8YYI
 
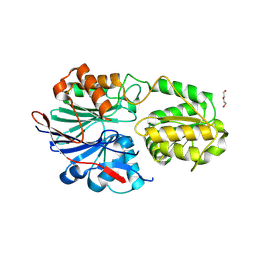 | | RNase J2 mutant E166A | | Descriptor: | DI(HYDROXYETHYL)ETHER, Ribonuclease J 2 | | Authors: | Singh, A.K, Chinnasamy, K, Gopal, B. | | Deposit date: | 2024-04-03 | | Release date: | 2025-02-12 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | A physicochemical rationale for the varied catalytic efficiency in RNase J paralogues.
J.Biol.Chem., 301, 2024
|
|
8YYF
 
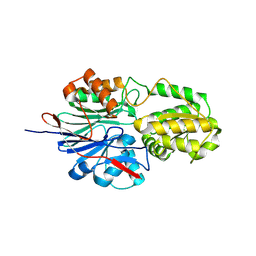 | | RNase J2 mutant H76A | | Descriptor: | Ribonuclease J 2 | | Authors: | Singh, A.K, Chinnasamy, K, Gopal, B. | | Deposit date: | 2024-04-03 | | Release date: | 2025-02-12 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | A physicochemical rationale for the varied catalytic efficiency in RNase J paralogues.
J.Biol.Chem., 301, 2024
|
|
8YYH
 
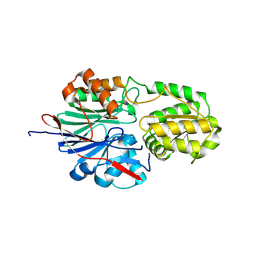 | | RNase J2 mutant H144A | | Descriptor: | Ribonuclease J 2 | | Authors: | Singh, A.K, Chinnasamy, K, Gopal, B. | | Deposit date: | 2024-04-03 | | Release date: | 2025-02-12 | | Method: | X-RAY DIFFRACTION (1.12 Å) | | Cite: | A physicochemical rationale for the varied catalytic efficiency in RNase J paralogues.
J.Biol.Chem., 301, 2024
|
|
4UMI
 
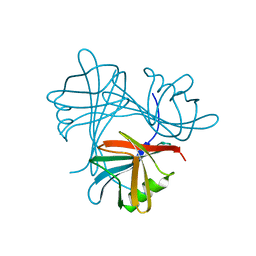 | |
7A2F
 
 | | Crystal structure of T. brucei PDE-B1 catalytic domain with inhibitor NPD-656 | | Descriptor: | 1-cycloheptyl-3-{4-methoxy-3-[2-(4-methoxyphenyl)ethoxy]phenyl}-4,4-dimethyl-4,5-dihydro-1H-pyrazol-5-one, FORMIC ACID, GLYCEROL, ... | | Authors: | Singh, A.K, Blaazer, A.R, Zara, L, de Esch, I.J.P, Leurs, R, Brown, D.G. | | Deposit date: | 2020-08-17 | | Release date: | 2021-08-25 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of T. brucei PDE-B1 catalytic domain with inhibitor NPD-656
To be published
|
|
7A28
 | | Crystal structure of T. brucei PDE-B1 catalytic domain with inhibitor NPD-617 | | Descriptor: | 1-cycloheptyl-3-(4-methoxy-3-{4-[4-(1H-1,2,3,4-tetrazol-5-yl)phenoxy]butoxy}phenyl)-4,4-dimethyl-4,5-dihydro-1H-pyrazol-5-one, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Singh, A.K, Blaazer, A.R, Zara, L, de Esch, I.J.P, Leurs, R, Brown, D.G. | | Deposit date: | 2020-08-16 | | Release date: | 2021-08-25 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Crystal structure of T. brucei PDE-B1 catalytic domain with inhibitor NPD-617
To be published
|
|
7P15
 
