2FXS
 
 | | Yeast HSP82 in complex with the novel HSP90 Inhibitor Radamide | | Descriptor: | ATP-dependent molecular chaperone HSP82, GLYCEROL, METHYL 3-CHLORO-2-{3-[(2,5-DIHYDROXY-4-METHOXYPHENYL)AMINO]-3-OXOPROPYL}-4,6-DIHYDROXYBENZOATE | | Authors: | Immormino, R.M, Gewirth, D.T. | | Deposit date: | 2006-02-06 | | Release date: | 2007-02-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Different poses for ligand and chaperone in inhibitor-bound Hsp90 and GRP94: implications for paralog-specific drug design.
J.Mol.Biol., 388, 2009
|
|
2FR1
 
 | |
2FYP
 
 | | GRP94 in complex with the novel HSP90 Inhibitor Radester amine | | Descriptor: | 2-(3-AMINO-2,5,6-TRIMETHOXYPHENYL)ETHYL 5-CHLORO-2,4-DIHYDROXYBENZOATE, Endoplasmin, PENTAETHYLENE GLYCOL, ... | | Authors: | Immormino, R.M, Gewirth, D.T. | | Deposit date: | 2006-02-08 | | Release date: | 2007-02-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Inhibittory Ligands Adopt Different Conformations When Bound to Hsp90 or GRP94: Implications for Paralog-specific Drug Design
To be Published
|
|
5DCH
 
 | | Crystal structure of Pseudomonas aeruginosa DsbA E82I in complex with MIPS-0000851 (3-[(2-METHYLBENZYL)SULFANYL]-4H-1,2,4-TRIAZOL-4-AMINE) | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 3-[(2-methylbenzyl)sulfanyl]-4H-1,2,4-triazol-4-amine, GLYCEROL, ... | | Authors: | McMahon, R.M, Martin, J.L. | | Deposit date: | 2015-08-24 | | Release date: | 2016-10-05 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.447 Å) | | Cite: | Fragment library screening identifies hits that bind to the non-catalytic surface of Pseudomonas aeruginosa DsbA1.
PLoS ONE, 12, 2017
|
|
3ZK7
 
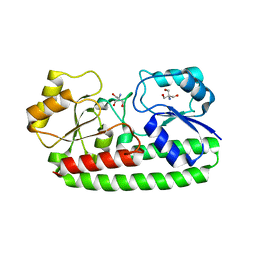 | | CRYSTAL STRUCTURE OF PNEUMOCOCCAL SURFACE ANTIGEN PSAA IN THE METAL-FREE, OPEN STATE | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, MANGANESE ABC TRANSPORTER SUBSTRATE-BINDING LIPOPROTEIN | | Authors: | Counago, R.M, Ween, M.P, Bajaj, M, Zuegg, J, Cooper, M.A, McEwan, A.G, Paton, J.C, Kobe, B, McDevitt, C.A. | | Deposit date: | 2013-01-22 | | Release date: | 2013-11-06 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Imperfect coordination chemistry facilitates metal ion release in the Psa permease.
Nat. Chem. Biol., 10, 2014
|
|
4LK3
 
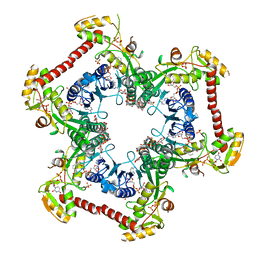 | | Crystal structure of Human UDP-xylose synthase R236A substitution | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, PYROPHOSPHATE 2-, SULFATE ION, ... | | Authors: | Walsh Jr, R.M, Polizzi, S.J, Wood, Z.A. | | Deposit date: | 2013-07-05 | | Release date: | 2013-08-07 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | Man o' war mutation in UDP-alpha-D-xylose synthase favors the abortive catalytic cycle and uncovers a latent potential for hexamer formation.
Biochemistry, 54, 2015
|
|
2F60
 
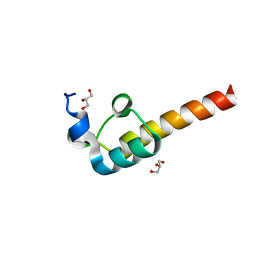 | | Crystal Structure of the Dihydrolipoamide Dehydrogenase (E3)-Binding Domain of Human E3-Binding Protein | | Descriptor: | GLYCEROL, Pyruvate dehydrogenase protein X component | | Authors: | Brautigam, C.A, Chuang, J.L, Wynn, R.M, Tomchick, D.R, Machius, M, Chuang, D.T. | | Deposit date: | 2005-11-28 | | Release date: | 2006-01-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural Insight into Interactions between Dihydrolipoamide Dehydrogenase (E3) and E3 Binding Protein of Human Pyruvate Dehydrogenase Complex.
Structure, 14, 2006
|
|
5EQG
 
