amino-acid composition: pathway flexibility within partly overlapping foldons | | Descriptor: | 30S RIBOSOMAL PROTEIN S6, SODIUM ION | | Authors: | Hansson, S, Olofsson, L, Hedberg, L, Oliveberg, M, Logan, D.T. | | Deposit date: | 2006-09-13 | | Release date: | 2006-10-25 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Folding of S6 Structures with Divergent Amino Acid Composition: Pathway Flexibility within Partly Overlapping Foldons.
J.Mol.Biol., 365, 2007
|
|
2IOF
 
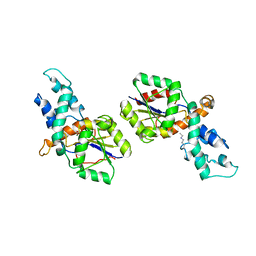 | | Crystal structure of phosphonoacetaldehyde hydrolase with sodium borohydride-reduced substrate intermediate | | Descriptor: | MAGNESIUM ION, PHOSPHATE ION, Phosphonoacetaldehyde hydrolase | | Authors: | Allen, K.A, Lahiri, S.D, Zhang, G, Dunaway-Mariano, D. | | Deposit date: | 2006-10-10 | | Release date: | 2007-07-17 | | Last modified: | 2019-07-24 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Diversification of function in the haloacid dehalogenase enzyme superfamily: The role of the cap domain in hydrolytic phosphoruscarbon bond cleavage.
Bioorg.Chem., 34, 2006
|
|
2JG7
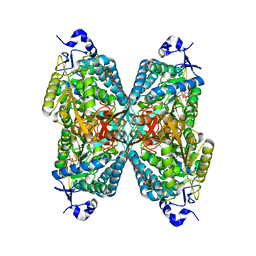 
 | | Crystal structure of Seabream Antiquitin and Elucidation of its substrate specificity | | Descriptor: | ANTIQUITIN, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Tang, W.K, Wong, K.B, Cha, S.S, Lee, H.S, Cheng, C.H.K, Fong, W.P. | | Deposit date: | 2007-02-09 | | Release date: | 2008-05-13 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.83 Å) | | Cite: | The Crystal Structure of Seabream Antiquitin Reveals the Structural Basis of its Substrate Specificity.
FEBS Lett., 582, 2008
|
|
2J6G
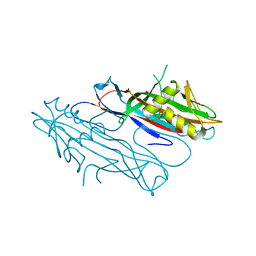 
 | | FaeG from F4ac ETEC strain 5_95, produced in tobacco plant chloroplast | | Descriptor: | ACETATE ION, FAEG | | Authors: | Van Molle, I, Joensuu, J.J, Buts, L, Panjikar, S, Kotiaho, M, Bouckaert, J, Wyns, L, Niklander-Teeri, V, De Greve, H. | | Deposit date: | 2006-09-28 | | Release date: | 2007-04-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Chloroplasts Assemble the Major Subunit Faeg of Escherichia Coli F4 (K88) Fimbriae Into Strand-Swapped Dimers
J.Mol.Biol., 368, 2007
|
|
5GLC
 
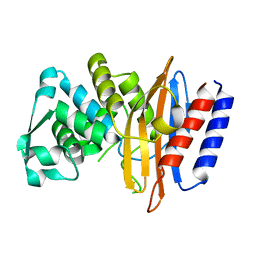 | | Crystal structure of the class A beta-lactamase PenL-tTR11 containing 20 residues insertion in omega-loop | | Descriptor: | Beta-lactamase | | Authors: | Choi, J.M, Yi, H, Kim, H.S, Lee, S.H. | | Deposit date: | 2016-07-10 | | Release date: | 2017-02-15 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.601 Å) | | Cite: | High adaptability of the omega loop underlies the substrate-spectrum-extension evolution of a class A beta-lactamase, PenL
Sci Rep, 6, 2016
|
|
5GIT
 
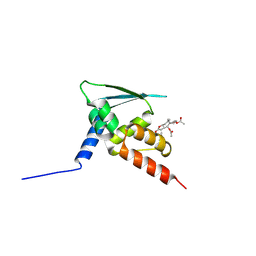 | | BTB domain of KEAP1 in complex with XX3 | | Descriptor: | Kelch-like ECH-associated protein 1, [(3aS,5R,5aS,6S,8S,8aS,9S,9aR)-9-acetyloxy-8-hydroxy-5,8a-dimethyl-1-methylidene-2-oxo-4,5,5a,6,7,8,9,9a-octahydro-3aH-azuleno[6,5-b]furan-6-yl] acetate | | Authors: | Zhu, L.L, Li, H.L, Wu, F.S, Xiong, R. | | Deposit date: | 2016-06-25 | | Release date: | 2017-06-28 | | Last modified: | 2019-01-23 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Britanin Ameliorates Cerebral Ischemia-Reperfusion Injury by Inducing the Nrf2 Protective Pathway.
Antioxid. Redox Signal., 27, 2017
|
|
8AFG
 
