2JAM
 
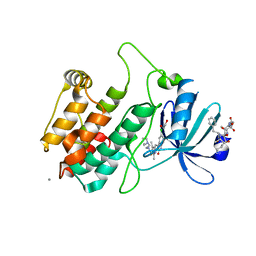 | | Crystal structure of human calmodulin-dependent protein kinase I G | | Descriptor: | 1,2-ETHANEDIOL, 5-[(E)-(5-CHLORO-2-OXO-1,2-DIHYDRO-3H-INDOL-3-YLIDENE)METHYL]-N-[2-(DIETHYLAMINO)ETHYL]-2,4-DIMETHYL-1H-PYRROLE-3-CARBOXAMIDE, CALCIUM ION, ... | | Authors: | Debreczeni, J.E, Bullock, A, Keates, T, Niesen, F.H, Salah, E, Shrestha, L, Smee, C, Sobott, F, Pike, A.C.W, Bunkoczi, G, von Delft, F, Turnbull, A, Weigelt, J, Arrowsmith, C.H, Edwards, A, Sundstrom, M, Knapp, S. | | Deposit date: | 2006-11-29 | | Release date: | 2007-03-13 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structure of Human Calmodulin-Dependent Protein Kinase I G
To be Published
|
|
2ZHR
 
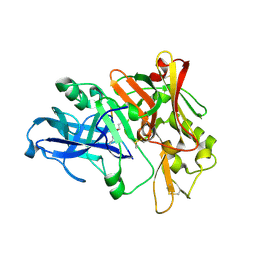 | |
2ZMM
 
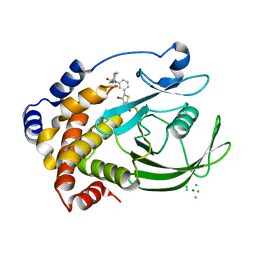 | | Crystal structure of PTP1B-inhibitor complex | | Descriptor: | 4-bromo-3-(carboxymethoxy)-5-{3-[cyclohexyl(methylcarbamoyl)amino]phenyl}thiophene-2-carboxylic acid, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Xu, W, Wu, J. | | Deposit date: | 2008-04-19 | | Release date: | 2008-10-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure-based optimization of protein tyrosine phosphatase-1 B inhibitors: capturing interactions with arginine 24
Chemmedchem, 3, 2008
|
|
2JG9
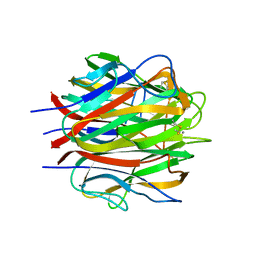 
 | | Crystallographic structure of human C1q globular heads (P1) | | Descriptor: | CALCIUM ION, Complement C1q subcomponent subunit A, Complement C1q subcomponent subunit B, ... | | Authors: | Paidassi, H, Tacnet-Delorme, P, Garlatti, V, Darnault, C, Ghebrehiwet, B, Gaboriaud, C, Arlaud, G.J, Frachet, P. | | Deposit date: | 2007-02-09 | | Release date: | 2008-02-19 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | C1Q Binds Phosphatidylserine and Likely Acts as a Multiligand-Bridging Molecule in Apoptotic Cell Recognition.
J.Immunol., 180, 2008
|
|
2ZIV
 
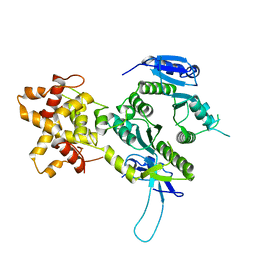 | | Crystal structure of the Mus81-Eme1 complex | | Descriptor: | Crossover junction endonuclease EME1, Mus81 protein | | Authors: | Chang, J.H, Kim, J.J, Choi, J.M, Lee, J.H, Cho, Y. | | Deposit date: | 2008-02-25 | | Release date: | 2008-04-29 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the Mus81-Eme1 complex
Genes Dev., 22, 2008
|
|
2ZN8
 
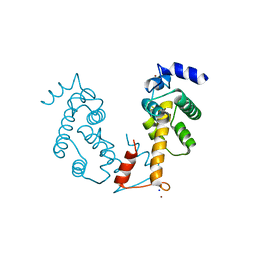 | | Crystal structure of Zn2+-bound form of ALG-2 | | Descriptor: | Programmed cell death protein 6, SODIUM ION, ZINC ION | | Authors: | Suzuki, H, Kawasaki, M, Inuzuka, T, Kakiuchi, T, Shibata, H, Wakatsuki, S, Maki, M. | | Deposit date: | 2008-04-22 | | Release date: | 2008-09-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural Basis for Ca(2+)-Dependent Formation of ALG-2/Alix Peptide Complex: Ca(2+)/EF3-Driven Arginine Switch Mechanism
Structure, 16, 2008
|
|
2ZND
 