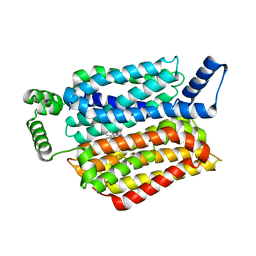 | | Human GLUT1 in complex with inhibitor (2~{S})-3-(4-fluorophenyl)-2-[2-(3-hydroxyphenyl)ethanoylamino]-~{N}-[(1~{S})-1-phenylethyl]propanamide | | Descriptor: | (2~{S})-3-(4-fluorophenyl)-2-[2-(3-hydroxyphenyl)ethanoylamino]-~{N}-[(1~{S})-1-phenylethyl]propanamide, Solute carrier family 2, facilitated glucose transporter member 1 | | Authors: | Kapoor, K, Finer-Moore, J, Pedersen, B.P, Caboni, L, Waight, A.B, Hillig, R, Bringmann, P, Heisler, I, Muller, T, Siebeneicher, H, Stroud, R.M. | | Deposit date: | 2015-11-12 | | Release date: | 2016-04-13 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Mechanism of inhibition of human glucose transporter GLUT1 is conserved between cytochalasin B and phenylalanine amides.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
4A07
 
 | | Human PDK1 Kinase Domain in Complex with Allosteric Activator PS171 Bound to the PIF-Pocket | | Descriptor: | (3S)-3-(4-CHLOROPHENYL)-4-(5,7-DICHLORO-1H-BENZIMIDAZOL-2-YL)BUTANOIC ACID, 3-PHOSPHOINOSITIDE-DEPENDENT PROTEIN KINASE 1, ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Schulze, J.O, Lopez-Garcia, L.A, Froehner, W, Zhang, H, Navratil, J, Hindie, V, Zeuzem, S, Alzari, P.M, Neimanis, S, Engel, M, Biondi, R.M. | | Deposit date: | 2011-09-08 | | Release date: | 2011-12-07 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Allosteric Regulation of Protein Kinase Pkczeta by the N-Terminal C1 Domain and Small Compounds to the Pif-Pocket.
Chem.Biol., 18, 2011
|
|
4A06
 
 | | Human PDK1 Kinase Domain in Complex with Allosteric Activator PS114 Bound to the PIF-Pocket | | Descriptor: | (3S)-4-(5-chloro-1H-benzimidazol-2-yl)-3-(4-chlorophenyl)butanoic acid, 3-PHOSPHOINOSITIDE-DEPENDENT PROTEIN KINASE 1, ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Schulze, J.O, Lopez-Garcia, L.A, Froehner, W, Zhang, H, Navratil, J, Hindie, V, Zeuzem, S, Alzari, P.M, Neimanis, S, Engel, M, Biondi, R.M. | | Deposit date: | 2011-09-08 | | Release date: | 2011-12-07 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Allosteric Regulation of Protein Kinase Pkczeta by the N-Terminal C1 Domain and Small Compounds to the Pif-Pocket.
Chem.Biol., 18, 2011
|
|
2EXL
 
 | | GRP94 N-terminal Domain bound to geldanamycin | | Descriptor: | Endoplasmin, GELDANAMYCIN, PENTAETHYLENE GLYCOL, ... | | Authors: | Reardon, P.N, Immormino, R.M, Gewirth, D.T. | | Deposit date: | 2005-11-08 | | Release date: | 2006-10-24 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Different poses for ligand and chaperone in inhibitor-bound Hsp90 and GRP94: implications for paralog-specific drug design.
J.Mol.Biol., 388, 2009
|
|
3CKE
 
 | | Crystal structure of aristolochene synthase in complex with 12,13-difluorofarnesyl diphosphate | | Descriptor: | (2E,6E)-12-fluoro-11-(fluoromethyl)-3,7-dimethyldodeca-2,6,10-trien-1-yl trihydrogen diphosphate, Aristolochene synthase, BETA-MERCAPTOETHANOL, ... | | Authors: | Shishova, E.Y, Yu, F, Miller, D.J, Faraldos, J.A, Zhao, Y, Coates, R.M, Allemann, R.K, Cane, D.E, Christianson, D.W. | | Deposit date: | 2008-03-14 | | Release date: | 2008-04-01 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | X-ray Crystallographic Studies of Substrate Binding to Aristolochene Synthase Suggest a Metal Ion Binding Sequence for Catalysis.
J.Biol.Chem., 283, 2008
|
|
5DUO
 
 | | Crystal structure of native translocator protein 18kDa (TSPO) from Rhodobacter sphaeroides (A139T Mutant) in C2 space group | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, FORMIC ACID, PROTOPORPHYRIN IX, ... | | Authors: | Li, F, Liu, J, Zheng, Y, Garavito, R.M, Ferguson-Miller, S. | | Deposit date: | 2015-09-20 | | Release date: | 2015-11-11 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Response to Comment on "Crystal structures of translocator protein (TSPO) and mutant mimic of a human polymorphism".
Science, 350, 2015
|
|
2EVQ
 | |
5E53
 
