5R96
 
 | |
5R9O
 
 | |
5RA0
 
 | |
6EGC
 
 | |
7OZD
 
 | | FGFR1 kinase domain (residues 458-765) with mutations C488A, C584S in complex with 34. | | Descriptor: | 1,2-ETHANEDIOL, Fibroblast growth factor receptor 1, N-[6-(4-hydroxyphenyl)-1H-indazol-3-yl]benzamide, ... | | Authors: | Trinh, C.H, Turner, L.D, Fishwick, C.W.G. | | Deposit date: | 2021-06-27 | | Release date: | 2021-12-01 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | From Fragment to Lead: De Novo Design and Development toward a Selective FGFR2 Inhibitor.
J.Med.Chem., 65, 2022
|
|
7OZB
 
 | | FGFR1 kinase domain (residues 458-765) with mutations C488A, C584S in complex with 38. | | Descriptor: | 1,2-ETHANEDIOL, 4-[3-(4-piperazin-4-ium-1-ylphenyl)-1H-indazol-6-yl]phenol, Fibroblast growth factor receptor 1, ... | | Authors: | Trinh, C.H, Turner, L.D, Fishwick, C.W.G. | | Deposit date: | 2021-06-27 | | Release date: | 2021-12-01 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | From Fragment to Lead: De Novo Design and Development toward a Selective FGFR2 Inhibitor.
J.Med.Chem., 65, 2022
|
|
7OZF
 
 | | FGFR1 kinase domain (residues 458-765) with mutations C488A, C584S in complex with 19. | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Fibroblast growth factor receptor 1, ... | | Authors: | Trinh, C.H, Turner, L.D, Fishwick, C.W.G. | | Deposit date: | 2021-06-28 | | Release date: | 2021-12-01 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | From Fragment to Lead: De Novo Design and Development toward a Selective FGFR2 Inhibitor.
J.Med.Chem., 65, 2022
|
|
7OZY
 
 | | FGFR2 kinase domain (residues 461-763) in complex with 38. | | Descriptor: | 1,2-ETHANEDIOL, 4-[3-(4-piperazin-4-ium-1-ylphenyl)-1H-indazol-6-yl]phenol, Fibroblast growth factor receptor 2, ... | | Authors: | Trinh, C.H, Turner, L.D, Fishwick, C.W.G. | | Deposit date: | 2021-06-29 | | Release date: | 2021-12-01 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | From Fragment to Lead: De Novo Design and Development toward a Selective FGFR2 Inhibitor.
J.Med.Chem., 65, 2022
|
|
7RWV
 
 | | Crystal structure of a metal-free RIDC1 variant | | Descriptor: | HEME C, Soluble cytochrome b562 | | Authors: | Kakkis, A, Golub, E. | | Deposit date: | 2021-08-20 | | Release date: | 2022-06-29 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Redox- and metal-directed structural diversification in designed metalloprotein assemblies.
Chem.Commun.(Camb.), 58, 2022
|
|
7RWX
 
 | |
7RWW
 
 | | Crystal structure of a Zn-bound RIDC1 variant | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Kakkis, A, Golub, E. | | Deposit date: | 2021-08-20 | | Release date: | 2022-06-29 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Redox- and metal-directed structural diversification in designed metalloprotein assemblies.
Chem.Commun.(Camb.), 58, 2022
|
|
7RWU
 
 | |
7RWY
 
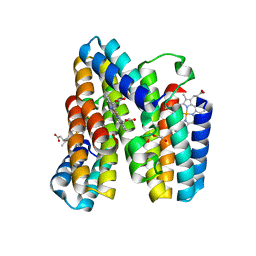 | |
4KVT
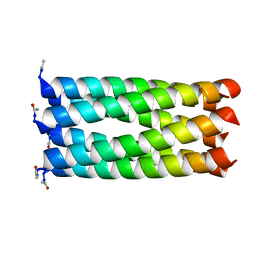 
 | | Crystal structure of a 6-helix coiled coil CC-Hex-L24C | | Descriptor: | 6-helix coiled coil CC-Hex-L24C peptide | | Authors: | Burton, A.J, Agnew, C, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2013-05-23 | | Release date: | 2013-08-21 | | Last modified: | 2013-09-11 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Accessibility, Reactivity, and Selectivity of Side Chains within a Channel of de Novo Peptide Assembly.
J.Am.Chem.Soc., 135, 2013
|
|
4KVV
 
