2BHF
 
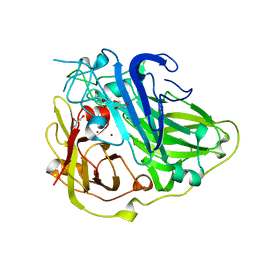 | | 3D structure of the reduced form of CotA | | Descriptor: | COPPER (I) ION, GLYCEROL, SPORE COAT PROTEIN A | | Authors: | Bento, I, Martins, L.O, Lopes, G.G, Carrondo, M.A, Lindley, P.F. | | Deposit date: | 2005-01-10 | | Release date: | 2005-10-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Dioxygen Reduction by Multi-Copper Oxidases; a Structural Perspective.
Dalton Trans., 7, 2005
|
|
2A8F
 
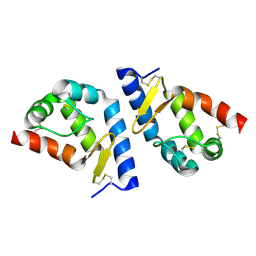 | | beta-cinnamomin after sterol removal | | Descriptor: | Beta-elicitin cinnamomin | | Authors: | Rodrigues, M.L, Archer, M, Martel, P, Miranda, S, Thomaz, M, Enguita, F.J, Baptista, R.P, Melo, E.P, Sousa, N, Cravador, A, Carrondo, M.A. | | Deposit date: | 2005-07-08 | | Release date: | 2006-01-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal structures of the free and sterol-bound forms of beta-cinnamomin
Biochim.Biophys.Acta, 1764, 2006
|
|
2AIB
 
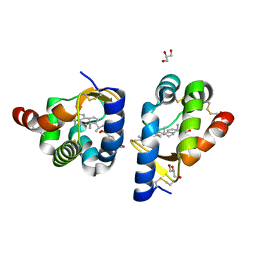 | | beta-cinnamomin in complex with ergosterol | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Beta-elicitin cinnamomin, ERGOSTEROL, ... | | Authors: | Rodrigues, M.L, Archer, M, Martel, P, Miranda, S, Thomaz, M, Enguita, F.J, Baptista, R.P, Melo, E.P, Sousa, N, Cravador, A, Carrondo, M.A. | | Deposit date: | 2005-07-29 | | Release date: | 2006-01-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Crystal structures of the free and sterol-bound forms of beta-cinnamomin
Biochim.Biophys.Acta, 1764, 2006
|
|
4C7U
 
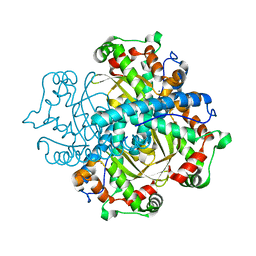 | | Crystal structure of manganese superoxide dismutase from Arabidopsis thaliana | | Descriptor: | MANGANESE (II) ION, SUPEROXIDE DISMUTASE [MN] 1, MITOCHONDRIAL | | Authors: | Marques, A, Santos, S.P, Rosa, M, Carrondo, M.A, Abreu, I.A, Romao, C.V, Frazao, C. | | Deposit date: | 2013-09-25 | | Release date: | 2014-10-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.951 Å) | | Cite: | Crystal Structure of the Arabidopsis Thaliana Manganese Superoxide Dismutase at 1.95 A Resolution
To be Published
|
|
4D02
 
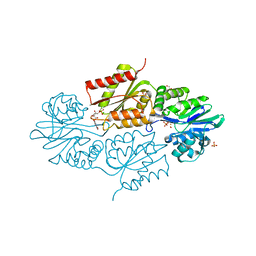 | | The crystallographic structure of Flavorubredoxin from Escherichia coli | | Descriptor: | ANAEROBIC NITRIC OXIDE REDUCTASE FLAVORUBREDOXIN, CHLORIDE ION, FE (III) ION, ... | | Authors: | Romao, C.V, Vicente, J.B, Bandeiras, T, Carrondo, M.A, Teixeira, M, Frazao, C. | | Deposit date: | 2014-04-23 | | Release date: | 2014-09-03 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.755 Å) | | Cite: | Structure of Escherichia Coli Flavodiiron Nitric Oxide Reductase.
J.Mol.Biol., 428, 2016
|
|
4COT
 
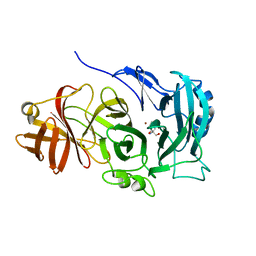 | | The importance of the Abn2 calcium cluster in the endo-1,5- arabinanase activity from Bacillus subtilis | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, EXTRACELLULAR ENDO-ALPHA-(1->5)-L-ARABINANASE 2, NICKEL (II) ION | | Authors: | McVey, C.E, Ferreira, M.J, Correia, B, Lahiri, S, deSanctis, D, Carrondo, M.A, Lindley, P.F, de Sa-Nogueira, I, Soares, C.M, Bento, I. | | Deposit date: | 2014-01-31 | | Release date: | 2014-03-05 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Importance of the Abn2 Calcium Cluster in the Endo-1,5-Arabinanase Activity from Bacillus Subtilis.
J.Biol.Inorg.Chem., 19, 2014
|
|
1LJP
 
