1CNH
 
 | |
4CAV
 
 | | Crystal structure of Aspergillus fumigatus N-myristoyl transferase in complex with myristoyl CoA and a benzofuran ligand R0-09-4879 | | Descriptor: | 3-[[3-methyl-2-[[2,3,4-tris(fluoranyl)phenoxy]methyl]-1-benzofuran-4-yl]oxy]-N-(pyridin-3-ylmethyl)propan-1-amine, CHLORIDE ION, GLYCYLPEPTIDE N-TETRADECANOYLTRANSFERASE, ... | | Authors: | Robinson, D.A, Fang, W, Raimi, O.G, Blair, D.E, Harrison, J, Ruda, G.F, Lockhart, D.E.A, Torrie, L.S, Wyatt, P.G, Gilbert, I.H, Van Aalten, D.M.F. | | Deposit date: | 2013-10-09 | | Release date: | 2014-09-17 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | N-Myristoyltransferase is a Cell Wall Target in Aspergillus Fumigatus.
Acs Chem.Biol., 10, 2015
|
|
4LW9
 
 | | Crystal structure of Vibrio cholera major pseudopilin EpsG | | Descriptor: | CALCIUM ION, CHLORIDE ION, PLATINUM (II) ION, ... | | Authors: | Vago, F.S, Raghunathan, K, Jens, J.C, Wedemeyer, W.J, Bagdasarian, M, Brunzelle, J.S, Arvidson, D.N. | | Deposit date: | 2013-07-26 | | Release date: | 2014-07-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of Vibrio cholera major pseudopilin EpsG
To be Published
|
|
1HY2
 
 | | MINIPROTEIN MP-1 COMPLEX WITH STREPTAVIDIN | | Descriptor: | MP-1, STREPTAVIDIN | | Authors: | Yang, H.W, Liu, D.Q, Fan, X, White, M.A, Fox, R.O. | | Deposit date: | 2001-01-17 | | Release date: | 2003-06-17 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Conformational Ensemble Analysis of Ligand Binding in Streptavidin Mini-Protein Complexes
To be Published
|
|
1P27
 
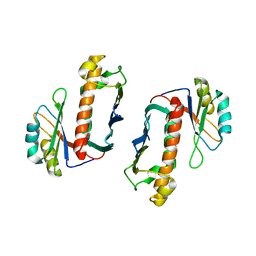 | | Crystal Structure of the Human Y14/Magoh complex | | Descriptor: | Mago nashi protein homolog, RNA-binding protein 8A | | Authors: | Lau, C.K, Diem, M.D, Dreyfuss, G, Van Duyne, G.D. | | Deposit date: | 2003-04-14 | | Release date: | 2003-08-19 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the y14-magoh core of the exon junction complex.
Curr.Biol., 13, 2003
|
|
4BRB
 
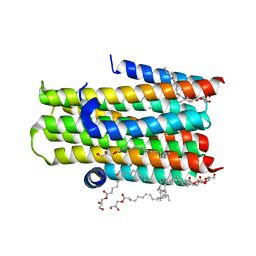 | | Crystal structure of the integral membrane enzyme DgkA-ref, delta 7 | | Descriptor: | (2R)-2,3-DIHYDROXYPROPYL(7Z)-PENTADEC-7-ENOATE, ACETATE ION, CITRATE ANION, ... | | Authors: | Li, D, Caffrey, M. | | Deposit date: | 2013-06-04 | | Release date: | 2014-06-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Renaturing Membrane Proteins in the Lipid Cubic Phase, a Nanoporous Membrane Mimetic.
Sci.Rep., 4, 2014
|
|
4Q32
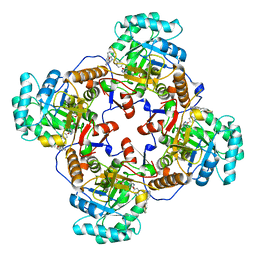 
 | | Crystal Structure of Inosine 5'-monophosphate Dehydrogenase from Clostridium perfringens Complexed with IMP and C91 | | Descriptor: | INOSINIC ACID, Inosine-5'-monophosphate dehydrogenase, N-(naphthalen-2-yl)-2-[2-(pyridin-2-yl)-1H-benzimidazol-1-yl]acetamide | | Authors: | Maltseva, N, Kim, Y, Makowska-Grzyska, M, Mulligan, R, Gu, M, Zhang, M, Mandapati, K, Gollapalli, D.R, Gorla, S.K, Hedstrom, L, Anderson, W.F, Joachimiak, A, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2014-04-10 | | Release date: | 2014-07-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.788 Å) | | Cite: | Crystal Structure of Inosine 5'-monophosphate Dehydrogenase from Clostridium perfringens Complexed with IMP and C91
To be Published
|
|
4Q5V
 
