4V01
 
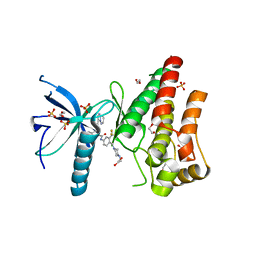 | | FGFR1 in complex with ponatinib (co-crystallisation). | | Descriptor: | 1,2-ETHANEDIOL, 3-(imidazo[1,2-b]pyridazin-3-ylethynyl)-4-methyl-N-{4-[(4-methylpiperazin-1-yl)methyl]-3-(trifluoromethyl)phenyl}benzamide, FIBROBLAST GROWTH FACTOR RECEPTOR 1 (FMS-RELATED TYROSINE KINASE 2, ... | | Authors: | Tucker, J, Klein, T, Breed, J, Breeze, A, Overman, R, Phillips, C, Norman, R.A. | | Deposit date: | 2014-09-10 | | Release date: | 2014-12-10 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Structural Insights Into Fgfr Kinase Isoform Selectivity: Diverse Binding Modes of Azd4547 and Ponatinib in Complex with Fgfr1 and Fgfr4
Structure, 22, 2014
|
|
4V05
 
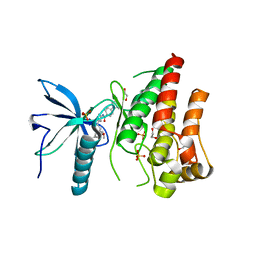 | | FGFR1 in complex with AZD4547. | | Descriptor: | 1,2-ETHANEDIOL, FIBROBLAST GROWTH FACTOR RECEPTOR 1 (FMS-RELATED TYROSINE KINASE 2, PFEIFFER SYNDROME), ... | | Authors: | Tucker, J, Klein, T, Breed, J, Breeze, A, Overman, R, Phillips, C, Norman, R.A. | | Deposit date: | 2014-09-10 | | Release date: | 2014-12-10 | | Last modified: | 2014-12-17 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | Structural Insights Into Fgfr Kinase Isoform Selectivity: Diverse Binding Modes of Azd4547 and Ponatinib in Complex with Fgfr1 and Fgfr4
Structure, 22, 2014
|
|
4UXQ
 
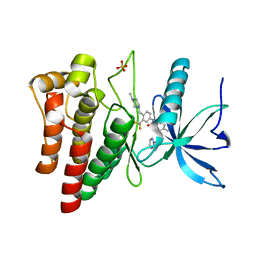 | | FGFR4 in complex with Ponatinib | | Descriptor: | 3-(imidazo[1,2-b]pyridazin-3-ylethynyl)-4-methyl-N-{4-[(4-methylpiperazin-1-yl)methyl]-3-(trifluoromethyl)phenyl}benzamide, FIBROBLAST GROWTH FACTOR RECEPTOR 4, SULFATE ION | | Authors: | Tucker, J, Klein, T, Breed, J, Breeze, A, Overman, R, Phillips, C, Norman, R.A. | | Deposit date: | 2014-08-27 | | Release date: | 2014-12-03 | | Last modified: | 2014-12-17 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural Insights Into Fgfr Kinase Isoform Selectivity: Diverse Binding Modes of Azd4547 and Ponatinib in Complex with Fgfr1 and Fgfr4
Structure, 22, 2014
|
|
5LHB
 
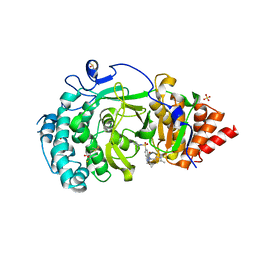 | | POLYADPRIBOSYL GLYCOSIDASE IN COMPLEX WITH PDD00017262 | | Descriptor: | 1-(cyclopropylmethyl)-6-[[(1-methylcyclopropyl)amino]-bis(oxidanyl)-$l^{4}-sulfanyl]-3-[(2-methyl-1,3-thiazol-5-yl)methyl]quinazoline-2,4-dione, DIMETHYL SULFOXIDE, Poly(ADP-ribose) glycohydrolase, ... | | Authors: | Tucker, J, Barkauskaite, E. | | Deposit date: | 2016-07-10 | | Release date: | 2016-10-12 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | First-in-Class Chemical Probes against Poly(ADP-ribose) Glycohydrolase (PARG) Inhibit DNA Repair with Differential Pharmacology to Olaparib.
ACS Chem. Biol., 11, 2016
|
|
4V04
 
