3P57
 
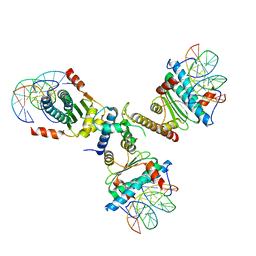 | | Crystal structure of the p300 TAZ2 domain bound to MEF2 on DNA | | Descriptor: | DNA (5'-D(*A*AP*AP*CP*TP*AP*TP*TP*TP*AP*TP*AP*AP*GP*A)-3'), DNA (5'-D(*TP*TP*CP*TP*TP*AP*TP*AP*AP*AP*TP*AP*GP*TP*T)-3'), Histone acetyltransferase p300, ... | | Authors: | He, J, Ye, J, Riquelme, C, Liu, J.O. | | Deposit date: | 2010-10-08 | | Release date: | 2011-08-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.1921 Å) | | Cite: | Structure of p300 bound to MEF2 on DNA reveals a mechanism of enhanceosome assembly.
Nucleic Acids Res., 39, 2011
|
|
6N4G
 
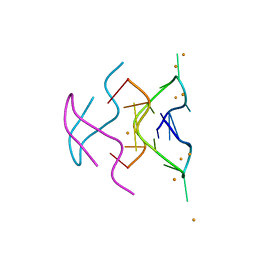 | |
1EN9
 | |
3AV4
 
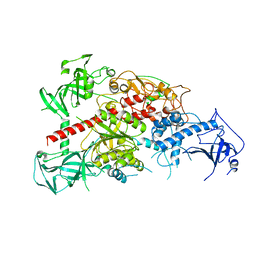 | | Crystal structure of mouse DNA methyltransferase 1 | | Descriptor: | DNA (cytosine-5)-methyltransferase 1, ZINC ION | | Authors: | Takeshita, K, Suetake, I, Yamashita, E, Suga, M, Narita, H, Nakagawa, A, Tajima, S. | | Deposit date: | 2011-02-22 | | Release date: | 2011-05-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural insight into maintenance methylation by mouse DNA methyltransferase 1 (Dnmt1).
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
6G3B
 
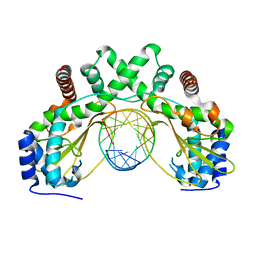 | | AvaII restriction endonuclease in complex with an RNA/DNA hybrid | | Descriptor: | DNA (5'-D(*CP*CP*AP*TP*GP*GP*TP*CP*CP*TP*A)-3'), RNA (5'-R(P*GP*UP*AP*GP*GP*AP*CP*CP*AP*UP*G)-3'), Type II site-specific deoxyribonuclease | | Authors: | Kisiala, M, Kowalska, M, Czapinska, H, Bochtler, M. | | Deposit date: | 2018-03-24 | | Release date: | 2019-04-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Restriction endonucleases that cleave RNA/DNA heteroduplexes bind dsDNA in A-like conformation
Nucleic Acids Res., 2020
|
|
6GVU
 
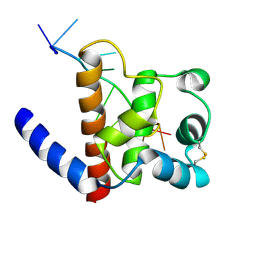 | |
6GZ7
 
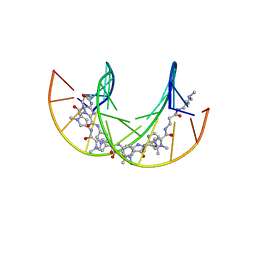 | | Polyamide - DNA complex NMR structure | | Descriptor: | DNA (5'-D(*CP*GP*AP*TP*GP*TP*AP*CP*AP*TP*CP*G)-3'), dimethyl-[3-[3-[[1-methyl-4-[[1-methyl-4-[[1-methyl-4-[[1-methyl-4-[4-[[1-methyl-4-[[1-methyl-4-[[1-methyl-4-[(1-propan-2-ylimidazol-2-yl)carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]butanoylamino]imidazol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]pyrrol-2-yl]carbonylamino]propanoylamino]propyl]azanium | | Authors: | Aman, K. | | Deposit date: | 2018-07-03 | | Release date: | 2018-11-28 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural and Kinetic Profiling of Allosteric Modulation of Duplex DNA Induced by DNA-Binding Polyamide Analogues.
Chemistry, 25, 2019
|
|
4GX9
 