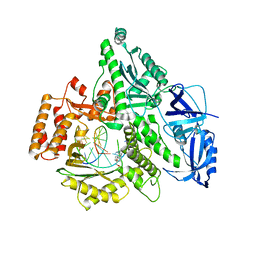 | | Crystal structure of the catalytic core of human DNA polymerase alpha in ternary complex with an RNA-primed DNA template and aphidicolin | | Descriptor: | (3R,4R,4aR,6aS,8R,9R,11aS,11bS)-4,9-bis(hydroxymethyl)-4,11b-dimethyltetradecahydro-8,11a-methanocyclohepta[a]naphthale ne-3,9-diol, DNA polymerase alpha catalytic subunit, DNA template, ... | | Authors: | Baranovskiy, A.G, Babayeva, N.D, Suwa, Y, Gu, J, Tahirov, T.H. | | Deposit date: | 2014-04-17 | | Release date: | 2014-11-26 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.52 Å) | | Cite: | Structural basis for inhibition of DNA replication by aphidicolin.
Nucleic Acids Res., 42, 2014
|
|
2WGM
 
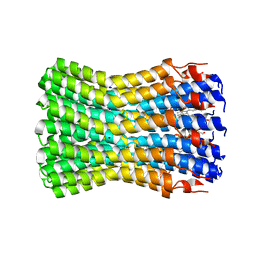 | | Complete ion-coordination structure in the rotor ring of Na-dependent F-ATP synthase | | Descriptor: | ATP SYNTHASE SUBUNIT C, SODIUM ION SPECIFIC, NONAN-1-OL, ... | | Authors: | Meier, T, Pogoryelov, D, Diederichs, K. | | Deposit date: | 2009-04-21 | | Release date: | 2009-06-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Complete Ion-Coordination Structure in the Rotor Ring of Na(+)-Dependent F-ATP Synthases.
J.Mol.Biol., 391, 2009
|
|
2GZO
 
 | | NMR structure of UPF0301 PROTEIN SO3346 from Shewanella oneidensis: Northeast Structural Genomics Consortium target SOR39 | | Descriptor: | UPF0301 protein SO3346 | | Authors: | Singarapu, K.K, Liu, G, Eletsky, A, Xu, D, Sukumaran, D.K, Mei, J, Xiao, R, Cunningham, K, Ma, L.C, Ritu, S, Acton, T.B, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2006-05-11 | | Release date: | 2006-06-20 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | NMR structure of UPF0301 PROTEIN SO3346 from Shewanella oneidensis: Northeast Structural Genomics Consortium target SOR39
To be Published
|
|
2H16
 
 | | Structure of human ADP-ribosylation factor-like 5 (ARL5) | | Descriptor: | ADP-ribosylation factor-like protein 5A, GUANOSINE-5'-DIPHOSPHATE, UNKNOWN ATOM OR ION | | Authors: | Rabeh, W.M, Tempel, W, Yaniw, D, Arrowsmith, C.H, Edwards, A.M, Sundstrom, M, Weigelt, J, Bochkarev, A, Park, H, Structural Genomics Consortium (SGC) | | Deposit date: | 2006-05-16 | | Release date: | 2006-06-13 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of human ADP-ribosylation factor-like 5 (ARL5)
To be Published
|
|
2E36
 
 | | L11 with SANS refinement | | Descriptor: | 50S ribosomal protein L11 | | Authors: | Lee, D, Walsh, J.D, Yu, P, Markus, M.A, Choli-Papadopoulous, T, Schwieters, C.D, Krueger, S, Draper, D.E, Wang, Y.X. | | Deposit date: | 2006-11-20 | | Release date: | 2007-06-19 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The structure of free L11 and functional dynamics of L11 in free, L11-rRNA(58 nt) binary and L11-rRNA(58 nt)-thiostrepton ternary complexes
J.Mol.Biol., 367, 2007
|
|
3SOH
 
 | | Architecture of the Flagellar Rotor | | Descriptor: | Flagellar motor switch protein FliG, Flagellar motor switch protein FliM | | Authors: | Koushik, P, Gonzalez-Bonet, G, Bilwes, A.M, Crane, B.R, Blair, D. | | Deposit date: | 2011-06-30 | | Release date: | 2011-07-27 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Architecture of the flagellar rotor.
Embo J., 30, 2011
|
|
1YAC
 
