1J0H
 
 | |
1J6Y
 
 | | Solution structure of Pin1At from Arabidopsis thaliana | | Descriptor: | peptidyl-prolyl cis-trans isomerase | | Authors: | Landrieu, I, Wieruszeski, J.M, Wintjens, R, Inze, D, Lippens, G. | | Deposit date: | 2001-05-15 | | Release date: | 2002-08-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the
Single-domain Prolyl Cis/Trans
Isomerase PIN1At from Arabidopsis thaliana
J.Mol.Biol., 320, 2002
|
|
1IQE
 
 | | Human coagulation factor Xa in complex with M55590 | | Descriptor: | 4-[(2R)-3-[[(6-CHLORO-2-NAPHTHALENYL)SULFONYL]AMINO]-1-OXO-2-[[[1-(4-PYRIDINYL)-4-PIPERIDINYL]METHYL]AMINO]PROPYL]-THIOMORPHOLINE-1,1-DIOXIDE, CALCIUM ION, coagulation Factor Xa | | Authors: | Shiromizu, I, Matsusue, T. | | Deposit date: | 2001-07-23 | | Release date: | 2003-09-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Factor Xa Specific Inhibitor that Induces the Novel Binding Model in Complex with Human Fxa
To be Published
|
|
1IQM
 
 | | Human coagulation factor Xa in complex with M54471 | | Descriptor: | 1-[[(1E)-2-(4-CHLOROPHENYL)ETHENYL]SULFONYL]-4-[[1-(4-PYRIDINYL)-4-PIPERIDINYL]METHYL]PIPERAZINE, CALCIUM ION, coagulation Factor Xa | | Authors: | Shiromizu, I, Matsusue, T. | | Deposit date: | 2001-07-23 | | Release date: | 2003-09-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Factor Xa Specific Inhibitor that Induces the Novel Binding Model in Complex with Human Fxa
To be Published
|
|
1IQK
 
 | | Human coagulation factor Xa in complex with M55113 | | Descriptor: | 4-[(6-CHLORO-2-NAPHTHALENYL)SULFONYL]-1-[[1-(4-PYRIDINYL)-4-PIPERIDINYL]METHYL]PIPERAZINONE, CALCIUM ION, coagulation Factor Xa | | Authors: | Shiromizu, I, Matsusue, T. | | Deposit date: | 2001-07-23 | | Release date: | 2003-09-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Factor Xa Specific Inhibitor that Induces the Novel Binding Model in Complex with Human Fxa
To be Published
|
|
2RG3
 
 | | Covalent complex structure of elastase | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Leukocyte elastase | | Authors: | Huang, W, Yamamoto, Y. | | Deposit date: | 2007-10-02 | | Release date: | 2008-07-01 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray snapshot of the mechanism of inactivation of human neutrophil elastase by 1,2,5-thiadiazolidin-3-one 1,1-dioxide derivatives.
J.Med.Chem., 51, 2008
|
|
1IQF
 
 | | Human coagulation factor Xa in complex with M55165 | | Descriptor: | (2R)-4-[(6-CHLORO-2-NAPHTHALENYL)SULFONYL]-6-OXO-1-[[1-(4-PYRIDINYL)-4-PIPERIDINYL]METHYL]-2-PIPERAZINECARBOXYLIC ACID ETHYL ESTER, CALCIUM ION, coagulation Factor Xa | | Authors: | Shiromizu, I, Matsusue, T. | | Deposit date: | 2001-07-23 | | Release date: | 2003-09-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Factor Xa Specific Inhibitor that Induces the Novel Binding Model in Complex with Human Fxa
To be Published
|
|
1IQL
 
 | | Human coagulation factor Xa in complex with M54476 | | Descriptor: | 4-[[(1E)-2-(4-CHLOROPHENYL)ETHENYL]SULFONYL]-1-[[1-(4-PYRIDINYL)-4-PIPERIDINYL]METHYL]PIPERAZINONE, CALCIUM ION, coagulation Factor Xa | | Authors: | Shiromizu, I, Matsusue, T. | | Deposit date: | 2001-07-23 | | Release date: | 2003-09-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Factor Xa Specific Inhibitor that Induces the Novel Binding Model in Complex with Human Fxa
To be Published
|
|
1J0I
 
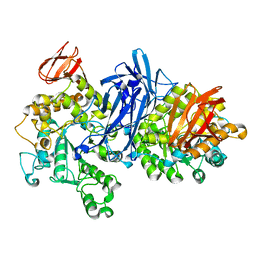 | | Crystal structure of neopullulanase complex with panose | | Descriptor: | alpha-D-glucopyranose-(1-6)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, neopullulanase | | Authors: | Hondoh, H, Kuriki, T, Matsuura, Y. | | Deposit date: | 2002-11-14 | | Release date: | 2003-01-28 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Three-dimensional structure and substrate binding of Bacillus stearothermophilus neopullulanase
J.Mol.Biol., 326, 2003
|
|
2TMN
 
