2BAY
 
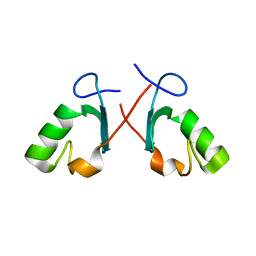 | | Crystal structure of the Prp19 U-box dimer | | Descriptor: | Pre-mRNA splicing factor PRP19 | | Authors: | Vander Kooi, C.W, Ohi, M.D, Rosenberg, J.A, Oldham, M.L, Newcomer, M.E, Gould, K.L, Chazin, W.J. | | Deposit date: | 2005-10-15 | | Release date: | 2006-01-10 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The Prp19 U-box Crystal Structure Suggests a Common Dimeric Architecture for a Class of Oligomeric E3 Ubiquitin Ligases.
Biochemistry, 45, 2006
|
|
3LRV
 
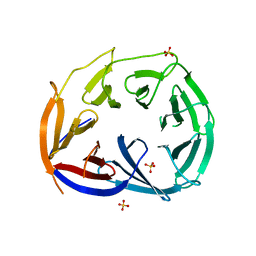 | |
3NME
 
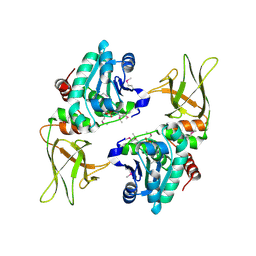 | | Structure of a plant phosphatase | | Descriptor: | PHOSPHATE ION, SEX4 glucan phosphatase | | Authors: | Vander Kooi, C.W. | | Deposit date: | 2010-06-22 | | Release date: | 2010-08-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for the glucan phosphatase activity of Starch Excess4.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
2ORZ
 
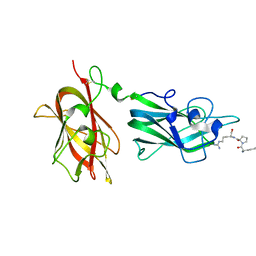 | | Structural Basis for Ligand Binding and Heparin Mediated Activation of Neuropilin | | Descriptor: | Neuropilin-1, Tuftsin | | Authors: | Vander Kooi, C.W, Jusino, M.A, Perman, B, Neau, D.B, Bellamy, H.D, Leahy, D.J. | | Deposit date: | 2007-02-05 | | Release date: | 2007-04-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structural basis for ligand and heparin binding to neuropilin B domains.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
2ORX
 
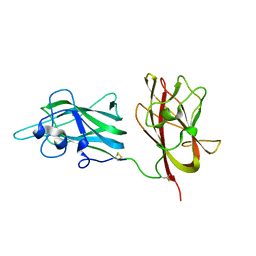 | | Structural Basis for Ligand Binding and Heparin Mediated Activation of Neuropilin | | Descriptor: | Neuropilin-1 | | Authors: | Vander Kooi, C.W, Jusino, M.A, Perman, B, Neau, D.B, Bellamy, H.D, Leahy, D.J. | | Deposit date: | 2007-02-05 | | Release date: | 2007-04-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for ligand and heparin binding to neuropilin B domains
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
4DEQ
 
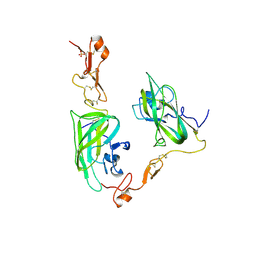 | | Structure of the Neuropilin-1/VEGF-A complex | | Descriptor: | Neuropilin-1, Vascular endothelial growth factor A, PHOSPHATE ION | | Authors: | Vander Kooi, C.W. | | Deposit date: | 2012-01-21 | | Release date: | 2012-02-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.649 Å) | | Cite: | Structural Basis for Selective Vascular Endothelial Growth Factor-A (VEGF-A) Binding to Neuropilin-1.
J.Biol.Chem., 287, 2012
|
|
5C1F
 