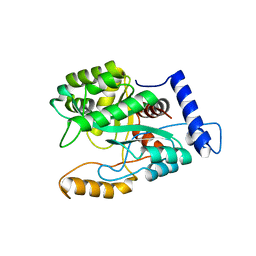 | | Crystal structure of a DNA polymerase III alpha-epsilon chimera | | Descriptor: | DNA polymerase III subunit epsilon,DNA polymerase III subunit alpha | | Authors: | Li, N, Horan, N, Xu, Z.-Q, Jacques, D, Dixon, N.E, Oakley, A.J. | | Deposit date: | 2012-09-04 | | Release date: | 2013-04-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Proofreading exonuclease on a tether: the complex between the E. coli DNA polymerase III subunits alpha, {varepsilon}, theta and beta reveals a highly flexible arrangement of the proofreading domain
Nucleic Acids Res., 41, 2013
|
|
1EN3
 | |
1ESG
 
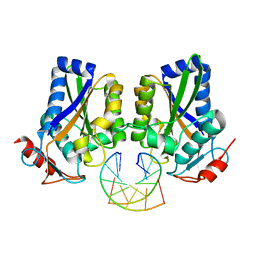 | |
5LS8
 
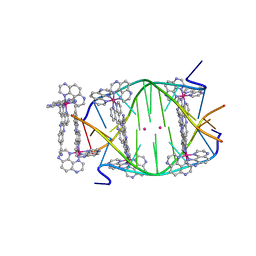 | | Light-activated ruthenium complex bound to a DNA quadruplex | | Descriptor: | DNA (5'-D(*TP*AP*GP*GP*GP*TP*TP*A)-3'), POTASSIUM ION, delta-[Ru(tap2-dppz-CN)]2+, ... | | Authors: | McQuaid, K.T, Abell, H, Hall, J.P, Cardin, C.J. | | Deposit date: | 2016-08-22 | | Release date: | 2018-01-17 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Structural Studies Reveal Enantiospecific Recognition of a DNA G-Quadruplex by a Ruthenium Polypyridyl Complex.
Angew.Chem.Int.Ed.Engl., 58, 2019
|
|
6M6K
 
 | |
6M6J
 | |
1C38
 
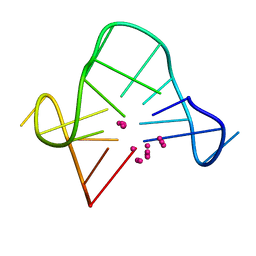 | | SOLUTION STRUCTURE OF A QUADRUPLEX FORMING DNA AND ITS INTERMEDIATE | | Descriptor: | DNA (5'-D(*GP*GP*TP*TP*GP*GP*TP*GP*TP*GP*GP*TP*TP*GP*G)-3'), POTASSIUM ION | | Authors: | Bolton, P.H, Marathias, V.M, Wang, K. | | Deposit date: | 1999-07-25 | | Release date: | 1999-08-25 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structures of the potassium-saturated, 2:1, and intermediate, 1:1, forms of a quadruplex DNA.
Nucleic Acids Res., 28, 2000
|
|
4K74
 
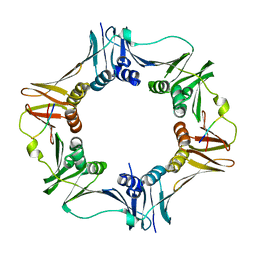 | |
6W89
 
 | | Structure of DNMT3A (R882H) in complex with CGA DNA | | Descriptor: | CGA DNA (25-MER), CITRIC ACID, DNA (cytosine-5)-methyltransferase 3-like, ... | | Authors: | Anteneh, H, Song, J. | | Deposit date: | 2020-03-20 | | Release date: | 2020-04-15 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.499 Å) | | Cite: | Structural basis for impairment of DNA methylation by the DNMT3A R882H mutation.
Nat Commun, 11, 2020
|
|
6W8J
 
