1F24
 
 | |
1F25
 
 | |
1F26
 
 | |
1EHG
 
 | |
1JFB
 
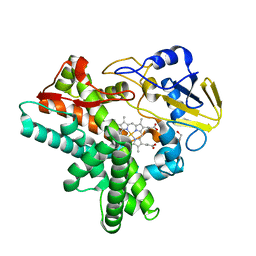 | | X-ray structure of nitric oxide reductase (cytochrome P450nor) in the ferric resting state at atomic resolution | | Descriptor: | GLYCEROL, PROTOPORPHYRIN IX CONTAINING FE, nitric-oxide reductase cytochrome P450 55A1 | | Authors: | Shimizu, H, Adachi, S, Park, S.Y, Shiro, Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2001-06-20 | | Release date: | 2001-12-20 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | X-ray structure of nitric oxide reductase (cytochrome P450nor) at atomic resolution.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1EHF
 
 | |
1EHE
 
 | |
1JFC
 
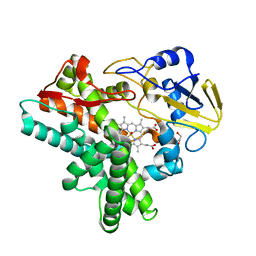 | | X-ray structure of nitric oxide reductase (cytochrome P450nor) in the ferrous CO state at atomic resolution | | Descriptor: | CARBON MONOXIDE, GLYCEROL, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Shimizu, H, Adachi, S, Park, S.Y, Shiro, Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2001-06-20 | | Release date: | 2001-12-20 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | X-ray structure of nitric oxide reductase (cytochrome P450nor) at atomic resolution.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1IQC
 
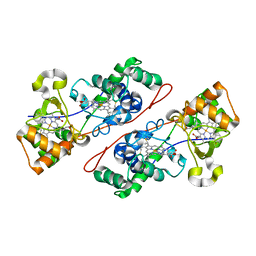 | | Crystal structure of Di-Heme Peroxidase from Nitrosomonas europaea | | Descriptor: | CALCIUM ION, GLYCEROL, HEME C, ... | | Authors: | Shimizu, H. | | Deposit date: | 2001-07-20 | | Release date: | 2002-01-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of Nitrosomonas europaea cytochrome c peroxidase and the structural basis for ligand switching in bacterial di-heme peroxidases
Biochemistry, 40, 2001
|
|
5AYQ
 
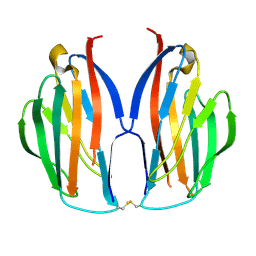 | |
5ZHX
 
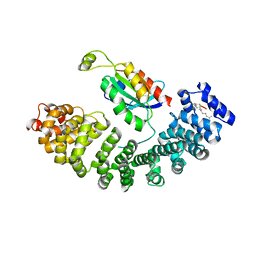 | |
5XGC
 
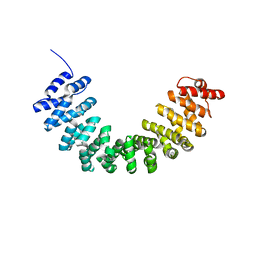 | | Crystal structure of SmgGDS-558 | | Descriptor: | Rap1 GTPase-GDP dissociation stimulator 1 | | Authors: | Shimizu, H, Toma-Fukai, S, Shimizu, T. | | Deposit date: | 2017-04-13 | | Release date: | 2017-06-28 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure-based analysis of the guanine nucleotide exchange factor SmgGDS reveals armadillo-repeat motifs and key regions for activity and GTPase binding
J. Biol. Chem., 292, 2017
|
|
6IZX
 
 | | The RNA-dependent RNA polymerase domain of dengue 2 NS5, bound with RK-0404678 | | Descriptor: | 2-oxo-2H-1,3-benzoxathiol-5-yl acetate, COBALT (II) ION, Genome polyprotein, ... | | Authors: | Shimizu, H, Sekine, S. | | Deposit date: | 2018-12-20 | | Release date: | 2019-12-25 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | Discovery of a small molecule inhibitor targeting dengue virus NS5 RNA-dependent RNA polymerase.
Plos Negl Trop Dis, 13, 2019
|
|
3ASX
 
 | | Human Squalene synthase in complex with 1-{4-[{4-chloro-2-[(2-chlorophenyl)(hydroxy)methyl]phenyl}(2,2-dimethylpropyl)amino]-4-oxobutanoyl}piperidine-3-carboxylic acid | | Descriptor: | (3R)-1-{4-[{4-chloro-2-[(S)-(2-chlorophenyl)(hydroxy)methyl]phenyl}(2,2-dimethylpropyl)amino]-4-oxobutanoyl}piperidine-3-carboxylic acid, PHOSPHATE ION, Squalene synthase | | Authors: | Shimizu, H, Suzuki, M, Katakura, S, Yamazaki, K, Higashihashi, N, Ichikawa, M, Yokomizo, A, Itoh, M, Sugita, K, Usui, H. | | Deposit date: | 2010-12-22 | | Release date: | 2011-12-21 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery of a new 2-aminobenzhydrol template for highly potent squalene synthase inhibitors
Bioorg.Med.Chem., 19, 2011
|
|
6IZZ
 