 | | FGFR1 in complex with ponatinib. | | Descriptor: | 1,2-ETHANEDIOL, 3-(imidazo[1,2-b]pyridazin-3-ylethynyl)-4-methyl-N-{4-[(4-methylpiperazin-1-yl)methyl]-3-(trifluoromethyl)phenyl}benzamide, FIBROBLAST GROWTH FACTOR RECEPTOR 1 (FMS-RELATED TYROSINE KINASE 2, ... | | Authors: | Tucker, J, Klein, T, Breed, J, Breeze, A, Overman, R, Phillips, C, Norman, R.A. | | Deposit date: | 2014-09-10 | | Release date: | 2014-12-10 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Structural Insights Into Fgfr Kinase Isoform Selectivity: Diverse Binding Modes of Azd4547 and Ponatinib in Complex with Fgfr1 and Fgfr4
Structure, 22, 2014
|
|
7PUS
 
 | | ERK5 in complex with Pyrrole Carboxamide scaffold | | Descriptor: | 4-[3,6-bis(chloranyl)-2-fluoranyl-phenyl]carbonyl-~{N}-(1-methylpyrazol-4-yl)-1~{H}-pyrrole-2-carboxamide, Mitogen-activated protein kinase 7 | | Authors: | Tucker, J.A, Martin, M.P, Endicott, J.A, Noble, M.E.N. | | Deposit date: | 2021-09-30 | | Release date: | 2022-05-11 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Parallel Optimization of Potency and Pharmacokinetics Leading to the Discovery of a Pyrrole Carboxamide ERK5 Kinase Domain Inhibitor.
J.Med.Chem., 65, 2022
|
|
5O7I
 
 | | ERK5 in complex with a pyrrole inhibitor | | Descriptor: | 4-(2-bromanyl-6-fluoranyl-phenyl)carbonyl-~{N}-pyridin-3-yl-1~{H}-pyrrole-2-carboxamide, DIMETHYL SULFOXIDE, Mitogen-activated protein kinase 7 | | Authors: | Tucker, J.A, Heptinstall, A, Myers, S. | | Deposit date: | 2017-06-08 | | Release date: | 2018-06-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Identification of a novel orally bioavailable ERK5 inhibitor with selectivity over p38 alpha and BRD4.
Eur.J.Med.Chem., 178, 2019
|
|
6TD4
 
 | | IRF4 DNA-binding domain surface entropy mutant apo structure | | Descriptor: | CHLORIDE ION, Interferon regulatory factor 4 | | Authors: | Tucker, J.A, Martin, M.P, Wang, L.Z, Jennings, C, Heath, R. | | Deposit date: | 2019-11-07 | | Release date: | 2020-11-18 | | Last modified: | 2021-06-23 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Cancer-associated mutations in the IRF4 DNA-binding domain confer no disadvantage in DNA-binding affinity and may increase transcriptional activity
To Be Published
|
|
6HMK
 
 | | POLYADPRIBOSYL GLYCOHYDROLASE IN COMPLEX WITH PDD00016690 | | Descriptor: | 1-methyl-~{N}-(1-methylcyclopropyl)-3-[(2-methyl-1,3-thiazol-5-yl)methyl]-2,4-bis(oxidanylidene)quinazoline-6-sulfonamide, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | Tucker, J.A, Barkauskaite, E. | | Deposit date: | 2018-09-12 | | Release date: | 2018-11-14 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Cell-Active Small Molecule Inhibitors of the DNA-Damage Repair Enzyme Poly(ADP-ribose) Glycohydrolase (PARG): Discovery and Optimization of Orally Bioavailable Quinazolinedione Sulfonamides.
J.Med.Chem., 61, 2018
|
|
6HML
 
 | | POLYADPRIBOSYL GLYCOSIDASE IN COMPLEX WITH PDD00017299 | | Descriptor: | 1-[(2,4-dimethyl-1,3-thiazol-5-yl)methyl]-6-[[(1-methylcyclopropyl)amino]-bis(oxidanyl)-$l^{4}-sulfanyl]-3-[(1-methylpyrazol-4-yl)methyl]quinazoline-2,4-dione, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | Tucker, J.A, Barkauskaite, E. | | Deposit date: | 2018-09-12 | | Release date: | 2018-11-14 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Cell-Active Small Molecule Inhibitors of the DNA-Damage Repair Enzyme Poly(ADP-ribose) Glycohydrolase (PARG): Discovery and Optimization of Orally Bioavailable Quinazolinedione Sulfonamides.
J.Med.Chem., 61, 2018
|
|
6HMM
 
 | | POLYADPRIBOSYL GLYCOHYDROLASE IN COMPLEX WITH PDD00013907 | | Descriptor: | DIMETHYL SULFOXIDE, GLYCEROL, Poly(ADP-ribose) glycohydrolase, ... | | Authors: | Tucker, J.A, Brassington, C, Hassall, G. | | Deposit date: | 2018-09-12 | | Release date: | 2018-11-14 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Cell-Active Small Molecule Inhibitors of the DNA-Damage Repair Enzyme Poly(ADP-ribose) Glycohydrolase (PARG): Discovery and Optimization of Orally Bioavailable Quinazolinedione Sulfonamides.
J.Med.Chem., 61, 2018
|
|
6HMN
 