 | | Crystal structure of an alkylated Cys mutant of CC-Hex | | Descriptor: | 6-HELIX COILED COIL CC-HEX-L24C PEPTIDE with an alkylated Cys mutation | | Authors: | Burton, A.J, Agnew, C, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2013-05-23 | | Release date: | 2013-08-21 | | Last modified: | 2013-09-11 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Accessibility, Reactivity, and Selectivity of Side Chains within a Channel of de Novo Peptide Assembly.
J.Am.Chem.Soc., 135, 2013
|
|
6YPI
 
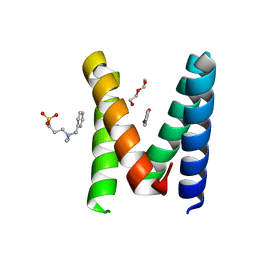 | | Structure of the engineered metallo-Diels-Alderase DA7 W16G,K58Q,L77R,T78R | | Descriptor: | 3-[BENZYL(DIMETHYL)AMMONIO]PROPANE-1-SULFONATE, BENZOIC ACID, DA7 W16G,K58Q,L77R,T78R, ... | | Authors: | Basler, S, Mori, T, Hilvert, D. | | Deposit date: | 2020-04-16 | | Release date: | 2021-04-28 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.479 Å) | | Cite: | Efficient Lewis acid catalysis of an abiological reaction in a de novo protein scaffold.
Nat.Chem., 13, 2021
|
|
6FL5
 
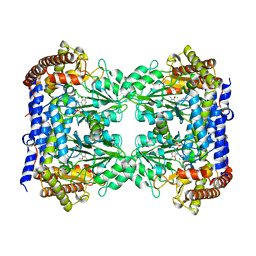 | | Structure of human SHMT1-H135N-R137A-E168N mutant at 3.6 Ang. resolution | | Descriptor: | CHLORIDE ION, PYRIDOXAL-5'-PHOSPHATE, Serine hydroxymethyltransferase, ... | | Authors: | Giardina, G, Cutruzzola, F, Lucchi, R. | | Deposit date: | 2018-01-25 | | Release date: | 2018-10-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | The catalytic activity of serine hydroxymethyltransferase is essential for de novo nuclear dTMP synthesis in lung cancer cells.
FEBS J., 285, 2018
|
|
5R87
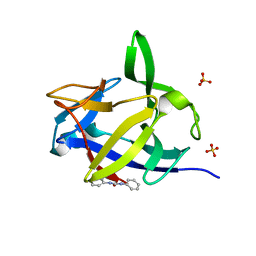 
 | |
5R8L
 
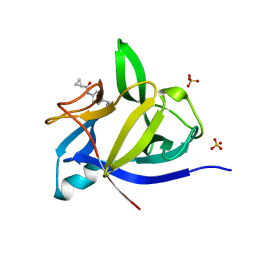 | |
5R8D
 
 | |
5R8J
 
 | |
5R8F
 
 | |
5R8A
 
 | | PanDDA analysis group deposition INTERLEUKIN-1 BETA -- Fragment Z1492796719 in complex with INTERLEUKIN-1 BETA | | Descriptor: | Interleukin-1 beta, SULFATE ION, ~{N}-[(3~{R})-1,2,3,4-tetrahydroquinolin-3-yl]ethanamide | | Authors: | De Nicola, G.F, Nichols, C.E. | | Deposit date: | 2020-03-03 | | Release date: | 2020-04-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | Mining the PDB for Tractable Cases Where X-ray Crystallography Combined with Fragment Screens Can Be Used to Systematically Design Protein-Protein Inhibitors: Two Test Cases Illustrated by IL1 beta-IL1R and p38 alpha-TAB1 Complexes.
J.Med.Chem., 63, 2020
|
|
5R8O
 
 | |
5R89
 
 | |