 | |
7OZW
 
 | |
5NBH
 | |
5NC1
 
 | |
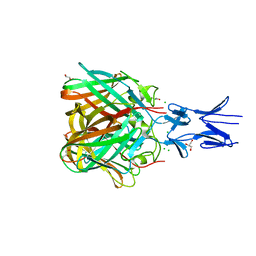
5N8D
 
 | |
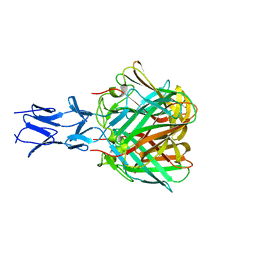
5N83
 
 | |
6FRD
 | |
6FT0
 
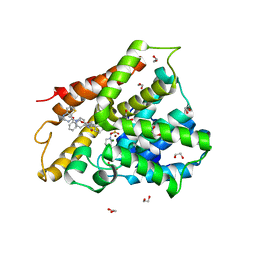 | | Crystal structure of human phosphodiesterase 4D2 catalytic domain with inhibitor NPD-425 | | Descriptor: | 1,2-ETHANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, 4-{4-methoxy-3-[(2-methoxyphenyl)methoxy]phenyl}-2-(1-{thieno[3,2-d]pyrimidin-4-yl}piperidin-4-yl)-1,2,4a,5,8,8a-hexahydrophthalazin-1-one, ... | | Authors: | Singh, A.K, Brown, D.G. | | Deposit date: | 2018-02-20 | | Release date: | 2019-03-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | hPDE4D2 structure with inhibitor NPD-425
To be published
|
|
6FTA
 
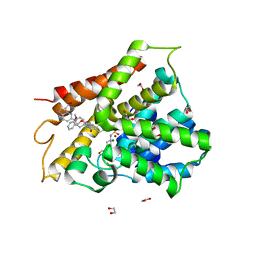 | | Crystal structure of human phosphodiesterase 4D2 catalytic domain with inhibitor NPD-3098 | | Descriptor: | (4~{a}~{S},8~{a}~{S})-4-[4-methoxy-3-[(2-methoxyphenyl)methoxy]phenyl]-2-[1-(3-nitroimidazo[1,2-b]pyridazin-6-yl)piperidin-4-yl]-4~{a},5,6,7,8,8~{a}-hexahydrophthalazin-1-one, 1,2-ETHANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Singh, A.K, Brown, D.G. | | Deposit date: | 2018-02-20 | | Release date: | 2019-03-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | hPDE4D2 structure with inhibitor NPD-3098
To be published
|
|
6PVQ
 
 | |
6PVN
 
 | |
6PVM
 
 | |
6PVL
 
 | |
6PVO
 
 | |
6QGU
 
 | | Crystal structure of T. brucei PDE-B1 catalytic domain with inhibitor NPD-1361 | | Descriptor: | 5-(3-(cyclopentyloxy)-4-methoxyphenyl)-2-isopropyl-4,4-dimethyl-2,4-dihydro-3H-pyrazol-3-one, FORMIC ACID, GLYCEROL, ... | | Authors: | Singh, A.K, Blaazer, A.R, Zara, L, de Esch, I.J.P, Leurs, R, Brown, D.G. | | Deposit date: | 2019-01-13 | | Release date: | 2020-02-05 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Crystal structure of T. brucei PDE-B1 catalytic domain with inhibitor NPD-1361
To be published
|
|
6PVP
 
 | |
6QGP
 
 | | Crystal structure of T. brucei PDE-B1 catalytic domain with inhibitor NPD-0769 | | Descriptor: | 1-cycloheptyl-3-[3-(cyclopentyloxy)-4-methoxyphenyl]-4,4-dimethyl-4,5-dihydro-1H-pyrazol-5-one, DI(HYDROXYETHYL)ETHER, FORMIC ACID, ... | | Authors: | Singh, A.K, Blaazer, A.R, Zara, L, de Esch, I.J.P, Leurs, R, Brown, D.G. | | Deposit date: | 2019-01-12 | | Release date: | 2020-02-05 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.942 Å) | | Cite: | Crystal structure of T. brucei PDE-B1 catalytic domain with inhibitor NPD-0769
To be published
|
|
7ABD
 
 | | Crystal structure of human phosphodiesterase 4D2 catalytic domain with inhibitor NPD-768 | | Descriptor: | 1,2-ETHANEDIOL, 3-(3-cyclopentyloxy-4-methoxy-phenyl)-4,4-dimethyl-1~{H}-pyrazol-5-one, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Singh, A.K, Blaazer, A.R, Zara, L, de Esch, I.J.P, Leurs, R, Brown, D.G. | | Deposit date: | 2020-09-07 | | Release date: | 2021-10-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | hPDE4D2 structure with inhibitor NPD-768
To be published
|
|
7A8Q
 
 | | Crystal structure of human phosphodiesterase 4D2 catalytic domain with inhibitor NPD-654 | | Descriptor: | 1,2-ETHANEDIOL, 2-cycloheptyl-5-[4-methoxy-3-[[4-(1~{H}-1,2,3,4-tetrazol-5-yl)phenyl]methoxy]phenyl]-4,4-dimethyl-pyrazolidin-3-one, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Singh, A.K, Blaazer, A.R, Zara, L, de Esch, I.J.P, Leurs, R, Brown, D.G. | | Deposit date: | 2020-08-30 | | Release date: | 2021-10-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | hPDE4D2 structure with inhibitor NPD-654
To be published
|
|
