2LSX
 
 | | Solution structure of a mini i-motif | | Descriptor: | DNA (5'-D(P*TP*CP*GP*TP*TP*TP*CP*GP*TP*T)-3') | | Authors: | Escaja, N, Viladoms, J, Garavis, M, Villasante, A, Pedroso, E, Gonzalez, C. | | Deposit date: | 2012-05-09 | | Release date: | 2012-10-17 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | A minimal i-motif stabilized by minor groove G:T:G:T tetrads.
Nucleic Acids Res., 40, 2012
|
|
2LA3
 
 | | The NMR structure of the protein NP_344798.1 reveals a CCA-adding enzyme head domain | | Descriptor: | Uncharacterized protein | | Authors: | Mohanty, B, Serrano, P, Geralt, M, Horst, R, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2011-03-01 | | Release date: | 2011-03-30 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the protein NP_344798.1
To be Published
|
|
2LF0
 
 | | Solution structure of sf3636, a two-domain unknown function protein from Shigella flexneri 2a, determined by joint refinement of NMR, residual dipolar couplings and small-angle X-ray scattering, NESG target SfR339/OCSP target sf3636 | | Descriptor: | Uncharacterized protein yibL | | Authors: | Wu, B, Lemak, A, Yee, A, Lee, H, Gutmanas, A, Semesi, A, Garcia, M, Fang, X, Wang, Y, Prestegard, J.H, Arrowsmith, C.H, Northeast Structural Genomics Consortium (NESG), Ontario Centre for Structural Proteomics (OCSP) | | Deposit date: | 2011-06-27 | | Release date: | 2011-07-13 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR, SOLUTION SCATTERING | | Cite: | Solution structure of sf3636, a two-domain unknown function protein from Shigella flexneri 2a, determined by joint refinement of NMR, residual dipolar couplings and small-angle X-ray scattering
To be Published
|
|
2LFO
 
 | | NMR structure of cl-BABP/SS complexed with glycochenodeoxycholic and glycocholic acids | | Descriptor: | Fatty acid-binding protein, liver, GLYCOCHENODEOXYCHOLIC ACID, ... | | Authors: | Tomaselli, S, Cogliati, C, Pagano, K, Zetta, L, Zanzoni, S, Assfalg, M, Molinari, H, Ragona, L. | | Deposit date: | 2011-07-07 | | Release date: | 2012-07-11 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | A disulfide bridge allows for site-selective binding in liver bile acid binding protein thereby stabilising the orientation of key amino acid side chains.
Chemistry, 18, 2012
|
|
2LFP
 
 | | Structure of bacteriophage SPP1 gp17 protein | | Descriptor: | Bacteriophage SPP1 complete nucleotide sequence | | Authors: | Chagot, B, Auzat, I, Gallopin, M, Petitpas, I, Gilquin, B, Tavares, P, Zinn-Justin, S. | | Deposit date: | 2011-07-07 | | Release date: | 2011-09-28 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure of gp17 from the Siphoviridae bacteriophage SPP1: Insights into its role in virion assembly.
Proteins, 80, 2012
|
|
2L6Y
 
 | | haddock model of GATA1NF:Lmo2LIM2-Ldb1LID | | Descriptor: | Erythroid transcription factor, LIM domain only 2, linker, ... | | Authors: | Wilkinson-White, L, Gamsjaeger, R, Dastmalchi, S, Wienert, B, Stokes, P.H, Crossley, M, Mackay, J.P, Matthews, J.M. | | Deposit date: | 2010-12-01 | | Release date: | 2011-08-31 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural basis of simultaneous recruitment of the transcriptional regulators LMO2 and FOG1/ZFPM1 by the transcription factor GATA1
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
2LMS
 | | A single GalNAc residue on Threonine-106 modifies the dynamics and the structure of Interferon alpha-2a around the glycosylation site | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-galactopyranose, Interferon alpha-2 | | Authors: | Ghasriani, H, Belcourt, P.J.F, Sauve, S, Hodgson, D.J, Gingras, G, Brochu, D, Gilbert, M, Aubin, Y. | | Deposit date: | 2011-12-12 | | Release date: | 2012-12-05 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | A single N-acetylgalactosamine residue at threonine 106 modifies the dynamics and structure of interferon alpha2a around the glycosylation site.
J.Biol.Chem., 288, 2013
|
|
2LHN
 
