3ZK6
 
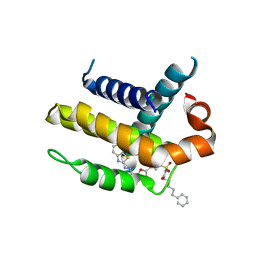 | | Crystal structure of Bcl-xL in complex with inhibitor (Compound 2). | | Descriptor: | BCL-2-LIKE PROTEIN 1, N-(3-(5-(1-(2-(benzo[d]thiazol-2-yl)hydrazono)ethyl)furan-2-yl)phenylsulfonyl)-6-phenylhexanamide | | Authors: | Czabotar, P.E, Lessene, G.L, Smith, B.J, Colman, P.M. | | Deposit date: | 2013-01-22 | | Release date: | 2013-04-24 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Structure-Guided Design of a Selective Bcl-Xl Inhibitor
Nat.Chem.Biol., 9, 2013
|
|
3ZLN
 
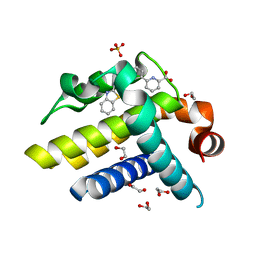 | | Crystal structure of BCL-XL in complex with inhibitor (Compound 3) | | Descriptor: | 1,2-ETHANEDIOL, 6-[(8E)-8-(1,3-benzothiazol-2-ylhydrazinylidene)-6,7-dihydro-5H-naphthalen-2-yl]pyridine-2-carboxylic acid, BCL-2-LIKE PROTEIN 1, ... | | Authors: | Czabotar, P.E, Lessene, G.L, Smith, B.J, Colman, P.M. | | Deposit date: | 2013-02-04 | | Release date: | 2013-04-24 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.288 Å) | | Cite: | Structure-Guided Design of a Selective Bcl-Xl Inhibitor
Nat.Chem.Biol., 9, 2013
|
|
4A1U
 
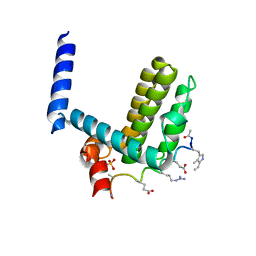 | | Crystal structure of alpha-beta-foldamer 2c in complex with Bcl-xL | | Descriptor: | ALPHA-BETA-FOLDAMER 2C, BCL-2-LIKE PROTEIN 1, CHLORIDE ION, ... | | Authors: | Boersma, M.D, Haase, H.S, Kaufman, K.J, Horne, W.S, Lee, E.F, Clarke, O.B, Smith, B.J, Colman, P.M, Gellman, S.H, Fairlie, W.D. | | Deposit date: | 2011-09-20 | | Release date: | 2011-12-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Evaluation of Diverse Alpha/Beta-Backbone Patterns for Functional Alpha-Helix Mimicry: Analogues of the Bim Bh3 Domain.
J.Am.Chem.Soc., 134, 2012
|
|
4A1W
 
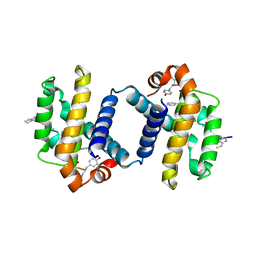 | | Crystal structure of alpha-beta foldamer 4c in complex with Bcl-xL | | Descriptor: | ALPHA-BETA-FOLDAMER 2C, BCL-2-LIKE PROTEIN 1 | | Authors: | Boersma, M.D, Haase, H.S, Kaufman, K.J, Horne, W.S, Lee, E.F, Clarke, O.B, Smith, B.J, Colman, P.M, Gellman, S.H, Fairlie, W.D. | | Deposit date: | 2011-09-20 | | Release date: | 2011-12-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.497 Å) | | Cite: | Evaluation of Diverse Alpha/Beta-Backbone Patterns for Functional Alpha-Helix Mimicry: Analogues of the Bim Bh3 Domain.
J.Am.Chem.Soc., 134, 2012
|
|
1FDY
 