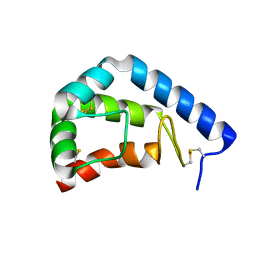 | | Crystal Structure of beta-Cinnamomin Elicitin | | Descriptor: | Beta-elicitin cinnamomin | | Authors: | Rodrigues, M.L, Archer, M, Martel, P, Jacquet, A, Cravador, A, Carrondo, M.A. | | Deposit date: | 2002-04-22 | | Release date: | 2002-07-31 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of beta-cinnamomin, a protein toxic to some plant species.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1NF6
 
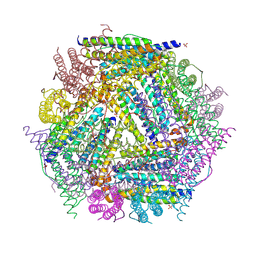 | | X-ray structure of the Desulfovibrio desulfuricans bacterioferritin: the diiron site in different catalytic states ("cycled" structure: reduced in solution and allowed to reoxidise before crystallisation) | | Descriptor: | 1,3,5,8-TETRAMETHYL-PORPHINE-2,4,6,7-TETRAPROPIONIC ACID FERROUS COMPLEX, FE (III) ION, GLYCEROL, ... | | Authors: | Macedo, S, Romao, C.V, Mitchell, E, Matias, P.M, Liu, M.Y, Xavier, A.V, LeGall, J, Teixeira, M, Lindley, P, Carrondo, M.A. | | Deposit date: | 2002-12-13 | | Release date: | 2003-04-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | The nature of the di-iron site in the bacterioferritin from
Desulfovibrio desulfuricans
NAT.STRUCT.BIOL., 10, 2003
|
|
1NF4
 
 | | X-Ray Structure of the Desulfovibrio desulfuricans bacterioferritin: the diiron site in different states (reduced structure) | | Descriptor: | 1,3,5,8-TETRAMETHYL-PORPHINE-2,4,6,7-TETRAPROPIONIC ACID FERROUS COMPLEX, FE (II) ION, SULFATE ION, ... | | Authors: | Macedo, S, Romao, C.V, Mitchell, E, Matias, P.M, Liu, M.Y, Xavier, A.V, LeGall, J, Teixeira, M, Lindley, P, Carrondo, M.A. | | Deposit date: | 2002-12-13 | | Release date: | 2003-04-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | The nature of the di-iron site in the bacterioferritin from
Desulfovibrio desulfuricans
NAT.STRUCT.BIOL., 10, 2003
|
|
1NFV
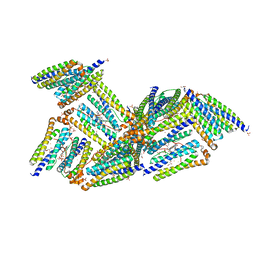 
 | | X-ray structure of Desulfovibrio desulfuricans bacterioferritin: the diiron centre in different catalytic states (as-isolated structure) | | Descriptor: | 1,3,5,8-TETRAMETHYL-PORPHINE-2,4,6,7-TETRAPROPIONIC ACID FERROUS COMPLEX, 3-HYDROXYPYRUVIC ACID, FE (III) ION, ... | | Authors: | Macedo, S, Romao, C.V, Mitchell, E, Matias, P.M, Liu, M.Y, Xavier, A.V, LeGall, J, Teixeira, M, Lindley, P, Carrondo, M.A. | | Deposit date: | 2002-12-16 | | Release date: | 2003-04-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The nature of the di-iron site in the bacterioferritin from
Desulfovibrio desulfuricans
NAT.STRUCT.BIOL., 10, 2003
|
|
1H1D
 
 | | Catechol O-Methyltransferase | | Descriptor: | 1-(3,4,DIHYDROXY-5-NITROPHENYL)-3-{4-[3-(TRIFLUOROMETHYL) PHENYL] PIPERAZIN-1-YL}PROPAN-1-ONE, CATECHOL-O-METHYLTRANSFERASE, MAGNESIUM ION, ... | | Authors: | Archer, M, Rodrigues, M.L, Matias, P.M, Bonifacio, M.J, Learmonth, D.A, Soares-da-Silva, P, Carrondo, M.A. | | Deposit date: | 2002-07-12 | | Release date: | 2003-07-17 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Kinetics and Crystal Structure of Catechol-O-Methyltransferase Complex with Co-Substrate and a Novel Inhibitor with Potential Therapeutic Application
Mol.Pharmacol., 62, 2002
|
|
2UXY
 