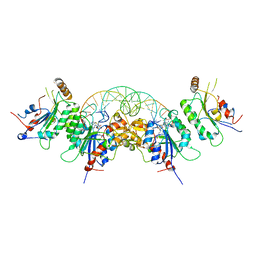 | | Structure of DNMT3A (R882H) in complex with CAG DNA | | Descriptor: | CAG DNA (25-MER), DNA (cytosine-5)-methyltransferase 3-like, DNA (cytosine-5)-methyltransferase 3A, ... | | Authors: | Anteneh, H, Song, J. | | Deposit date: | 2020-03-20 | | Release date: | 2020-04-15 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.445 Å) | | Cite: | Structural basis for impairment of DNA methylation by the DNMT3A R882H mutation.
Nat Commun, 11, 2020
|
|
6W8D
 
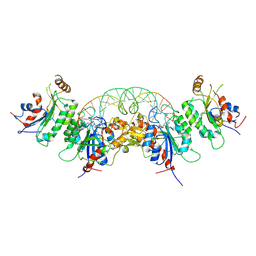 | | Structure of DNMT3A (R882H) in complex with CGT DNA | | Descriptor: | CGT DNA (25-MER), DNA (cytosine-5)-methyltransferase 3-like, DNA (cytosine-5)-methyltransferase 3A, ... | | Authors: | Anteneh, H, Song, J. | | Deposit date: | 2020-03-20 | | Release date: | 2020-04-15 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.598 Å) | | Cite: | Structural basis for impairment of DNA methylation by the DNMT3A R882H mutation.
Nat Commun, 11, 2020
|
|
7OHE
 
 | | A self-complementary DNA dodecamer duplex contaning 5-hydroxymethylcitosine | | Descriptor: | DNA (5'-D(*CP*GP*CP*GP*TP*CP*GP*AP*CP*GP*CP*G)-3') | | Authors: | Battistini, F, Dans, P.D, Terrazas, M, Castellazzi, C.L, Portella, G, Labrador, M, Villegas, N, Brun-Heath, I, Gonzalez, C, Orozco, M. | | Deposit date: | 2021-05-10 | | Release date: | 2021-11-03 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | The Impact of the HydroxyMethylCytosine epigenetic signature on DNA structure and function.
Plos Comput.Biol., 17, 2021
|
|
7OHM
 
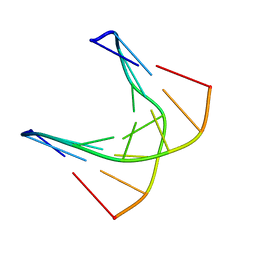 | | A self-complementary DNA dodecamer duplex contaning 5-hydroxymethylcitosine | | Descriptor: | DNA (5'-D(*CP*GP*AP*(DH)P*GP*TP*CP*G)-3') | | Authors: | Battistini, F, Dans, P.D, Terrazas, M, Castellazzi, C.L, Portella, G, Labrador, M, Villegas, N, Brun-Heath, I, Gonzalez, C, Orozco, M. | | Deposit date: | 2021-05-11 | | Release date: | 2021-11-03 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | The Impact of the HydroxyMethylCytosine epigenetic signature on DNA structure and function.
Plos Comput.Biol., 17, 2021
|
|
7OHJ
 
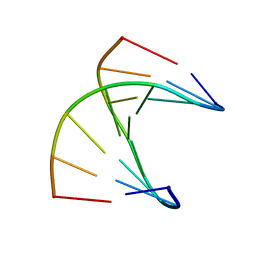 | | A self-complementary DNA dodecamer duplex contaning 5-hydroxymethylcitosine | | Descriptor: | DNA (5'-D(*CP*GP*AP*CP*GP*TP*CP*G)-3') | | Authors: | Battistini, F, Dans, P.D, Terrazas, M, Castellazzi, C.L, Portella, G, Labrador, M, Villegas, N, Brun-Heath, I, Gonzalez, C, Orozco, M. | | Deposit date: | 2021-05-11 | | Release date: | 2021-11-03 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | The Impact of the HydroxyMethylCytosine epigenetic signature on DNA structure and function.
Plos Comput.Biol., 17, 2021
|
|
7OGV
 