 | | POLYADPRIBOSYL GLYCOSIDASE IN COMPLEX WITH PDD00014909 | | Descriptor: | 3-methyl-6-[[(1-methylcyclopropyl)amino]-bis(oxidanyl)-$l^{4}-sulfanyl]-1-(phenylmethyl)quinazoline-2,4-dione, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | Tucker, J.A, Brassington, C, Hassall, G. | | Deposit date: | 2018-09-12 | | Release date: | 2018-11-14 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.87 Å) | | Cite: | Cell-Active Small Molecule Inhibitors of the DNA-Damage Repair Enzyme Poly(ADP-ribose) Glycohydrolase (PARG): Discovery and Optimization of Orally Bioavailable Quinazolinedione Sulfonamides.
J.Med.Chem., 61, 2018
|
|
4ZSG
 
 | | MITOGEN ACTIVATED PROTEIN KINASE 7 IN COMPLEX WITH INHIBITOR | | Descriptor: | 3-amino-5-[(4-chlorophenyl)amino]-N-(propan-2-yl)-1H-1,2,4-triazole-1-carboxamide, GLYCEROL, Mitogen-activated protein kinase 7 | | Authors: | Tucker, J, Ogg, D.J. | | Deposit date: | 2015-05-13 | | Release date: | 2016-05-04 | | Last modified: | 2016-05-11 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Discovery of a novel allosteric inhibitor-binding site in ERK5: comparison with the canonical kinase hinge ATP-binding site.
Acta Crystallogr D Struct Biol, 72, 2016
|
|
4ZSJ
 
 | | MITOGEN ACTIVATED PROTEIN KINASE 7 IN COMPLEX WITH INHIBITOR | | Descriptor: | 3-amino-5-[(4-chloro-3-methylphenyl)amino]-N-(propan-2-yl)-1H-1,2,4-triazole-1-carboxamide, GLYCEROL, Mitogen-activated protein kinase 7 | | Authors: | Tucker, J, Ogg, D.J. | | Deposit date: | 2015-05-13 | | Release date: | 2016-05-04 | | Last modified: | 2016-05-11 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Discovery of a novel allosteric inhibitor-binding site in ERK5: comparison with the canonical kinase hinge ATP-binding site.
Acta Crystallogr D Struct Biol, 72, 2016
|
|
4AJK
 
 | | rat LDHA in complex with N-(2-(methylamino)-1,3-benzothiazol-6-yl) acetamide | | Descriptor: | GLYCEROL, L-LACTATE DEHYDROGENASE A CHAIN, MALONATE ION, ... | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Vogtherr, M, Ward, R, Tart, J, Davies, G, Greenwood, R. | | Deposit date: | 2012-02-16 | | Release date: | 2012-03-21 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
4AJL
 | | rat LDHA in complex with 3-(ethylcarbamoylamino)-N-(2-methyl-1,3- benzothiazol-6-yl)propanamide | | Descriptor: | DIMETHYL SULFOXIDE, GLYCEROL, L-LACTATE DEHYDROGENASE A CHAIN, ... | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Vogtherr, M, Ward, R, Tart, J, Davies, G, Addie, M, Ferguson, A. | | Deposit date: | 2012-02-16 | | Release date: | 2012-03-21 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
4AJ2
 
 | | rat LDHA in complex with 5-(2-chlorophenyl)-1H-tetrazole | | Descriptor: | 5-(2-CHLOROPHENYL)-1H-TETRAZOLE, GLYCEROL, L-LACTATE DEHYDROGENASE A CHAIN, ... | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Vogtherr, M, Ward, R, Tart, J, Davies, G, Frazer, M. | | Deposit date: | 2012-02-15 | | Release date: | 2012-03-21 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
4AJN
 | | rat LDHA in complex with 2-((4-(2-((3-((2-methyl-1,3-benzothiazol-6- yl)amino)-3-oxo-propyl)carbamoylamino)ethyl)phenyl)methyl) propanedioic acid | | Descriptor: | (4-{2-[({3-[(2-METHYL-1,3-BENZOTHIAZOL-6-YL)AMINO]-3-OXOPROPYL}CARBAMOYL)AMINO]ETHYL}BENZYL)PROPANEDIOIC ACID, GLYCEROL, L-LACTATE DEHYDROGENASE A CHAIN | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Watson, M, Ward, R, Tart, J, Davies, G, Caputo, A, Pearson, S. | | Deposit date: | 2012-02-16 | | Release date: | 2012-03-21 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
4AJ1
 