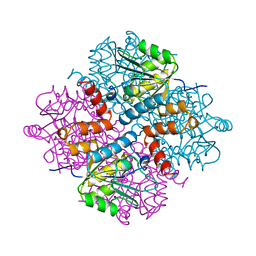 | |
2H62
 
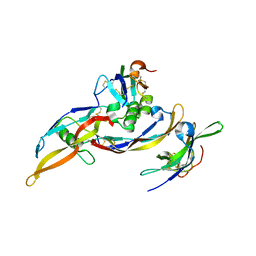 | | Crystal structure of a ternary ligand-receptor complex of BMP-2 | | Descriptor: | Acvr2b protein, Bone morphogenetic protein 2, Bone morphogenetic protein receptor type IA | | Authors: | Mueller, T.D. | | Deposit date: | 2006-05-30 | | Release date: | 2007-04-10 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | A silent H-bond can be mutationally activated for high-affinity interaction of BMP-2 and activin type IIB receptor.
Bmc Struct.Biol., 7, 2007
|
|
1I1B
 
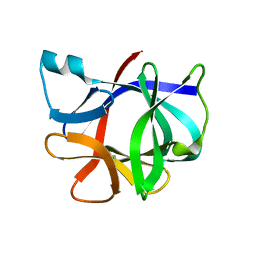 | |
1P3D
 
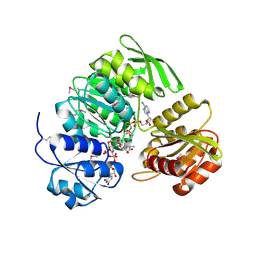 | | Crystal Structure of UDP-N-acetylmuramic acid:L-alanine ligase (MurC) in Complex with UMA and ANP. | | Descriptor: | MANGANESE (II) ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, UDP-N-acetylmuramate--alanine ligase, ... | | Authors: | Mol, C.D, Brooun, A, Dougan, D.R, Hilgers, M.T, Tari, L.W, Wijnands, R.A, Knuth, M.W, McRee, D.E, Swanson, R.V. | | Deposit date: | 2003-04-17 | | Release date: | 2003-07-15 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structures of Active Fully Assembled Substrate- and Product-Bound Complexes of UDP-N-Acetylmuramic Acid:L-Alanine Ligase (MurC) from Haemophilus influenzae.
J.Bacteriol., 185, 2003
|
|
2W9N
 
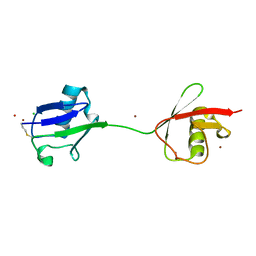 | | crystal structure of linear di-ubiquitin | | Descriptor: | CHLORIDE ION, UBIQUITIN, ZINC ION | | Authors: | Komander, D, Reyes-Turcu, F, Wilkinson, K.D, Barford, D. | | Deposit date: | 2009-01-27 | | Release date: | 2009-04-28 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Molecular Discrimination of Structurally Equivalent Lys 63-Linked and Linear Polyubiquitin Chains.
Embo Rep., 10, 2009
|
|
2GQR
 
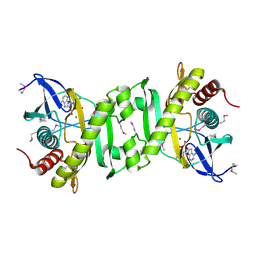 | | SAICAR Synthetase Complexed with ADP-Mg2+ | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Phosphoribosylaminoimidazole-succinocarboxamide synthase | | Authors: | Ginder, N.D, Honzatko, R.B. | | Deposit date: | 2006-04-21 | | Release date: | 2006-05-16 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Nucleotide Complexes of Escherichia coli Phosphoribosylaminoimidazole Succinocarboxamide Synthetase.
J.Biol.Chem., 281, 2006
|
|
2KC7
 