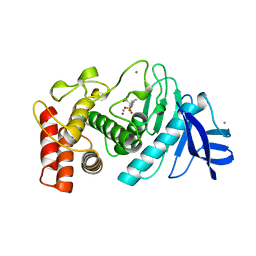 | | CRYSTALLOGRAPHIC STRUCTURAL ANALYSIS OF PHOSPHORAMIDATES AS INHIBITORS AND TRANSITION-STATE ANALOGS OF THERMOLYSIN | | Descriptor: | CALCIUM ION, N~2~-phosphono-L-leucinamide, Thermolysin, ... | | Authors: | Tronrud, D.E, Monzingo, A.F, Matthews, B.W. | | Deposit date: | 1987-06-29 | | Release date: | 1989-01-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystallographic structural analysis of phosphoramidates as inhibitors and transition-state analogs of thermolysin.
Eur.J.Biochem., 157, 1986
|
|
2RHA
 
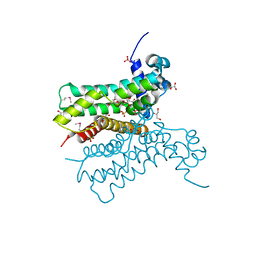 | |
1GZG
 
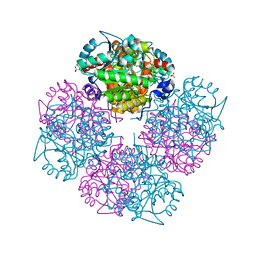 | | Complex of a Mg2-dependent porphobilinogen synthase from Pseudomonas aeruginosa (mutant D139N) with 5-fluorolevulinic acid | | Descriptor: | 5-FLUOROLEVULINIC ACID, DELTA-AMINOLEVULINIC ACID DEHYDRATASE, MAGNESIUM ION, ... | | Authors: | Frere, F, Schubert, W.-D, Stauffer, F, Frankenberg, N, Neier, R, Jahn, D, Heinz, D.W. | | Deposit date: | 2002-05-21 | | Release date: | 2002-06-27 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Structure of porphobilinogen synthase from Pseudomonas aeruginosa in complex with 5-fluorolevulinic acid suggests a double Schiff base mechanism.
J. Mol. Biol., 320, 2002
|
|
1HPX
 
 | | HIV-1 PROTEASE COMPLEXED WITH THE INHIBITOR KNI-272 | | Descriptor: | (4R)-N-tert-butyl-3-[(2S,3S)-2-hydroxy-3-({N-[(isoquinolin-5-yloxy)acetyl]-S-methyl-L-cysteinyl}amino)-4-phenylbutanoyl]-1,3-thiazolidine-4-carboxamide, HIV-1 PROTEASE | | Authors: | Bhat, T.N, Erickson, J.W. | | Deposit date: | 1995-05-18 | | Release date: | 1996-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of HIV-1 protease with KNI-272, a tight-binding transition-state analog containing allophenylnorstatine.
Structure, 3, 1995
|
|
2TSC
 
 | | STRUCTURE, MULTIPLE SITE BINDING, AND SEGMENTAL ACCOMODATION IN THYMIDYLATE SYNTHASE ON BINDING D/UMP AND AN ANTI-FOLATE | | Descriptor: | 10-PROPARGYL-5,8-DIDEAZAFOLIC ACID, 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, THYMIDYLATE SYNTHASE | | Authors: | Montfort, W.R, Stroud, R.M. | | Deposit date: | 1991-07-03 | | Release date: | 1991-10-15 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structure, multiple site binding, and segmental accommodation in thymidylate synthase on binding dUMP and an anti-folate.
Biochemistry, 29, 1990
|
|
2SGA
 
 | |
2VKR
 
 | | 3Fe-4S, 4Fe-4S plus Zn Acidianus ambivalens ferredoxin | | Descriptor: | FE3-S4 CLUSTER, IRON/SULFUR CLUSTER, ZINC ION, ... | | Authors: | Frazao, C, Aragao, D, Coelho, R, Leal, S.S, Gomes, C.M, Teixeira, M, Carrondo, M.A. | | Deposit date: | 2007-12-23 | | Release date: | 2008-03-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Crystallographic analysis of the intact metal centres [3Fe-4S](1+/0) and [4Fe-4S](2+/1+) in a Zn(2+) -containing ferredoxin.
FEBS Lett., 582, 2008
|
|
1JET
 
 | | OLIGO-PEPTIDE BINDING PROTEIN (OPPA) COMPLEXED WITH KAK | | Descriptor: | OLIGO-PEPTIDE BINDING PROTEIN, PEPTIDE LYS ALA LYS, URANYL (VI) ION | | Authors: | Tame, J, Wilkinson, A.J. | | Deposit date: | 1996-07-03 | | Release date: | 1997-05-15 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | The role of water in sequence-independent ligand binding by an oligopeptide transporter protein.
Nat.Struct.Biol., 3, 1996
|
|
2V1T
 