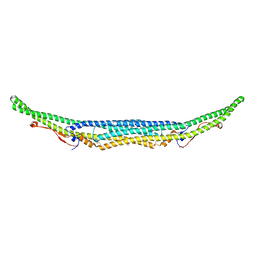 | | Structure of the Imp2 F-BAR domain | | Descriptor: | FORMIC ACID, Septation protein imp2 | | Authors: | Vander Kooi, C.W. | | Deposit date: | 2015-06-13 | | Release date: | 2016-01-27 | | Last modified: | 2019-12-25 | | Method: | X-RAY DIFFRACTION (2.3551 Å) | | Cite: | The Tubulation Activity of a Fission Yeast F-BAR Protein Is Dispensable for Its Function in Cytokinesis.
Cell Rep, 14, 2016
|
|
4RKK
 
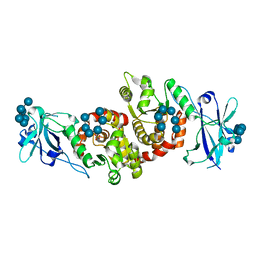 | | Structure of a product bound phosphatase | | Descriptor: | Laforin, PHOSPHATE ION, alpha-D-glucopyranose, ... | | Authors: | Vander Kooi, C.W. | | Deposit date: | 2014-10-13 | | Release date: | 2015-01-07 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural mechanism of laforin function in glycogen dephosphorylation and lafora disease.
Mol.Cell, 57, 2015
|
|
1N87
 
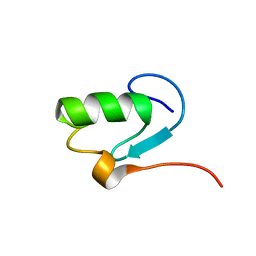 | |
3BXL
 
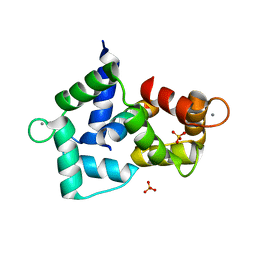 | | Crystal structure of the R-type calcium channeL (CaV2.3) IQ domain and CA2+calmodulin complex | | Descriptor: | CALCIUM ION, Calmodulin, SULFATE ION, ... | | Authors: | Mori, M.X, Vander Kooi, C.W, Leahy, D.J, Yue, D.T. | | Deposit date: | 2008-01-14 | | Release date: | 2008-03-25 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the CaV2 IQ domain in complex with Ca2+/calmodulin
To be Published
|
|
3BXK
 
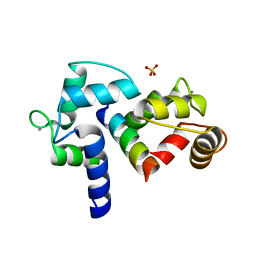 | | Crystal structure of the P/Q-type calcium channel (CaV2.1) IQ domain and CA2+calmodulin complex | | Descriptor: | CALCIUM ION, Calmodulin, SULFATE ION, ... | | Authors: | Mori, M.X, Vander Kooi, C.W, Leahy, D.J, Yue, D.T. | | Deposit date: | 2008-01-14 | | Release date: | 2008-03-25 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Crystal structure of the P/Q-type calcium channel (CaV2.1) IQ domain and CA2+calmodulin complex
To be Published
|
|
4KYQ
 
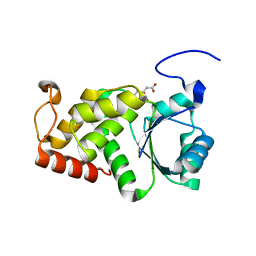 | | Structure of a product bound plant phosphatase | | Descriptor: | CITRATE ANION, Phosphoglucan phosphatase LSF2, chloroplastic | | Authors: | Meekins, D.A, Guo, H.-F, Husodo, S, Paasch, B.C, Bridges, T.M, Santelia, D, Kotting, O, Vander Kooi, C.W, Gentry, M.S. | | Deposit date: | 2013-05-29 | | Release date: | 2013-07-24 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Structure of the Arabidopsis Glucan Phosphatase LIKE SEX FOUR2 Reveals a Unique Mechanism for Starch Dephosphorylation.
Plant Cell, 25, 2013
|
|
4KYR
 