 | | Crystal structure of chicken CNTN1 FN1-FN3 domains | | Descriptor: | Contactin-1, IMIDAZOLE | | Authors: | Nikolaienko, R.M, Bouyain, S. | | Deposit date: | 2015-10-07 | | Release date: | 2016-08-31 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.497 Å) | | Cite: | Structural Basis for Interactions Between Contactin Family Members and Protein-tyrosine Phosphatase Receptor Type G in Neural Tissues.
J.Biol.Chem., 291, 2016
|
|
5E5R
 
 | |
1PXZ
 
 | | 1.7 Angstrom Crystal Structure of jun a 1, the major allergen from cedar pollen | | Descriptor: | Major pollen allergen Jun a 1 | | Authors: | Czerwinski, E.W, White, M.A, Midoro-Horiuti, T, Brooks, E.G, Goldblum, R.M. | | Deposit date: | 2003-07-07 | | Release date: | 2004-11-16 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of Jun a 1, the major cedar pollen allergen from Juniperus ashei, reveals a parallel beta-helical core.
J.Biol.Chem., 280, 2005
|
|
2ESA
 
 | |
2F5Z
 
 | | Crystal Structure of Human Dihydrolipoamide Dehydrogenase (E3) Complexed to the E3-Binding Domain of Human E3-Binding Protein | | Descriptor: | Dihydrolipoyl dehydrogenase, FLAVIN-ADENINE DINUCLEOTIDE, Pyruvate dehydrogenase protein X component, ... | | Authors: | Brautigam, C.A, Chuang, J.L, Wynn, R.M, Tomchick, D.R, Machius, M, Chuang, D.T. | | Deposit date: | 2005-11-28 | | Release date: | 2006-01-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Structural Insight into Interactions between Dihydrolipoamide Dehydrogenase (E3) and E3 Binding Protein of Human Pyruvate Dehydrogenase Complex.
Structure, 14, 2006
|
|
5E01
 
 | | Crystal structure of HiNmlR, a MerR family regulator lacking the sensor domain, bound to palyndromic promoter DNA | | Descriptor: | 5'-D(*CP*TP*TP*AP*GP*AP*GP*TP*GP*CP*AP*CP*TP*CP*TP*AP*AP*G)-3', Uncharacterized HTH-type transcriptional regulator HI_0186 | | Authors: | Counago, R.M, Chang, C.W, Chen, N.H, Djoko, K.Y, McEwan, A.G, Kobe, B. | | Deposit date: | 2015-09-26 | | Release date: | 2016-06-29 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis of thiol-based regulation of formaldehyde detoxification in H. influenzae by a MerR regulator with no sensor region.
Nucleic Acids Res., 44, 2016
|
|
5GIM
 
 | | Crystal structure of thrombin-avathrin complex | | Descriptor: | C-terminal peptide from Putative uncharacterized protein avahiru, N-terminal peptide from Putative uncharacterized protein avahiru, Thrombin light chain, ... | | Authors: | Kini, R.M, Koh, C.Y, Iyer, J.K, Swaminathan, K. | | Deposit date: | 2016-06-24 | | Release date: | 2017-05-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Avathrin: a novel thrombin inhibitor derived from a multicopy precursor in the salivary glands of the ixodid tick, Amblyomma variegatum.
FASEB J., 31, 2017
|
|
4JUL
 | | Crystal structure of H5N1 influenza virus hemagglutinin, clade 2.3.4 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin HA1, ... | | Authors: | DuBois, R.M, Zaraket, H, Reddivari, M, Coop, T, Heath, R.J, White, S.W, Russell, C.J. | | Deposit date: | 2013-03-25 | | Release date: | 2014-05-07 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.7927 Å) | | Cite: | To be published
To be Published
|
|
2GF7
 
 | | Double tudor domain structure | | Descriptor: | Jumonji domain-containing protein 2A, SULFATE ION | | Authors: | Huang, Y, Fang, J, Bedford, M.T, Zhang, Y, Xu, R.M. | | Deposit date: | 2006-03-21 | | Release date: | 2006-05-02 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Recognition of histone H3 lysine-4 methylation by the double tudor domain of JMJD2A
Science, 312, 2006
|
|
4J6E
 | | Structure of LPXI D225A Mutant | | Descriptor: | (2R,3R,4R,5S,6R)-2-{[(S)-{[(S)-{[(2R,3S,4R,5R)-5-(2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)-3,4-dihydroxytetrahydrofuran-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}-5-hydroxy-6-(hydroxymethyl)-3-{[(3R)-3-hydroxytetradecanoyl]amino}tetrahydro-2H-pyran-4-yl (3R)-3-hydroxytetradecanoate, UDP-2,3-diacylglucosamine pyrophosphatase LpxI | | Authors: | Metzger IV, L.E, Lee, J.K, Finer-Moore, J.S, Raetz, C.R.H, Stroud, R.M, Center for Structures of Membrane Proteins (CSMP) | | Deposit date: | 2013-02-11 | | Release date: | 2013-05-08 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.52 Å) | | Cite: | LpxI structures reveal how a lipid A precursor is synthesized.
Nat.Struct.Mol.Biol., 19, 2012
|
|
2FWZ
 | |