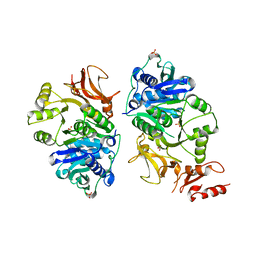 | |
5GLD
 
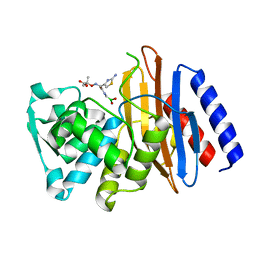 | | Crystal structure of the class A beta-lactamase PenL-tTR11 in complex with CBA | | Descriptor: | Beta-lactamase, PINACOL[[2-AMINO-ALPHA-(1-CARBOXY-1-METHYLETHOXYIMINO)-4-THIAZOLEACETYL]AMINO]METHANEBORONATE | | Authors: | Choi, J.M, Yi, H, Kim, H.S, Lee, S.H. | | Deposit date: | 2016-07-10 | | Release date: | 2017-02-15 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | High adaptability of the omega loop underlies the substrate-spectrum-extension evolution of a class A beta-lactamase, PenL
Sci Rep, 6, 2016
|
|
5GOT
 
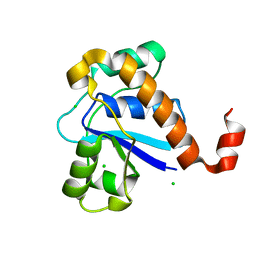 | | Crystal structure of SP-PTP, low molecular weight protein tyrosine phosphatase from Streptococcus pyogenes | | Descriptor: | CHLORIDE ION, Low molecular weight phosphotyrosine phosphatase family protein | | Authors: | Ku, B, Keum, C.W, KIim, S.J. | | Deposit date: | 2016-07-29 | | Release date: | 2016-09-07 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.902 Å) | | Cite: | Crystal structure of SP-PTP, a low molecular weight protein tyrosine phosphatase from Streptococcus pyogenes
Biochem.Biophys.Res.Commun., 478, 2016
|
|
2IIT
 
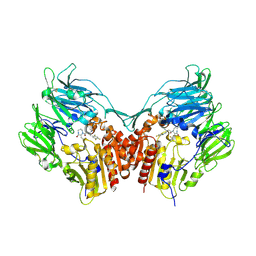 | | Human dipeptidyl peptidase 4 in complex with a diazepan-2-one inhibitor | | Descriptor: | (3R)-4-[(3R)-3-AMINO-4-(2,4,5-TRIFLUOROPHENYL)BUTANOYL]-3-(2,2,2-TRIFLUOROETHYL)-1,4-DIAZEPAN-2-ONE, 2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Scapin, G, Biftu, T, Weber, A.E. | | Deposit date: | 2006-09-28 | | Release date: | 2006-11-28 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | (3R)-4-[(3R)-3-Amino-4-(2,4,5-trifluorophenyl)butanoyl]-3-(2,2,2-trifluoroethyl)-1,4-diazepan-2-one, a selective dipeptidyl peptidase IV inhibitor for the treatment of type 2 diabetes
Bioorg.Med.Chem.Lett., 17, 2007
|
|
8AUZ
 
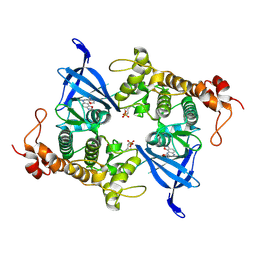 | | Crystal structure of GSK3 beta (GSK3b) in complex with FL291. | | Descriptor: | 8-morpholin-4-yl-2-pyridin-3-yl-[1,3]oxazolo[5,4-f]quinoxaline, Glycogen synthase kinase-3 beta, SULFATE ION | | Authors: | Chaikuad, A, Mongin, F, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2022-08-26 | | Release date: | 2023-04-19 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.66 Å) | | Cite: | Oxazolo[5,4-f]quinoxaline-type selective inhibitors of glycogen synthase kinase-3 alpha (GSK-3 alpha ): Development and impact on temozolomide treatment of glioblastoma cells.
Bioorg.Chem., 134, 2023
|
|
8AV1
 
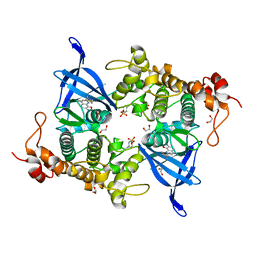 | | Crystal structure of GSK3 beta (GSK3b) in complex with CD7. | | Descriptor: | 1,2-ETHANEDIOL, 2-pyridin-3-yl-8-thiomorpholin-4-yl-[1,3]oxazolo[5,4-f]quinoxaline, Glycogen synthase kinase-3 beta, ... | | Authors: | Chaikuad, A, Mongin, F, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2022-08-26 | | Release date: | 2023-04-19 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Oxazolo[5,4-f]quinoxaline-type selective inhibitors of glycogen synthase kinase-3 alpha (GSK-3 alpha ): Development and impact on temozolomide treatment of glioblastoma cells.
Bioorg.Chem., 134, 2023
|
|
8DT0
 
 | |