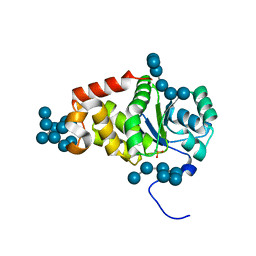 | | Structure of a product bound plant phosphatase | | Descriptor: | PHOSPHATE ION, Phosphoglucan phosphatase LSF2, chloroplastic, ... | | Authors: | Meekins, D.A, Guo, H.-F, Husodo, S, Paasch, B.C, Bridges, T.M, Santelia, D, Kotting, O, Vander Kooi, C.W, Gentry, M.S. | | Deposit date: | 2013-05-29 | | Release date: | 2013-07-24 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of the Arabidopsis Glucan Phosphatase LIKE SEX FOUR2 Reveals a Unique Mechanism for Starch Dephosphorylation.
Plant Cell, 25, 2013
|
|
4QDQ
 
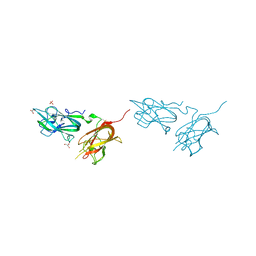 | | Physical basis for Nrp2 ligand binding | | Descriptor: | GLYCEROL, Neuropilin-2, SULFATE ION | | Authors: | Parker, M.W, Vander Kooi, C.W. | | Deposit date: | 2014-05-14 | | Release date: | 2015-04-15 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural Basis for VEGF-C Binding to Neuropilin-2 and Sequestration by a Soluble Splice Form.
Structure, 23, 2015
|
|
4PYH
 
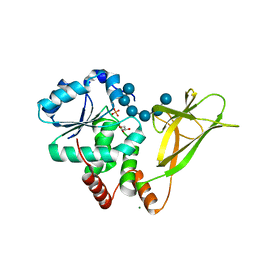 | |
4Q5U
 
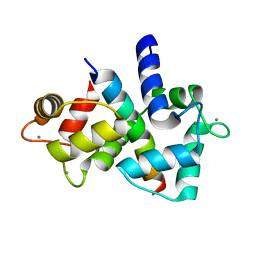 | | Structure of calmodulin bound to its recognition site from calcineurin | | Descriptor: | CALCIUM ION, Calmodulin, Serine/threonine-protein phosphatase 2B catalytic subunit alpha isoform | | Authors: | Guo, H, Dunlap, T.B, Creamer, T.P, Vander Kooi, C.W. | | Deposit date: | 2014-04-17 | | Release date: | 2014-09-03 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Stoichiometry of the calcineurin regulatory domain-calmodulin complex.
Biochemistry, 53, 2014
|
|
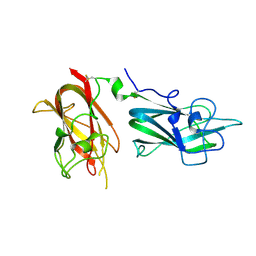
4QDR
 | |
4QDS
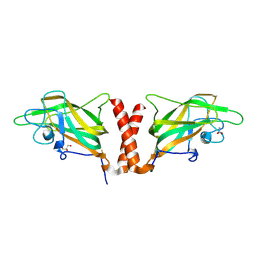 
 | | Physical basis for Nrp2 ligand binding | | Descriptor: | ACETATE ION, GLYCEROL, Neuropilin-2 | | Authors: | Parker, M.W, Vander Kooi, C.W. | | Deposit date: | 2014-05-14 | | Release date: | 2015-04-15 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural Basis for VEGF-C Binding to Neuropilin-2 and Sequestration by a Soluble Splice Form.
Structure, 23, 2015
|
|
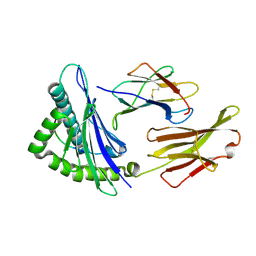
6UJO
 
