4HPS
 
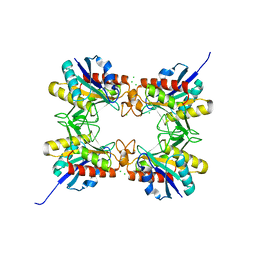 | |
2VRF
 
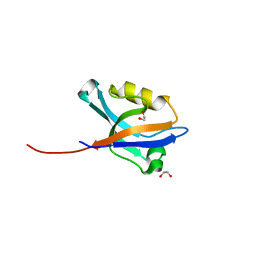 | | CRYSTAL STRUCTURE OF THE HUMAN BETA-2-SYNTROPHIN PDZ DOMAIN | | Descriptor: | 1,2-ETHANEDIOL, BETA-2-SYNTROPHIN | | Authors: | Sun, Z, Roos, A.K, Pike, A.C.W, Pilka, E.S, Cooper, C, Elkins, J.M, Murray, J, Arrowsmith, C.H, Doyle, D, Edwards, A, von Delft, F, Bountra, C, Oppermann, U. | | Deposit date: | 2008-03-31 | | Release date: | 2008-04-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structure of the Human Beta-2-Syntrophin Pdz Domain
To be Published
|
|
2H9I
 
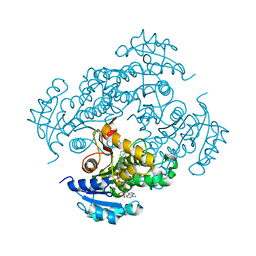 | | Mycobacterium tuberculosis InhA bound with ETH-NAD adduct | | Descriptor: | Enoyl-[acyl-carrier-protein] reductase [NADH], {(2R,3S,4R,5R)-5-[(4S)-3-(AMINOCARBONYL)-4-(2-ETHYLISONICOTINOYL)PYRIDIN-1(4H)-YL]-3,4-DIHYDROXYTETRAHYDROFURAN-2-YL}METHYL [(2R,3S,4R,5R)-5-(6-AMINO-9H-PURIN-9-YL)-3,4-DIHYDROXYTETRAHYDROFURAN-2-YL]METHYL DIHYDROGEN DIPHOSPHATE | | Authors: | Wang, F, Sacchettini, J.C. | | Deposit date: | 2006-06-09 | | Release date: | 2007-01-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Mechanism of thioamide drug action against tuberculosis and leprosy.
J.Exp.Med., 204, 2007
|
|
3DV1
 
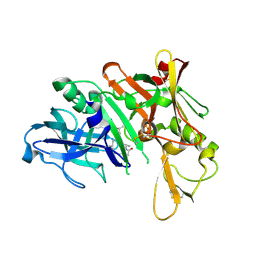 | | Crystal structure of human beta-secretase in complex with NVP-ARV999 | | Descriptor: | (2R,4S)-N-butyl-4-[(2S,5S,7R)-2,7-dimethyl-3,15-dioxo-1,4-diazacyclopentadecan-5-yl]-4-hydroxy-2-methylbutanamide, Beta-secretase 1 | | Authors: | Rondeau, J.-M. | | Deposit date: | 2008-07-18 | | Release date: | 2009-02-24 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Macrocyclic peptidomimetic beta-secretase (BACE-1) inhibitors with activity in vivo.
Bioorg.Med.Chem.Lett., 19, 2009
|
|
1JQ0
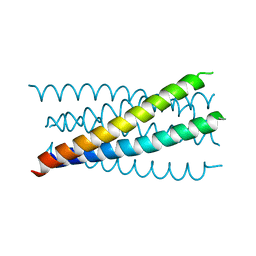 
 | | Mutation that destabilize the gp41 core: determinants for stabilizing the SIV/CPmac envelope glycoprotein complex. Mutant structure. | | Descriptor: | gp41 envelope protein | | Authors: | Liu, J, Wang, S, LaBranche, C.C, Hoxie, J.A, Lu, M. | | Deposit date: | 2001-08-03 | | Release date: | 2002-04-24 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Mutations that destabilize the gp41 core are determinants for stabilizing the simian immunodeficiency virus-CPmac envelope glycoprotein complex.
J.Biol.Chem., 277, 2002
|
|
2O34
 