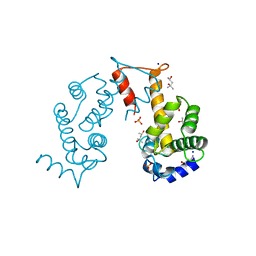 | | Crystal structure of Ca2+-free form of des3-20ALG-2 | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, PHOSPHATE ION, Programmed cell death protein 6, ... | | Authors: | Suzuki, H, Kawasaki, M, Inuzuka, T, Kakiuchi, T, Shibata, H, Wakatsuki, S, Maki, M. | | Deposit date: | 2008-04-22 | | Release date: | 2008-09-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Basis for Ca(2+)-Dependent Formation of ALG-2/Alix Peptide Complex: Ca(2+)/EF3-Driven Arginine Switch Mechanism
Structure, 16, 2008
|
|
2ZP0
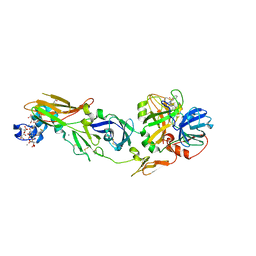 
 | | Human factor viia-tissue factor complexed with benzylsulfonamide-D-ile-gln-P-aminobenzamidine | | Descriptor: | (2S)-N-[(4-carbamimidoylphenyl)methyl]-2-[[(2R,3R)-3-methyl-2-(phenylmethylsulfonylamino)pentanoyl]amino]pentanediamide, CALCIUM ION, Factor VII heavy chain, ... | | Authors: | Kadono, S, Sakamoto, A, Kikuchi, Y, Oh-eda, M, Yabuta, N, Koga, T, Hattori, K, Shiraishi, T, Haramura, M, Kodama, H. | | Deposit date: | 2008-06-20 | | Release date: | 2008-07-15 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Peptide Mimetic Factor VIIa Inhibitor: Importance of Hydrophilic Pocket in S2 Site to Improve Selectivity aganist Thrombin
LETT.DRUG DES.DISCOVERY, 2, 2005
|
|
3B8Q
 
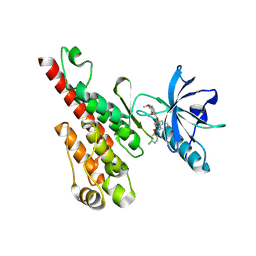 | | Crystal structure of the VEGFR2 kinase domain in complex with a naphthamide inhibitor | | Descriptor: | N-(4-chlorophenyl)-6-[(6,7-dimethoxyquinolin-4-yl)oxy]naphthalene-1-carboxamide, Vascular endothelial growth factor receptor 2 | | Authors: | Whittington, D.A, Long, A.M, Gu, Y, Zhao, H. | | Deposit date: | 2007-11-01 | | Release date: | 2008-04-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Naphthamides as Novel and Potent Vascular Endothelial Growth Factor
Receptor Tyrosine Kinase Inhibitors: Design, Synthesis and Evaluation
J.Med.Chem., 51, 2008
|
|
3B96
 
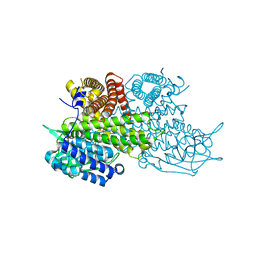 | | Structural Basis for Substrate Fatty-Acyl Chain Specificity: Crystal Structure of Human Very-Long-Chain Acyl-CoA Dehydrogenase | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, TETRADECANOYL-COA, Very long-chain specific acyl-CoA dehydrogenase | | Authors: | McAndrew, R.P, Wang, Y, Mohsen, A.W, He, M, Vockley, J, Kim, J.J. | | Deposit date: | 2007-11-02 | | Release date: | 2008-02-12 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Structural basis for substrate fatty acyl chain specificity: crystal structure of human very-long-chain acyl-CoA dehydrogenase.
J.Biol.Chem., 283, 2008
|
|
3BIA
 
 | | Tim-4 in complex with sodium potassium tartrate | | Descriptor: | L(+)-TARTARIC ACID, SODIUM ION, T-cell immunoglobulin and mucin domain-containing protein 4 | | Authors: | Santiago, C, Ballesteros, A, Kaplan, G.G, Freeman, G.J, Casasnovas, J.M. | | Deposit date: | 2007-11-30 | | Release date: | 2008-01-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structures of T Cell Immunoglobulin Mucin Protein 4 Show a Metal-Ion-Dependent Ligand Binding Site where Phosphatidylserine Binds.
Immunity, 27, 2007
|
|
3BAU
 