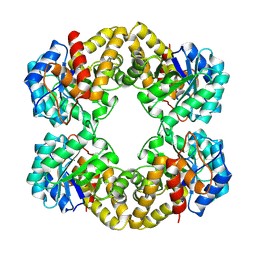 | | N-ACETYLNEURAMINATE LYASE IN COMPLEX WITH HYDROXYPYRUVATE | | Descriptor: | 3-HYDROXYPYRUVIC ACID, N-ACETYLNEURAMINATE LYASE | | Authors: | Lawrence, M.C, Barbosa, J.A.R.G, Smith, B.J, Hall, N.E, Pilling, P.A, Ooi, H.C, Marcuccio, S.M. | | Deposit date: | 1996-07-08 | | Release date: | 1997-10-22 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Structure and mechanism of a sub-family of enzymes related to N-acetylneuraminate lyase.
J.Mol.Biol., 266, 1997
|
|
1FDZ
 
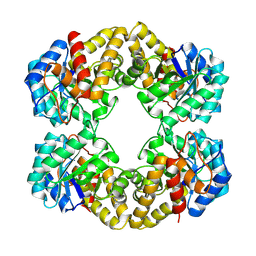 | | N-ACETYLNEURAMINATE LYASE IN COMPLEX WITH PYRUVATE VIA BOROHYDRIDE REDUCTION | | Descriptor: | N-ACETYLNEURAMINATE LYASE, PYRUVIC ACID | | Authors: | Lawrence, M.C, Barbosa, J.A.R.G, Smith, B.J, Hall, N.E, Pilling, P.A, Ooi, H.C, Marcuccio, S.M. | | Deposit date: | 1996-07-08 | | Release date: | 1997-10-22 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure and mechanism of a sub-family of enzymes related to N-acetylneuraminate lyase.
J.Mol.Biol., 266, 1997
|
|
1G5G
 
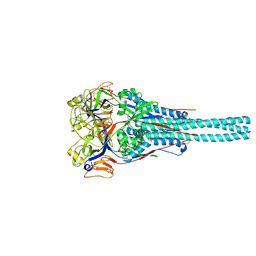 | | FRAGMENT OF FUSION PROTEIN FROM NEWCASTLE DISEASE VIRUS | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Fusion glycoprotein F0, ... | | Authors: | Lawrence, M.C, Smith, B.J. | | Deposit date: | 2000-11-01 | | Release date: | 2002-02-27 | | Last modified: | 2022-12-21 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | The structure of the fusion glycoprotein of Newcastle disease virus suggests a novel paradigm for the molecular mechanism of membrane fusion.
Structure, 9, 2001
|
|
6V4M
 
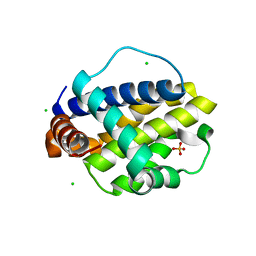 | | Trichuris suis BCL-2 | | Descriptor: | BCL-2, CHLORIDE ION, SULFATE ION | | Authors: | Fairlie, W.D, Lee, E.F, Smith, B.J. | | Deposit date: | 2019-11-28 | | Release date: | 2020-10-07 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.599 Å) | | Cite: | Diversity in the intrinsic apoptosis pathway of nematodes.
Commun Biol, 3, 2020
|
|
5VAX
 
 | |
5VAY
 
 | |
5VAU
 
 | |
4WAT
 
 | |
4QVX
 
 | |
7LK4
 
 | |
5J3H
 | | Human insulin receptor domains L1-CR in complex with peptide S519C16 and 83-7 Fv | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lawrence, M, Menting, J, Lawrence, C. | | Deposit date: | 2016-03-30 | | Release date: | 2016-06-15 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.27 Å) | | Cite: | Insulin Mimetic Peptide Disrupts the Primary Binding Site of the Insulin Receptor.
J.Biol.Chem., 291, 2016
|
|
5JYQ
 
 | | Structure of Conus Geographus insulin G1 | | Descriptor: | Insulin 1, Insulin 1b, SULFATE ION | | Authors: | Lawrence, M.C, Menting, J, Norton, R.S, Safavi-Hemami, H. | | Deposit date: | 2016-05-15 | | Release date: | 2016-09-07 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | A minimized human insulin-receptor-binding motif revealed in a Conus geographus venom insulin.
Nat.Struct.Mol.Biol., 23, 2016
|
|
7KD6
 
 | |
7N9L
 | | KirBac3.1 C71S C262S | | Descriptor: | 1,2-DIOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, Inward rectifier potassium channel Kirbac3.1, POTASSIUM ION, ... | | Authors: | Gulbis, J.M, Black, K.A. | | Deposit date: | 2021-06-18 | | Release date: | 2021-12-29 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Ion currents through Kir potassium channels are gated by anionic lipids.
Nat Commun, 13, 2022
|
|
7N9K
 
 | | KirBac3.1 L124M mutant | | Descriptor: | 1,2-DIOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, Inward rectifier potassium channel Kirbac3.1, N-OCTANE, ... | | Authors: | Black, T.A, Gulbis, J.M. | | Deposit date: | 2021-06-18 | | Release date: | 2021-12-29 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.72 Å) | | Cite: | Ion currents through Kir potassium channels are gated by anionic lipids.
Nat Commun, 13, 2022
|
|
1YWG
 