 | | RNA-binding zinc finger protein | | Descriptor: | Nuclear polyadenylated RNA-binding protein NAB2, ZINC ION | | Authors: | Brockmann, C, Neuhaus, D, Stewart, M. | | Deposit date: | 2011-08-12 | | Release date: | 2012-06-27 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural Basis for Polyadenosine-RNA Binding by Nab2 Zn Fingers and Its Function in mRNA Nuclear Export.
Structure, 20, 2012
|
|
2LJQ
 
 | | (C9S, C14S)-leucocin A | | Descriptor: | Bacteriocin leucocin-A | | Authors: | Sit, C.S, Lohans, C.T, van Belkum, M.J, Campbell, C.D, Miskolzie, M, Vederas, J.C. | | Deposit date: | 2011-09-22 | | Release date: | 2012-01-18 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Substitution of a Conserved Disulfide in the Type IIa Bacteriocin, Leucocin A, with L-Leucine and L-Serine Residues: Effects on Activity and Three-Dimensional Structure.
Chembiochem, 13, 2012
|
|
2LNL
 
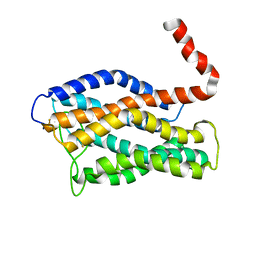 | | Structure of human CXCR1 in phospholipid bilayers | | Descriptor: | C-X-C chemokine receptor type 1 | | Authors: | Park, S, Das, B.B, Casagrande, F, Nothnagel, H, Chu, M, Kiefer, H, Maier, K, De Angelis, A, Marassi, F.M, Opella, S.J. | | Deposit date: | 2011-12-31 | | Release date: | 2012-10-17 | | Last modified: | 2016-04-27 | | Method: | SOLID-STATE NMR | | Cite: | Structure of the chemokine receptor CXCR1 in phospholipid bilayers.
Nature, 491, 2012
|
|
2LOD
 
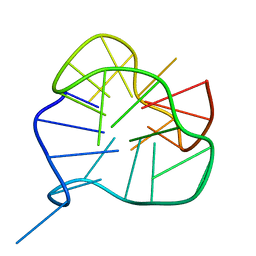 | | Solution-state structure of an intramolecular G-quadruplex with propeller, diagonal and edgewise loops | | Descriptor: | DNA (5'-D(*GP*GP*GP*AP*TP*GP*GP*GP*AP*CP*AP*CP*AP*GP*GP*GP*GP*AP*CP*GP*GP*G)-3') | | Authors: | Marusic, M, Sket, P, Bauer, L, Viglasky, V, Plavec, J. | | Deposit date: | 2012-01-23 | | Release date: | 2012-05-30 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution-state structure of an intramolecular G-quadruplex with propeller, diagonal and edgewise loops.
Nucleic Acids Res., 40, 2012
|
|
2L38
 | | R29Q Sticholysin II mutant | | Descriptor: | Sticholysin-2 | | Authors: | Castrillo, I, Alegre-Cebollada, J, Martinez-del-Pozo, A, Gavilanes, J, Bruix, M. | | Deposit date: | 2010-09-10 | | Release date: | 2010-09-22 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of StnIIR29Q, a defective lipid binding mutant of the sea anemone actinoporin Sticholysin II
To be Published
|
|
2L3N
 
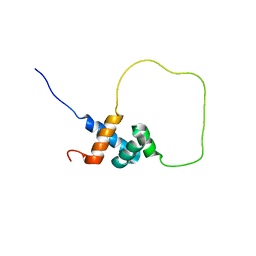 | | Solution structure of Rap1-Taz1 fusion protein | | Descriptor: | DNA-binding protein rap1,Telomere length regulator taz1 | | Authors: | Zhou, Z.R, Wang, F, Chen, Y, Lei, M, Hu, H. | | Deposit date: | 2010-09-19 | | Release date: | 2011-01-12 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A conserved motif within RAP1 has diversified roles in telomere protection and regulation in different organisms.
Nat.Struct.Mol.Biol., 18, 2011
|
|
2L4X
 