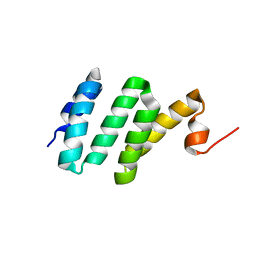 | | Solution NMR structure of Bacteroides fragilis protein BF1650. Northeast Structural Genomics Consortium target BfR218 | | Descriptor: | bfr218_protein | | Authors: | Eletsky, A, Wu, Y, Sukumaran, D, Lee, H, Lee, D.Y, Jiang, M, Foote, E.L, Xiao, R, Nair, R, Everett, J.K, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Prestegard, J.H, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-12-17 | | Release date: | 2009-01-13 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of Bacteroides fragilis protein BF1650. Northeast Structural Genomics Consortium target BfR218
To be Published
|
|
2GJ4
 
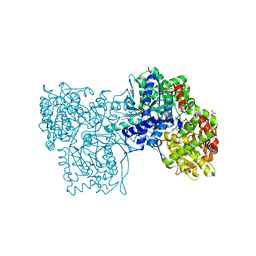 | | Structure of rabbit muscle glycogen phosphorylase in complex with ligand | | Descriptor: | (5-HYDROXY-4,6-DIMETHYLPYRIDIN-3-YL)METHYL DIHYDROGEN PHOSPHATE, 2-CHLORO-N-[(1R,2R)-1-HYDROXY-2,3-DIHYDRO-1H-INDEN-2-YL]-6H-THIENO[2,3-B]PYRROLE-5-CARBOXAMIDE, Glycogen phosphorylase, ... | | Authors: | Otterbein, L.R, Pannifer, A.D, Tucker, J, Breed, J, Oikonomakos, N.G, Rowsell, S, Pauptit, R.A, Claire, M. | | Deposit date: | 2006-03-30 | | Release date: | 2007-02-13 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Novel thienopyrrole glycogen phosphorylase inhibitors: synthesis, in vitro SAR and crystallographic studies.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
2ZJP
 
 | | Thiopeptide antibiotic Nosiheptide bound to the large ribosomal subunit of Deinococcus radiodurans | | Descriptor: | 4-(hydroxymethyl)-3-methyl-1H-indole-2-carboxylic acid, 50S RIBOSOMAL PROTEIN L11, 50S RIBOSOMAL PROTEIN L13, ... | | Authors: | Harms, J.M, Wilson, D.N, Schluenzen, F, Connell, S.R, Stachelhaus, T, Zaborowska, Z, Spahn, C.M.T, Fucini, P. | | Deposit date: | 2008-03-07 | | Release date: | 2008-06-17 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | Translational Regulation Via L11: Molecular Switches on the Ribosome Turned on and Off by Thiostrepton and Micrococcin.
Mol.Cell, 30, 2008
|
|
1I3P
 
 | |
2ZM0
 
 | | Structure of 6-aminohexanoate-dimer hydrolase, G181D/H266N/D370Y mutant | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 6-aminohexanoate-dimer hydrolase, GLYCEROL, ... | | Authors: | Ohki, T, Shibata, N, Higuchi, Y, Kawashima, Y, Takeo, M, Kato, D, Nego, S. | | Deposit date: | 2008-04-10 | | Release date: | 2009-04-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Molecular design of a nylon-6 byproduct-degrading enzyme from a carboxylesterase with a beta-lactamase fold.
Febs J., 276, 2009
|
|
2KMC
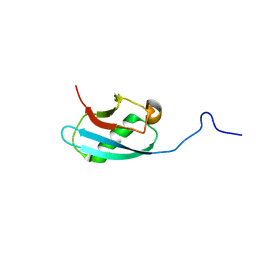 
 | | Solution Structure of the N-terminal domain of kindlin-1 | | Descriptor: | Fermitin family homolog 1 | | Authors: | Goult, B.T, Bate, N, Roberts, G.C, Barsukov, I.L, Critchley, D.R. | | Deposit date: | 2009-07-27 | | Release date: | 2009-10-06 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The Structure of the N-Terminus of Kindlin-1: A Domain Important for alphaIIbbeta3 Integrin Activation
J.Mol.Biol., 394, 2009
|
|