 | |
3BCJ
 
 | | Crystal structure of Aldose Reductase complexed with 2S4R (Stereoisomer of Fidarestat, 2S4S) at 0.78 A | | Descriptor: | (2S,4R)-2-AMINOFORMYL-6-FLUORO-SPIRO[CHROMAN-4,4'-IMIDAZOLIDINE]-2',5'-DIONE, Aldose reductase, CITRIC ACID, ... | | Authors: | Zhao, H.T, El-Kabbani, O. | | Deposit date: | 2007-11-13 | | Release date: | 2008-04-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (0.78 Å) | | Cite: | Unusual Binding Mode of the 2S4R Stereoisomer of the Potent Aldose Reductase Cyclic Imide Inhibitor Fidarestat (2S4S) in the 15 K Crystal Structure of the Ternary Complex Refined at 0.78 A Resolution: Implications for the Inhibition Mechanism
J.Med.Chem., 51, 2008
|
|
2W23
 
 | | Structure of mutant W169Y of Pleurotus eryngii versatile peroxidase (VP) | | Descriptor: | CALCIUM ION, GLYCEROL, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Ruiz-Duenas, F.J, Pogni, R, Morales, M, Giansanti, S, Mate, M.J, Romero, A, Martinez, M.J, Basosi, R, Martinez, A.T. | | Deposit date: | 2008-10-23 | | Release date: | 2009-01-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Protein Radicals in Fungal Versatile Peroxidase: Catalytic Tryptophan Radical in Both Compound I and Compound II and Studies on W164Y, W164H, and W164S Variants.
J.Biol.Chem., 284, 2009
|
|
3BEA
 
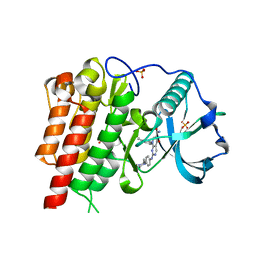 | | cFMS tyrosine kinase (tie2 KID) in complex with a pyrimidinopyridone inhibitor | | Descriptor: | 8-(2,3-dihydro-1H-inden-5-yl)-2-({4-[(3R,5S)-3,5-dimethylpiperazin-1-yl]phenyl}amino)-5-oxo-5,8-dihydropyrido[2,3-d]pyrimidine-6-carboxamide, Macrophage colony-stimulating factor 1 receptor, SULFATE ION | | Authors: | Schubert, C. | | Deposit date: | 2007-11-16 | | Release date: | 2008-07-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Design and synthesis of a pyrido[2,3-d]pyrimidin-5-one class of anti-inflammatory FMS inhibitors.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
3BEJ
 
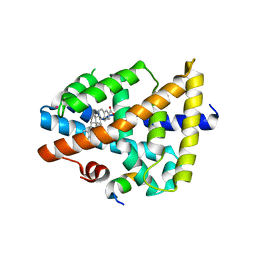 | | Structure of human FXR in complex with MFA-1 and co-activator peptide | | Descriptor: | (8alpha,10alpha,13alpha,17beta)-17-[(4-hydroxyphenyl)carbonyl]androsta-3,5-diene-3-carboxylic acid, Bile acid receptor, Nuclear receptor coactivator 1, ... | | Authors: | Soisson, S.M, Parthasarathy, G, Becker, J.W. | | Deposit date: | 2007-11-19 | | Release date: | 2008-03-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Identification of a potent synthetic FXR agonist with an unexpected mode of binding and activation.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
3J6R
 
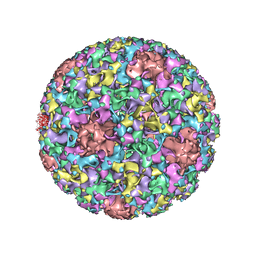 | | Electron cryo-microscopy of Human Papillomavirus Type 16 capsid | | Descriptor: | Major capsid protein L1 | | Authors: | Cardone, G, Moyer, A.L, Cheng, N, Thompson, C.D, Dvoretzky, I, Lowy, D.R, Schiller, J.T, Steven, A.C, Buck, C.B, Trus, B.L. | | Deposit date: | 2014-03-20 | | Release date: | 2014-07-23 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (9.1 Å) | | Cite: | Maturation of the human papillomavirus 16 capsid.
MBio, 5, 2014
|
|
3BEW
 
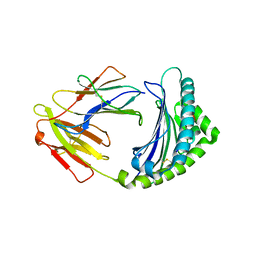 | | 10mer Crystal Structure of chicken MHC class I haplotype B21 | | Descriptor: | 10-mer from Tubulin beta-6 chain, Beta-2-microglobulin, Major histocompatibility complex class I glycoprotein haplotype B21 | | Authors: | Koch, M, Camp, S, Collen, T, Avila, D, Salomonsen, J, Wallny, H.J, van Hateren, A, Hunt, L, Jacob, J.P, Johnston, F, Marston, D.A, Shaw, I, Dunbar, P.R, Cerundolo, V, Jones, E.Y, Kaufman, J. | | Deposit date: | 2007-11-20 | | Release date: | 2008-01-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structures of an MHC class I molecule from b21 chickens illustrate promiscuous Peptide binding
Immunity, 27, 2007
|
|
3B2T
 