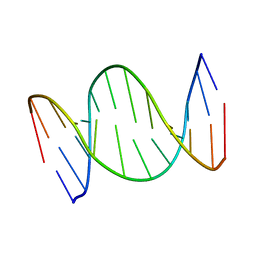 | | A self-complementary DNA dodecamer duplex contaning 5-hydroxymethylcitosine | | Descriptor: | DNA (5'-D(*(P*GP*CP*GP*TP*(DH)P*GP*AP*CP*GP*CP*G-3') | | Authors: | Battistini, F, Dans, P.D, Terrazas, M, Castellazzi, C.L, Portella, G, Labrador, M, Villegas, N, Brun-Heath, I, Gonzalez, C, Orozco, M. | | Deposit date: | 2021-05-07 | | Release date: | 2021-11-03 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | The Impact of the HydroxyMethylCytosine epigenetic signature on DNA structure and function.
Plos Comput.Biol., 17, 2021
|
|
8KGS
 | | Structure of African swine fever virus topoisomerase II in complex with dsDNA | | Descriptor: | DNA topoisomerase 2, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Cong, J, Xin, Y, Li, X, Chen, Y. | | Deposit date: | 2023-08-19 | | Release date: | 2024-04-03 | | Last modified: | 2024-08-14 | | Method: | X-RAY DIFFRACTION (2.603 Å) | | Cite: | Structural insights into the DNA topoisomerase II of the African swine fever virus.
Nat Commun, 15, 2024
|
|
8H7A
 
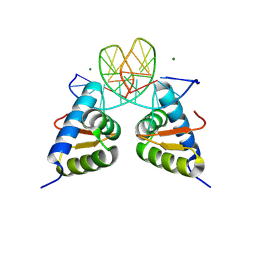 | | Crystal structure of the dimer form KAT6A WH domain with its bound double stranded DNA | | Descriptor: | DNA (5'-D(*GP*GP*TP*CP*CP*GP*AP*CP*GP*GP*AP*CP*C)-3'), DNA (5'-D(*GP*GP*TP*CP*CP*GP*TP*CP*GP*GP*AP*CP*C)-3'), Histone acetyltransferase KAT6A, ... | | Authors: | Wang, Z, Cao, Y. | | Deposit date: | 2022-10-19 | | Release date: | 2023-01-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | The histone acetyltransferase KAT6A is recruited to unmethylated CpG islands via a DNA binding winged helix domain.
Nucleic Acids Res., 51, 2023
|
|
7TZ1
 
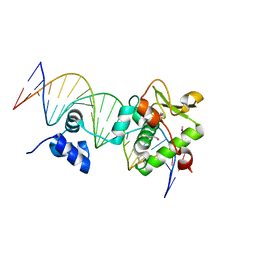 | | Crystal Structure of a Mycobacteriophage Cluster A2 Immunity Repressor:DNA Complex | | Descriptor: | DNA (5'-D(P*CP*CP*CP*GP*CP*TP*TP*GP*AP*CP*AP*GP*CP*CP*AP*CP*CP*GP*AP*AP*A)-3'), DNA (5'-D(P*TP*TP*TP*CP*GP*GP*TP*GP*GP*CP*TP*GP*TP*CP*AP*AP*GP*CP*GP*GP*G)-3'), Immunity repressor | | Authors: | McGinnis, R.J, Brambley, C.A, Stamey, B, Green, W.C, Gragg, K.N, Cafferty, E.R, Terwilliger, T.C, Hammel, M, Hollis, T.J, Miller, J.M, Gainey, M.D, Wallen, J.R. | | Deposit date: | 2022-02-15 | | Release date: | 2022-07-20 | | Last modified: | 2022-07-27 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | A monomeric mycobacteriophage immunity repressor utilizes two domains to recognize an asymmetric DNA sequence.
Nat Commun, 13, 2022
|
|