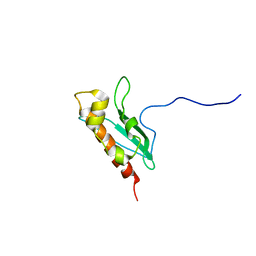 | | Solution Structure of apo-IscU(WT) | | Descriptor: | Iron-sulfur cluster assembly scaffold protein | | Authors: | Kim, J.H, Tonelli, M, Markley, J.L. | | Deposit date: | 2010-10-19 | | Release date: | 2011-12-07 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Three-Dimensional Structure and Determinants of Stability of the Iron-Sulfur Cluster Scaffold Protein IscU from Escherichia coli.
Biochemistry, 51, 2012
|
|
2L41
 
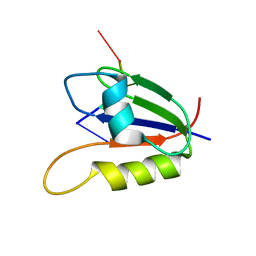 | | Nab3 RRM - UCUU complex | | Descriptor: | RNA (5'-R(P*UP*CP*UP*U)-3'), RRM domain from Nuclear polyadenylated RNA-binding protein 3 | | Authors: | Stefl, R, Pergoli, R, Hobor, F, Kubicek, K, Zimmermann, M, Pasulka, J, Hofr, C. | | Deposit date: | 2010-09-28 | | Release date: | 2010-11-17 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Recognition of transcription termination signal by the nuclear polyadenylated RNA-binding (NAB) 3 protein
J.Biol.Chem., 286, 2011
|
|
2L8M
 
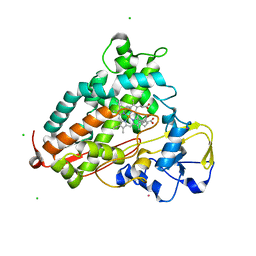 | | Reduced and CO-bound cytochrome P450cam (CYP101A1) | | Descriptor: | CAMPHOR, CARBON MONOXIDE, CHLORIDE ION, ... | | Authors: | Pochapsky, T.C, Pochapsky, S.S, Dang, M, Asciutto, E, Madura, J. | | Deposit date: | 2011-01-19 | | Release date: | 2011-02-16 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Experimentally Restrained Molecular Dynamics Simulations for Characterizing the Open States of Cytochrome P450(cam).
Biochemistry, 50, 2011
|
|
2LBH
 
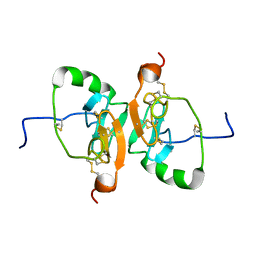 | | Solution Structure of the Dimeric Form of a Unliganded Bovine Neurophysin, Minimized Average Structure | | Descriptor: | Neurophysin 1 | | Authors: | Lee, H, Naik, M, Nguyen, T, Bracken, C, Breslow, E. | | Deposit date: | 2011-03-31 | | Release date: | 2012-04-04 | | Method: | SOLUTION NMR | | Cite: | Structural Basis of the Dimerization-Induced Increase in Neurophysin-Hormone Affinity: Interplay of Inter-Domain and Inter-Subunit Interactions
To be Published
|
|
2LHW
 