 | | CRYSTAL STRUCTURE OF RAT TOM20-ALDH PRESEQUENCE COMPLEX | | Descriptor: | ALDEHYDE DEHYDROGENASE, MITOCHONDRIAL IMPORT RECEPTOR SUBUNIT TOM20 HOMOLOG | | Authors: | Obita, T, Igura, M, Ose, T, Endo, T, Maenaka, K, Kohda, D. | | Deposit date: | 2007-05-29 | | Release date: | 2007-06-12 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Tom20 Recognizes Mitochondrial Presequences Through Dynamic Equilibrium Among Multiple Bound States.
Embo J., 26, 2007
|
|
2V27
 
 | | Structure of the cold active phenylalanine hydroxylase from Colwellia psychrerythraea 34H | | Descriptor: | FE (III) ION, PHENYLALANINE HYDROXYLASE, SULFATE ION | | Authors: | Leiros, H.-K.S, Pey, A.L, Innselset, M, Moe, E, Leiros, I, Steen, I.H, Martinez, A. | | Deposit date: | 2007-06-03 | | Release date: | 2007-06-19 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structure of Phenylalanine Hydroxylase from Colwellia Psychrerythraea 34H, a Monomeric Cold Active Enzyme with Local Flexibility Around the Active Site and High Overall Stability.
J.Biol.Chem., 282, 2007
|
|
1JR2
 
 | | Structure of Uroporphyrinogen III Synthase | | Descriptor: | UROPORPHYRINOGEN-III SYNTHASE | | Authors: | Mathews, M.A, Schubert, H.L, Whitby, F.G, Alexander, K.J, Schadick, K, Bergonia, H.A, Phillips, J.D, Hill, C.P. | | Deposit date: | 2001-08-10 | | Release date: | 2001-11-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Crystal structure of human uroporphyrinogen III synthase.
EMBO J., 20, 2001
|
|
2VBM
 
 | | Tailspike protein of bacteriophage Sf6 complexed with tetrasaccharide | | Descriptor: | MAGNESIUM ION, MANGANESE (II) ION, PHOSPHATE ION, ... | | Authors: | Mueller, J.J, Barbirz, S, Freiberg, A, Seckler, R, Heinemann, U. | | Deposit date: | 2007-09-14 | | Release date: | 2008-04-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | An Intersubunit Active Site between Supercoiled Parallel Beta Helices in the Trimeric Tailspike Endorhamnosidase of Shigella Flexneri Phage Sf6.
Structure, 16, 2008
|
|
2V28
 
 | | Apo structure of the cold active phenylalanine hydroxylase from Colwellia psychrerythraea 34H | | Descriptor: | PHENYLALANINE-4-HYDROXYLASE, SULFATE ION | | Authors: | Leiros, H.-K.S, Pey, A.L, Innselset, M, Moe, E, Leiros, I, Steen, I.H, Martinez, A. | | Deposit date: | 2007-06-04 | | Release date: | 2007-06-19 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure of Phenylalanine Hydroxylase from Colwellia Psychrerythraea 34H, a Monomeric Cold Active Enzyme with Local Flexibility Around the Active Site and High Overall Stability.
J.Biol.Chem., 282, 2007
|
|
1I8J
 
 | | CRYSTAL STRUCTURE OF PORPHOBILINOGEN SYNTHASE COMPLEXED WITH THE INHIBITOR 4,7-DIOXOSEBACIC ACID | | Descriptor: | 4,7-DIOXOSEBACIC ACID, MAGNESIUM ION, PORPHOBILINOGEN SYNTHASE, ... | | Authors: | Kervinen, J, Jaffe, E.K, Stauffer, F, Neier, R, Wlodawer, A, Zdanov, A. | | Deposit date: | 2001-03-14 | | Release date: | 2001-06-20 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Mechanistic basis for suicide inactivation of porphobilinogen synthase by 4,7-dioxosebacic acid, an inhibitor that shows dramatic species selectivity.
Biochemistry, 40, 2001
|
|
2R61
 
 | |
1HLF
 
 | | BINDING OF GLUCOPYRANOSYLIDENE-SPIRO-THIOHYDANTOIN TO GLYCOGEN PHOSPHORYLASE B: KINETIC AND CRYSTALLOGRAPHIC STUD | | Descriptor: | (5S,7R,8S,9S,10R)-8,9,10-trihydroxy-7-(hydroxymethyl)-2-thioxo-6-oxa-1,3-diazaspiro[4.5]decan-4-one, GLYCOGEN PHOSPHORYLASE, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Oikonomakos, N.G, Skamnaki, V.T, Docsa, T, Toth, B, Gergely, P, Osz, E, Szilagyi, L, Somsak, L. | | Deposit date: | 2000-12-01 | | Release date: | 2000-12-13 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Kinetic and crystallographic studies of glucopyranosylidene spirothiohydantoin
binding to glycogen phosphorylase B
BIOORG.MED.CHEM., 10, 2002
|
|