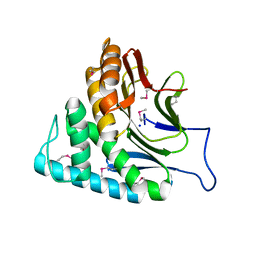 | | Crystal structure of protein DVU1097 from Desulfovibrio vulgaris Hildenborough, Pfam DUF375 | | Descriptor: | Hypothetical protein, SODIUM ION | | Authors: | Malashkevich, V.N, Toro, R, Sauder, J.M, Schwinn, K.D, Thompson, D.A, Rutter, M.E, Dickey, M, Groshong, C, Bain, K.T, Adams, J.M, Reyes, C, Rooney, I, Powell, A, Boice, A, Gheyi, T, Ozyurt, S, Atwell, S, Wasserman, S.R, Emtage, S, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2006-11-30 | | Release date: | 2006-12-12 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure of the Hypothetical Protein from Desulfovibrio vulgaris Hildenborough
To be Published
|
|
1MD7
 
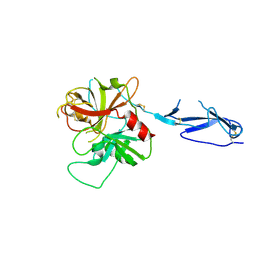 | | Monomeric structure of the zymogen of complement protease C1r | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, C1R COMPLEMENT SERINE PROTEASE | | Authors: | Budayova-Spano, M, Grabarse, W, Thielens, N.M, Hillen, H, Lacroix, M, Schmidt, M, Fontecilla-Camps, J, Arlaud, G.J, Gaboriaud, C. | | Deposit date: | 2002-08-07 | | Release date: | 2003-08-07 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Monomeric structures of the zymogen and active catalytic domain of complement protease c1r: further insights into the c1 activation mechanism
Structure, 10, 2002
|
|
3DWK
 
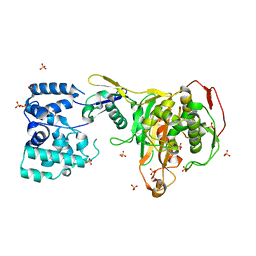 | |
3RJE
 
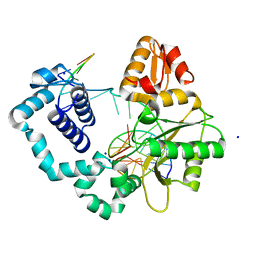 | | Ternary complex of DNA Polymerase Beta with a gapped DNA containing 8odG at template position | | Descriptor: | DNA (5'-D(*CP*CP*GP*AP*CP*(8OG)P*TP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3'), DNA (5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*A)-3'), DNA (5'-D(P*GP*TP*CP*GP*G)-3'), ... | | Authors: | Batra, V.K, Beard, W.A, Wilson, S.H. | | Deposit date: | 2011-04-15 | | Release date: | 2012-01-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Binary complex crystal structure of DNA polymerase beta reveals multiple conformations of the templating 8-oxoguanine lesion
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
2B3J
 
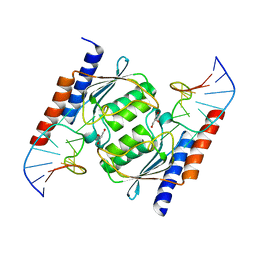 | | Crystal Structure of Staphylococcus aureus tRNA Adenosine Deaminase, TadA, in Complex with RNA | | Descriptor: | GLYCEROL, ZINC ION, anticodon stem-loop of t-RNA-Arg2 (nucleotides 27-42), ... | | Authors: | Losey, H.C, Ruthenburg, A.J, Verdine, G.L. | | Deposit date: | 2005-09-20 | | Release date: | 2006-01-24 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of Staphylococcus aureus tRNA adenosine deaminase TadA in complex with RNA.
Nat.Struct.Mol.Biol., 13, 2006
|
|
1JSO
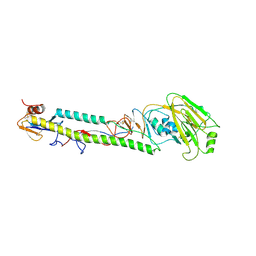 
 | | STRUCTURE OF AVIAN H5 HAEMAGGLUTININ BOUND TO LSTC RECEPTOR ANALOG | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, HAEMAGGLUTININ (HA1 CHAIN), ... | | Authors: | Ha, Y, Stevens, D.J, Skehel, J.J, Wiley, D.C. | | Deposit date: | 2001-08-17 | | Release date: | 2001-09-26 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | X-ray structures of H5 avian and H9 swine influenza virus hemagglutinins bound to avian and human receptor analogs.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
2VSP
 