 | | Tri-O-GalNAc glycosylated Mucin sequence based on MUC2 Mucin glycoprotein tandem repeat | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-galactopyranose, MUC2 Mucin Domain Peptide | | Authors: | Borgert, A, Heimburg-Molinaro, J, Lasanajak, Y, Ju, T, Liu, M, Thompson, P, Ragupathi, G, Barany, G, Cummings, R, Smith, D, Live, D. | | Deposit date: | 2011-08-18 | | Release date: | 2012-04-04 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Deciphering structural elements of mucin glycoprotein recognition.
Acs Chem.Biol., 7, 2012
|
|
2LI2
 | | Mono-O-GalNAc glycosylated Mucin sequence based on MUC2 Mucin glycoprotein tandem repeat | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-galactopyranose, MUC2 Mucin Domain Peptide | | Authors: | Borgert, A, Heimburg-Molinaro, J, Lasanajak, Y, Ju, T, Liu, M, Thompson, P, Ragupathi, G, Barany, G, Cummings, R, Smith, D, Live, D. | | Deposit date: | 2011-08-18 | | Release date: | 2012-04-04 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Deciphering structural elements of mucin glycoprotein recognition.
Acs Chem.Biol., 7, 2012
|
|
2LQI
 
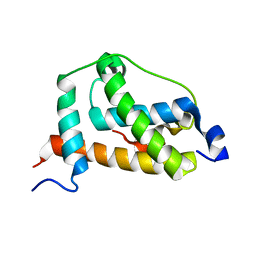 | | NMR structure of FOXO3a transactivation domains (CR2C-CR3) in complex with CBP KIX domain (2l3b conformation) | | Descriptor: | CREB-binding protein, Forkhead box O3 | | Authors: | Wang, F, Marshall, C.B, Yamamoto, K, Li, G.B, Gasmi-Seabrook, G.M.C, Okada, H, Mak, T.W, Ikura, M. | | Deposit date: | 2012-03-06 | | Release date: | 2012-05-16 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structures of KIX domain of CBP in complex with two FOXO3a transactivation domains reveal promiscuity and plasticity in coactivator recruitment.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
2LR4
 
 | |
2LRN
 
 | | Solution structure of a thiol:disulfide interchange protein from Bacteroides sp. | | Descriptor: | Thiol:disulfide interchange protein | | Authors: | Harris, R, Bandaranayake, A.D, Banu, R, Bonanno, J.B, Calarese, D.A, Celikgil, A, Chamala, S, Chan, M.K, Chaparro, R, Evans, B, Garforth, S, Gizzi, A, Hillerich, B, Kar, A, Lafleur, J, Lim, S, Love, J, Matikainen, B, Patel, H, Seidel, R.D, Smith, B, Stead, M, Girvin, M.E, Almo, S.C, New York Structural Genomics Research Consortium (NYSGRC) | | Deposit date: | 2012-04-10 | | Release date: | 2012-04-25 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure of a thiol:disulfide interchange protein from Bacteroides sp.
To be Published
|
|
2LST
 
 | | Solution structure of a thioredoxin from Thermus thermophilus | | Descriptor: | Thioredoxin | | Authors: | Harris, R, Bandaranayake, A.D, Banu, R, Bonanno, J.B, Calarese, D.A, Celikgil, A, Chamala, S, Chan, M.K, Chaparro, R, Evans, B, Garforth, S, Gizzi, A, Hillerich, B, Kar, A, Lafleur, J, Lim, S, Love, J, Matikainen, B, Patel, H, Seidel, R.D, Smith, B, Stead, M, Girvin, M.E, Almo, S.C, New York Structural Genomics Research Consortium (NYSGRC) | | Deposit date: | 2012-05-04 | | Release date: | 2012-05-16 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure of a thioredoxin from Thermus thermophilus
To be Published
|
|
2LM8
 | |
2L0W
 
 | | Solution NMR structure of the N-terminal PAS domain of HERG potassium channel | | Descriptor: | Potassium voltage-gated channel, subfamily H (Eag-related), member 2, ... | | Authors: | Ng, C.A, Hunter, M.J, Mobli, M, King, G.F, Kuchel, P.W, Vandenberg, J.I. | | Deposit date: | 2010-07-19 | | Release date: | 2011-01-26 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The N-Terminal Tail of hERG Contains an Amphipathic alpha-Helix That Regulates Channel Deactivation
PLoS ONE, 6, 2011
|
|