 | |
1DG4
 
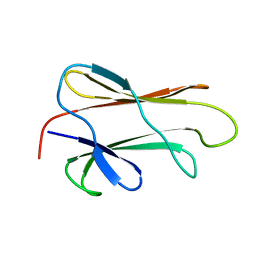 | | NMR STRUCTURE OF THE SUBSTRATE BINDING DOMAIN OF DNAK IN THE APO FORM | | Descriptor: | DNAK | | Authors: | Pellecchia, M, Montgomery, D.L, Stevens, S.Y, Van der Kooi, C.W, Feng, H, Gierasch, L.M, Zuiderweg, E.R.P. | | Deposit date: | 1999-11-23 | | Release date: | 1999-12-08 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural insights into substrate binding by the molecular chaperone DnaK.
Nat.Struct.Biol., 7, 2000
|
|
5E9D
 
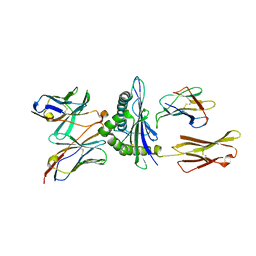 | |
5JZI
 
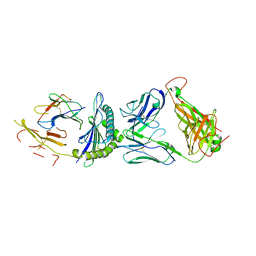 | | Crystal structure of 1406 TCR bound to HLA-A2 with HCV 1406-1415 antigen peptide | | Descriptor: | Beta-2-microglobulin, HCV1406 TCR alpha chain, HCV1406 TCR beta chain, ... | | Authors: | Wang, Y, Piepenbrink, K.H, Baker, B.M. | | Deposit date: | 2016-05-16 | | Release date: | 2017-05-31 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | How an alloreactive T-cell receptor achieves peptide and MHC specificity.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
6D78
 
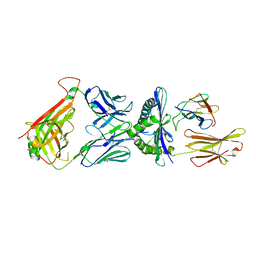 | | The complex between high-affinity TCR DMF5(alpha-D26Y,beta-L98W) and human Class I MHC HLA-A2 with the bound MART-1(27-35)peptide | | Descriptor: | Beta-2-microglobulin, DMF5 alpha chain,DMF5 alpha chain, DMF5 beta chain,DMF5 beta chain, ... | | Authors: | Hellman, L.M, Singh, N.K. | | Deposit date: | 2018-04-24 | | Release date: | 2019-04-03 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.347 Å) | | Cite: | Improving T Cell Receptor On-Target Specificity via Structure-Guided Design.
Mol. Ther., 27, 2019
|
|
6DKP
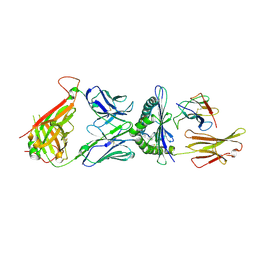 
 | | The complex among DMF5(alpha-D26Y, alpha-Y50A,beta-L98W) TCR, human Class I MHC HLA-A2 and MART-1(26-35)(A27L) peptide | | Descriptor: | Beta-2-microglobulin, DMF5 T-cell Receptor Alpha Chain fusion, DMF5 T-cell Receptor Beta Chain fusion, ... | | Authors: | Hellman, L.M, Singh, N.K. | | Deposit date: | 2018-05-30 | | Release date: | 2019-04-10 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.966 Å) | | Cite: | Improving T Cell Receptor On-Target Specificity via Structure-Guided Design.
Mol. Ther., 27, 2019
|
|
4L3E
 
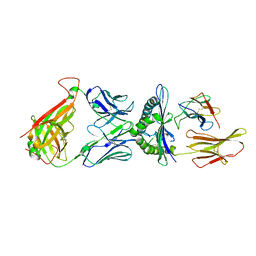 | |