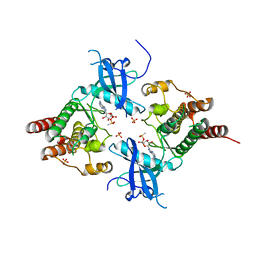 | | Structure of phosphotransferase | | Descriptor: | 5'-O-[(S)-hydroxy{[(S)-hydroxy(methyl)phosphoryl]oxy}phosphoryl]adenosine, Fibroblast growth factor receptor 2, PHOSPHATE ION | | Authors: | Lew, E.D, Bae, J.H, Rohmann, E, Wollnik, B, Schlessinger, J. | | Deposit date: | 2007-10-19 | | Release date: | 2008-02-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for reduced FGFR2 activity in LADD syndrome: Implications for FGFR autoinhibition and activation.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
2W3K
 
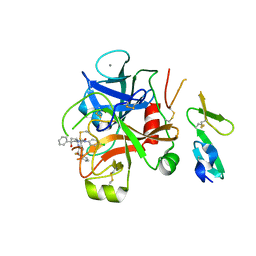 | | Crystal Structure of FXa in complex with 4,4-disubstituted pyrrolidine-1,2-dicarboxamide inhibitor 1 | | Descriptor: | (2R,4S)-N^1^-(4-chlorophenyl)-N^2^-[2-fluoro-4-(2-oxopyridin-1(2H)-yl)phenyl]-4-hydroxy-4-phenylpyrrolidine-1,2-dicarboxamide, CALCIUM ION, COAGULATION FACTOR X, ... | | Authors: | Zhang, E, Mochalkin, I, Casimiro-Garcia, A, Van Huis, C.A. | | Deposit date: | 2008-11-12 | | Release date: | 2009-04-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Exploration of 4,4-Disubstituted Pyrrolidine-1,2-Dicarboxamides as Potent, Orally Active Factor Xa Inhibitors with Extended Duration of Action.
Bioorg.Med.Chem., 17, 2009
|
|
3B3C
 
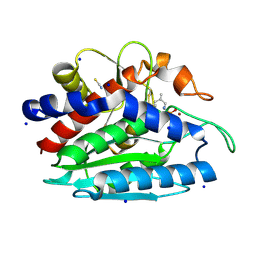 | | Crystal structure of the M180A mutant of the aminopeptidase from Vibrio proteolyticus in complex with leucine phosphonic acid | | Descriptor: | Bacterial leucyl aminopeptidase, LEUCINE PHOSPHONIC ACID, POTASSIUM ION, ... | | Authors: | Ataie, N.J, Hoang, Q.Q, Petsko, G.A, Ringe, D. | | Deposit date: | 2007-10-19 | | Release date: | 2007-11-27 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | Zinc coordination geometry and ligand binding affinity: the structural and kinetic analysis of the second-shell serine 228 residue and the methionine 180 residue of the aminopeptidase from Vibrio proteolyticus.
Biochemistry, 47, 2008
|
|
3B67
 
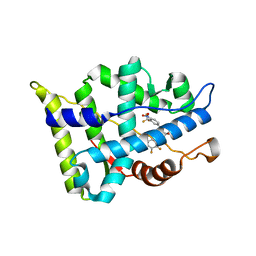 | |
3B3S
 
 | | Crystal structure of the M180A mutant of the aminopeptidase from Vibrio proteolyticus in complex with leucine | | Descriptor: | Bacterial leucyl aminopeptidase, LEUCINE, SODIUM ION, ... | | Authors: | Ataie, N.J, Hoang, Q.Q, Petsko, G.A, Ringe, D. | | Deposit date: | 2007-10-22 | | Release date: | 2007-11-27 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.18 Å) | | Cite: | Zinc coordination geometry and ligand binding affinity: the structural and kinetic analysis of the second-shell serine 228 residue and the methionine 180 residue of the aminopeptidase from Vibrio proteolyticus.
Biochemistry, 47, 2008
|
|
3B75
 
 | | Crystal Structure of Glycated Human Haemoglobin | | Descriptor: | Hemoglobin subunit alpha, Hemoglobin subunit beta, OXYGEN MOLECULE, ... | | Authors: | Saraswathi, N.T, Syakhovich, V.E, Bokut, S.B, Moras, D, Ruff, M. | | Deposit date: | 2007-10-30 | | Release date: | 2008-10-14 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The effect of hemoglobin glycosylation on diabete linked oxidative stress
To be Published
|
|
3B65
 
 | |