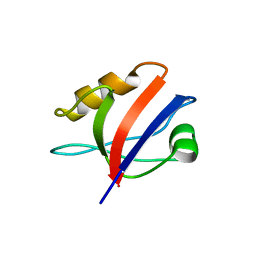 | | Crystal structure of the fourth PDZ domain of PDZ domain-containing protein 1 | | Descriptor: | PDZ DOMAIN-CONTAINING PROTEIN 1 | | Authors: | Yue, W.W, Shafqat, N, Pilka, E.S, Johansson, C, Murray, J.W, Elkins, J, Roos, A, Cooper, C, Phillips, C, Salah, E, von Delft, F, Doyle, D, Edwards, A, Wikstrom, M, Arrowsmith, C, Bountra, C, Oppermann, U. | | Deposit date: | 2008-04-28 | | Release date: | 2009-03-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Crystal Structure of the Fourth Pdz Domain of Pdz Domain-Containing Protein 1
To be Published
|
|
1V4K
 
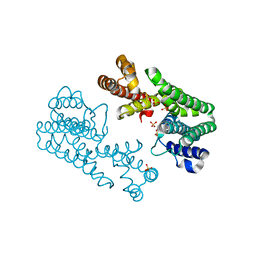 | | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima S77F mutant | | Descriptor: | SULFATE ION, octoprenyl-diphosphate synthase | | Authors: | Guo, R.T, Kuo, C.J, Chou, C.C, Ko, T.P, Shr, H.L, Liang, P.H, Wang, A.H.-J. | | Deposit date: | 2003-11-14 | | Release date: | 2004-03-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima and Mechanism of Product Chain Length Determination
J.Biol.Chem., 279, 2004
|
|
3DFT
 
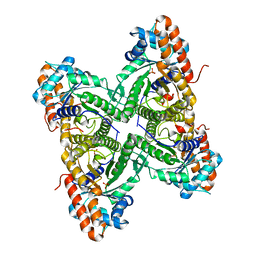 | |
1E48
 
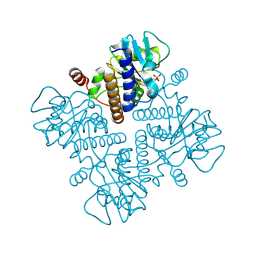 | |
2DKK
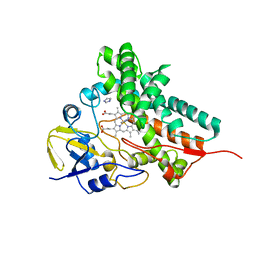 
 | |
1JPZ
 
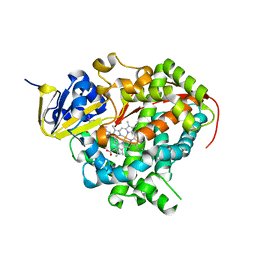 | | Crystal structure of a complex of the heme domain of P450BM-3 with N-Palmitoylglycine | | Descriptor: | BIFUNCTIONAL P-450:NADPH-P450 REDUCTASE, N-PALMITOYLGLYCINE, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Haines, D.C, Tomchick, D.R, Machius, M, Peterson, J.A. | | Deposit date: | 2001-08-03 | | Release date: | 2001-11-09 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Pivotal role of water in the mechanism of P450BM-3.
Biochemistry, 40, 2001
|
|
1E4B
 
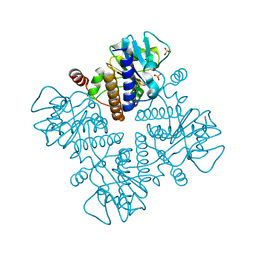 | |
2VSW
 
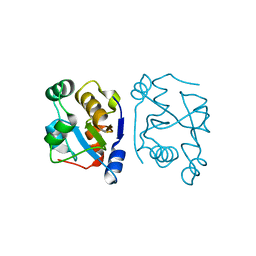 | | The structure of the rhodanese domain of the human dual specificity phosphatase 16 | | Descriptor: | DUAL SPECIFICITY PROTEIN PHOSPHATASE 16 | | Authors: | Murray, J.W, Barr, A, Pike, A.C.W, Elkins, J, Phillips, C, Wang, J, Savitsky, P, Roos, A, Bishop, S, Wickstroem, M, Bountra, C, Edwards, A.M, Arrowsmith, C.H, Burgess-Brown, N, Pantic, N, Bray, J, von Delft, F, Gileadi, O, Knapp, S. | | Deposit date: | 2008-04-30 | | Release date: | 2008-07-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Structure of the Rhodanese Domain of the Human Dual Specifity Phosphatase 16
To be Published
|
|
4KFE
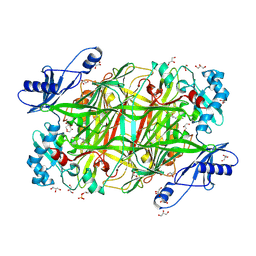 
 | | Crystal structure of Hansenula polymorpha copper amine oxidase-1 reduced by methylamine at pH 7.0 | | Descriptor: | COPPER (II) ION, FORMYL GROUP, GLYCEROL, ... | | Authors: | Johnson, B.J, Yukl, E.T, Klema, V.J, Wilmot, C.M. | | Deposit date: | 2013-04-26 | | Release date: | 2013-08-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural evidence for the semiquinone in a copper amine oxidase from Hansenula polymorpha: implications for the catalytic mechanism
J.Biol.Chem., 2013
|
|
2J2S
 