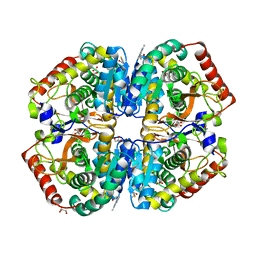 | | rat LDHA in complex with N-(2-(methylamino)-1,3-benzothiazol-6-yl) acetamide | | Descriptor: | GLYCEROL, L-LACTATE DEHYDROGENASE A CHAIN, MALONATE ION, ... | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Vogtherr, M, Ward, R, Tart, J, Davies, G. | | Deposit date: | 2012-02-15 | | Release date: | 2012-03-21 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
4AJJ
 | | rat LDHA in complex with 2-((3,4-dimethoxyphenyl)methyl))propanedioic acid and N-(2-methyl-1,3-benzothiazol-6-yl)-3-ureido-propanamide | | Descriptor: | 2-((3,4-DIMETHOXYPHENYL)METHYL))PROPANEDIOIC ACID, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Vogtherr, M, Ward, R, Tart, J, Davies, G, Patel, J, Greenwood, R. | | Deposit date: | 2012-02-16 | | Release date: | 2012-03-21 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
4AJI
 | | rat LDHA in complex with 2-((3,4-dimethoxyphenyl)methyl))propanedioic acid | | Descriptor: | 2-((3,4-DIMETHOXYPHENYL)METHYL))PROPANEDIOIC ACID, L-LACTATE DEHYDROGENASE A CHAIN | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Vogtherr, M, Ward, R, Tart, J, Davies, G, Debreczeni, J. | | Deposit date: | 2012-02-16 | | Release date: | 2012-03-21 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
4AJH
 | | rat LDHA in complex with N-(2-methyl-1,3-benzothiazol-6-yl)-3-ureido- propanamide and 2-(4-bromophenoxy)propanedioic acid | | Descriptor: | 2-(4-BROMANYLPHENOXY)PROPANEDIOIC ACID, GLYCEROL, L-LACTATE DEHYDROGENASE A CHAIN, ... | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Watson, M, Ward, R, Tart, J, Davies, G, Greenwood, R, Pearson, S, Debreczeni, J. | | Deposit date: | 2012-02-16 | | Release date: | 2012-03-21 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
4AL4
 
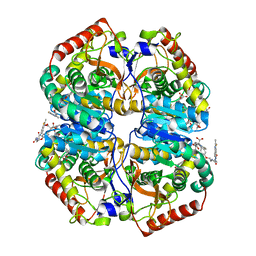 | | rat LDHA in complex with 2-((4-(2-((3-((2-methyl-1,3-benzothiazol-6- yl)amino)3-oxo-propyl)carbamoylamino)ethoxy)phenyl)methylpropanedioic acid | | Descriptor: | 2-[[4-[2-[[3-[(2-methyl-1,3-benzothiazol-6-yl)amino]-3-oxidanylidene-propyl]carbamoylamino]ethoxy]phenyl]methyl]propanedioic acid, GLYCEROL, L-LACTATE DEHYDROGENASE A CHAIN | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Ward, R, Tart, J, Davies, G, Hohson, M, Pearson, S. | | Deposit date: | 2012-03-01 | | Release date: | 2012-03-21 | | Last modified: | 2012-04-25 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
4AJE
 
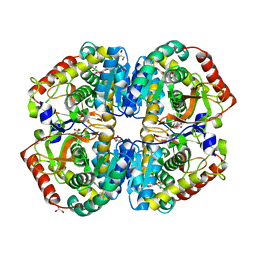 | | rat LDHA in complex with 2-(4-bromophenoxy)propanedioic acid | | Descriptor: | 2-(4-BROMANYLPHENOXY)PROPANEDIOIC ACID, GLYCEROL, L-LACTATE DEHYDROGENASE A CHAIN, ... | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Vogtherr, M, Ward, R, Tart, J, Davies, G, Pearson, S. | | Deposit date: | 2012-02-16 | | Release date: | 2012-03-21 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
4AJ4
 | | rat LDHA in complex with 4-((2-allylsulfanyl-1,3-benzothizol-6-yl) amino)-4-oxo-butanoic acid | | Descriptor: | 4-oxo-4-{[2-(prop-2-en-1-ylsulfanyl)-1,3-benzothiazol-6-yl]amino}butanoic acid, L-LACTATE DEHYDROGENASE A CHAIN, MALONATE ION | | Authors: | Tucker, J.A, Brassington, C, Hassall, G, Vogtherr, M, Ward, R, Tart, J, Davies, G. | | Deposit date: | 2012-02-15 | | Release date: | 2012-03-21 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Design and Synthesis of Novel Lactate Dehydrogenase a Inhibitors by Fragment-Based Lead Generation
J.Med.Chem., 55, 2012
|
|