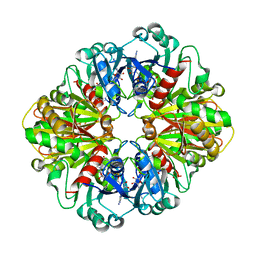 | | The structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, glyceraldehyde-3-phosphate dehydrogenase | | Authors: | Satchell, J.F, Klonis, N, Malby, R.L, Adisa, A, Alpyurek, A.E, Colman, P.M, Tilley, L. | | Deposit date: | 2005-02-17 | | Release date: | 2005-11-29 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
6UXO
 
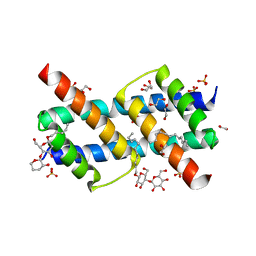 | | Crystal structure of BAK core domain BH3-groove-dimer in complex with DDM | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Bcl-2 homologous antagonist/killer, ... | | Authors: | Cowan, A.D, Colman, P.M, Czabotar, P.E. | | Deposit date: | 2019-11-07 | | Release date: | 2020-09-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.799 Å) | | Cite: | BAK core dimers bind lipids and can be bridged by them.
Nat.Struct.Mol.Biol., 27, 2020
|
|
6UXQ
 
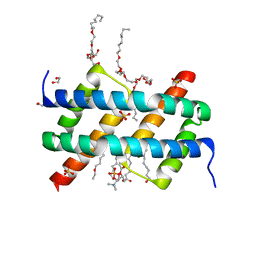 | | Crystal structure of BAK core domain BH3-groove-dimer in complex with POPC and C8E4 | | Descriptor: | (HYDROXYETHYLOXY)TRI(ETHYLOXY)OCTANE, 1,2-ETHANEDIOL, 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine, ... | | Authors: | Cowan, A.D, Colman, P.M, Czabotar, P.E. | | Deposit date: | 2019-11-07 | | Release date: | 2020-09-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.696 Å) | | Cite: | BAK core dimers bind lipids and can be bridged by them.
Nat.Struct.Mol.Biol., 27, 2020
|
|
6VEP
 
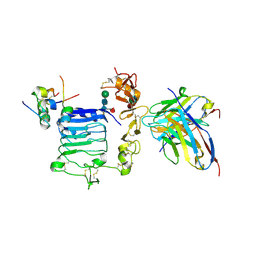 | | Human insulin in complex with the human insulin microreceptor in turn in complex with Fv 83-7 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lawrence, M.C, Menting, J.G.T. | | Deposit date: | 2020-01-02 | | Release date: | 2020-06-03 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | A structurally minimized yet fully active insulin based on cone-snail venom insulin principles.
Nat.Struct.Mol.Biol., 27, 2020
|
|
6VET
 | | Human insulin analog: [GluB10,HisA8,ArgA9,TyrB20]-DOI | | Descriptor: | Insulin A chain, Insulin B chain | | Authors: | Menting, J.G, Chou, D.H.-C, Lawrence, M.C, Xiong, X. | | Deposit date: | 2020-01-02 | | Release date: | 2020-06-03 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | A structurally minimized yet fully active insulin based on cone-snail venom insulin principles.
Nat.Struct.Mol.Biol., 27, 2020
|
|
6UXN
 
 | | Crystal structure of BAK core domain BH3-groove-dimer in complex with phosphatidylserine | | Descriptor: | Bcl-2 homologous antagonist/killer, GLYCEROL, O-[(R)-{[(2R)-2,3-bis(octanoyloxy)propyl]oxy}(hydroxy)phosphoryl]-L-serine, ... | | Authors: | Cowan, A.D, Colman, P.M, Czabotar, P.E. | | Deposit date: | 2019-11-07 | | Release date: | 2020-09-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | BAK core dimers bind lipids and can be bridged by them.
Nat.Struct.Mol.Biol., 27, 2020
|
|