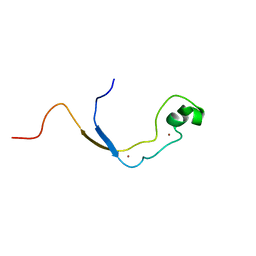 | | Solution structure of the nonmethyl-CpG-binding CXXC domain of the leukaemia-associated MLL histone methyltransferase | | Descriptor: | ZINC FINGER PROTEIN HRX, ZINC ION | | Authors: | Allen, M.D, Grummitt, C.G, Hilcenko, C, Young-Min, S, Tonkin, L.M, Johnson, C.M, Bycroft, M, Warren, A.J. | | Deposit date: | 2006-08-17 | | Release date: | 2006-08-21 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Nonmethyl-Cpg-Binding Cxxc Domain of the Leukaemia-Associated Mll Histone Methyltransferase
Embo J., 25, 2006
|
|
1PZ5
 
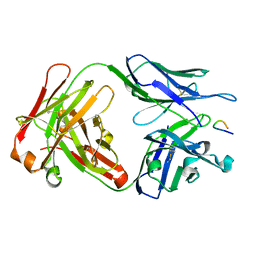 | | Structural basis of peptide-carbohydrate mimicry in an antibody combining site | | Descriptor: | Heavy chain of Fab (SYA/J6), Light chain of Fab (SYA/J6), Octapeptide (MDWNMHAA) | | Authors: | Vyas, N.K, Vyas, M.N, Chervenak, M.C, Bundle, D.R, Pinto, B.M, Quiocho, F.A. | | Deposit date: | 2003-07-09 | | Release date: | 2003-11-18 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis of peptide-carbohydrate mimicry in an antibody combining site.
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
2DUW
 
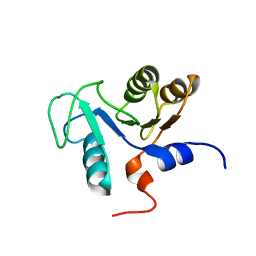 | | Solution structure of putative CoA-binding protein of Klebsiella pneumoniae | | Descriptor: | putative CoA-binding protein | | Authors: | Hung, K.W, Lin, Y.C, Cheng, C.C, Chang, C.F, Tsai, S.F, Huang, T.H. | | Deposit date: | 2006-07-27 | | Release date: | 2007-08-14 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of putative CoA-binding protein of Klebsiella pneumoniae
To be Published
|
|
4BZ9
 
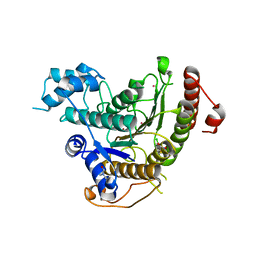 | | Crystal structure of Schistosoma mansoni HDAC8 complexed with J1075 | | Descriptor: | 3-chlorobenzothiophene-2-carbohydroxamic acid, DIMETHYLFORMAMIDE, GLYCEROL, ... | | Authors: | Marek, M, Romier, C. | | Deposit date: | 2013-07-24 | | Release date: | 2013-08-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for the Inhibition of Histone Deacetylase 8 (Hdac8), a Key Epigenetic Player in the Blood Fluke Schistosoma Mansoni.
Plos Pathog., 9, 2013
|
|
1TK9
 
 | | Crystal Structure of Phosphoheptose isomerase 1 | | Descriptor: | Phosphoheptose isomerase 1 | | Authors: | Rajashankar, K.R, Solorzano, V, Kniewel, R, Lima, C.D, Burley, S.K, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2004-06-08 | | Release date: | 2004-06-22 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structures of two putative phosphoheptose isomerases.
Proteins, 63, 2006
|
|