 | |
6J00
 
 | |
6IZY
 
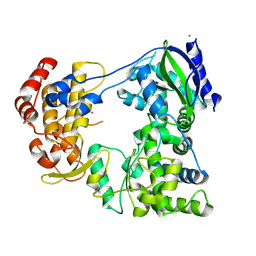 | |
2DDD
 
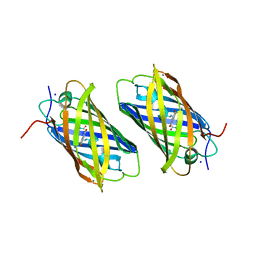 | | Unique behavior of a histidine responsible for an engineered green-to-red photoconversion process | | Descriptor: | MAGNESIUM ION, SODIUM ION, photoconvertible fluorescent protein | | Authors: | Shimizu, H, Tsutsui, H, Nukina, N, Miyawaki, A. | | Deposit date: | 2006-01-27 | | Release date: | 2006-03-07 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | The E1 mechanism in photo-induced beta-elimination reactions for green-to-red conversion of fluorescent proteins
Chem.Biol., 16, 2009
|
|
2DDC
 
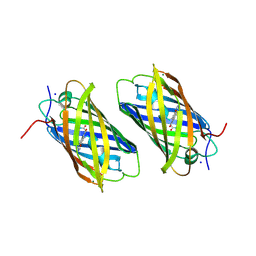 | | Unique behavior of a histidine responsible for an engineered green-to-red photoconversion process | | Descriptor: | MAGNESIUM ION, SODIUM ION, photoconvertible fluorescent protein | | Authors: | Shimizu, H, Tsutsui, H, Nukina, N, Miyawaki, A. | | Deposit date: | 2006-01-27 | | Release date: | 2006-03-07 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | The E1 mechanism in photo-induced beta-elimination reactions for green-to-red conversion of fluorescent proteins.
Chem.Biol., 16, 2009
|
|
5Z06
 
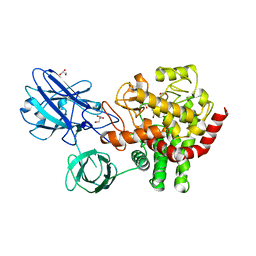 | | Crystal structure of beta-1,2-glucanase from Parabacteroides distasonis | | Descriptor: | BDI_3064 protein, CALCIUM ION, GLYCEROL | | Authors: | Shimizu, H, Nakajima, M, Miyanaga, A, Takahashi, Y, Tanaka, N, Kobayashi, K, Sugimoto, N, Nakai, H, Taguchi, H. | | Deposit date: | 2017-12-18 | | Release date: | 2018-05-30 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Characterization and Structural Analysis of a Novel exo-Type Enzyme Acting on beta-1,2-Glucooligosaccharides from Parabacteroides distasonis
Biochemistry, 57, 2018
|
|
2ZHR
 
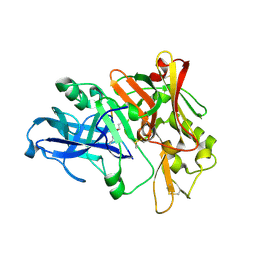 | |
5XAW
 
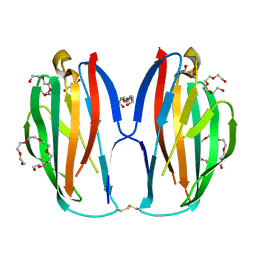 | | Parallel homodimer structures of voltage-gated sodium channel beta4 for cell-cell adhesion | | Descriptor: | 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, GLYCEROL, Sodium channel subunit beta-4, ... | | Authors: | Shimizu, H, Yokoyama, S. | | Deposit date: | 2017-03-15 | | Release date: | 2017-07-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.101 Å) | | Cite: | Parallel homodimer structures of the extracellular domains of the voltage-gated sodium channel beta 4 subunit explain its role in cell-cell adhesion
J. Biol. Chem., 292, 2017
|
|
5XAX
 
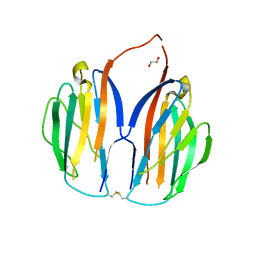 | |
2ZHU
 
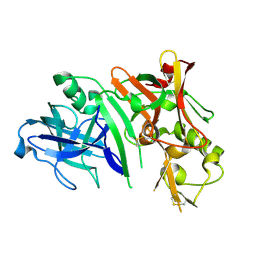 | | Crystal structure of BACE1 at pH 5.0 | | Descriptor: | Beta-secretase 1 | | Authors: | Shimizu, H, Nukina, N. | | Deposit date: | 2008-02-08 | | Release date: | 2008-04-22 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of an active form of BACE1, an enzyme responsible for amyloid beta protein production
Mol.Cell.Biol., 28, 2008
|
|
2ZHV
 
 | | Crystal structure of BACE1 at pH 7.0 | | Descriptor: | Beta-secretase 1 | | Authors: | Shimizu, H, Nukina, N. | | Deposit date: | 2008-02-08 | | Release date: | 2008-04-22 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structure of an active form of BACE1, an enzyme responsible for amyloid beta protein production
Mol.Cell.Biol., 28, 2008
|
|