 | | Aliphatic amidase | | Descriptor: | ALIPHATIC AMIDASE, SULFATE ION | | Authors: | Andrade, J, KArmali, A, Carrondo, M.A, Frazao, C. | | Deposit date: | 2007-04-02 | | Release date: | 2007-04-17 | | Last modified: | 2019-05-08 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Structure of Amidase from Pseudomonas Aeruginosa Showing a Trapped Acyl Transfer Reaction Intermediate State.
J.Biol.Chem., 282, 2007
|
|
2V5Q
 
 | | CRYSTAL STRUCTURE OF WILD-TYPE PLK-1 KINASE DOMAIN IN COMPLEX WITH A SELECTIVE DARPIN | | Descriptor: | DESIGN ANKYRIN REPEAT PROTEIN, SERINE/THREONINE-PROTEIN KINASE PLK1 | | Authors: | Bandeiras, T.M, Hillig, R.C, Matias, P.M, Eberspaecher, U, Fanghaenel, J, Thomaz, M, Miranda, S, Crusius, K, Puetter, V, Amstutz, P, Gulotti-Georgieva, M, Binz, H.K, Holz, C, Schmitz, A.A.P, Lang, C, Donner, P, Egner, U, Carrondo, M.A, Mueller-Tiemann, B. | | Deposit date: | 2007-07-08 | | Release date: | 2008-04-01 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of wild-type Plk-1 kinase domain in complex with a selective DARPin.
Acta Crystallogr. D Biol. Crystallogr., 64, 2008
|
|
2VKR
 
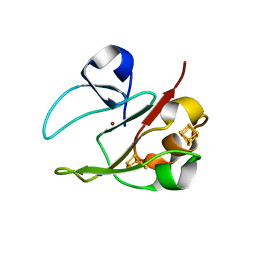 | | 3Fe-4S, 4Fe-4S plus Zn Acidianus ambivalens ferredoxin | | Descriptor: | FE3-S4 CLUSTER, IRON/SULFUR CLUSTER, ZINC ION, ... | | Authors: | Frazao, C, Aragao, D, Coelho, R, Leal, S.S, Gomes, C.M, Teixeira, M, Carrondo, M.A. | | Deposit date: | 2007-12-23 | | Release date: | 2008-03-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Crystallographic analysis of the intact metal centres [3Fe-4S](1+/0) and [4Fe-4S](2+/1+) in a Zn(2+) -containing ferredoxin.
FEBS Lett., 582, 2008
|
|
2W48
 
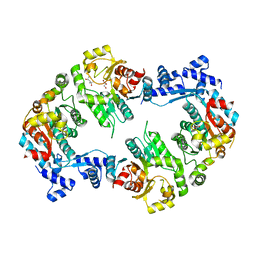 | | Crystal structure of the Full-length Sorbitol Operon Regulator SorC from Klebsiella pneumoniae | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, CHLORIDE ION, ... | | Authors: | de Sanctis, D, McVey, C.E, Enguita, F.J, Carrondo, M.A. | | Deposit date: | 2008-11-21 | | Release date: | 2009-05-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal Structure of the Full-Length Sorbitol Operon Regulator Sorc from Klebsiella Pneumoniae: Structural Evidence for a Novel Transcriptional Regulation Mechanism.
J.Mol.Biol., 387, 2009
|
|
1OFW
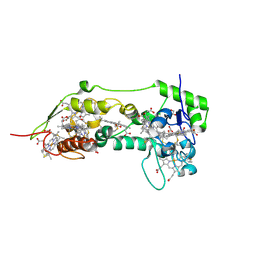 
 | | Three dimensional structure of the oxidized form of nine heme cytochrome c at PH 7.5 | | Descriptor: | ACETATE ION, GLYCEROL, HEME C, ... | | Authors: | Bento, I, Teixeira, V.H, Baptista, A.M, Soares, C.M, Matias, P.M, Carrondo, M.A. | | Deposit date: | 2003-04-22 | | Release date: | 2003-09-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Redox-Bohr and Other Cooperativity Effects in the Nine-Heme Cytochrome C from Desulfovibrio Desulfuricans Atcc 27774: Crystallographic and Modeling Studies
J.Biol.Chem., 278, 2003
|
|
3ZDW
 
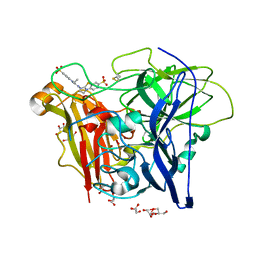 | | Substrate and dioxygen binding to the endospore coat laccase CotA from Bacillus subtilis | | Descriptor: | 3-ETHYL-2-[(2Z)-2-(3-ETHYL-6-SULFO-1,3-BENZOTHIAZOL-2(3H)-YLIDENE)HYDRAZINO]-6-SULFO-3H-1,3-BENZOTHIAZOL-1-IUM, COPPER (I) ION, COTA LACCASE, ... | | Authors: | Enguita, F.J, Marcal, D, Grenha, R, Lindley, P.F, Carrondo, M.A. | | Deposit date: | 2012-12-01 | | Release date: | 2012-12-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Substrate and Dioxygen Binding to the Endospore Coat Laccase from Bacillus Subtilis.
J.Biol.Chem., 279, 2004
|
|
4AJB
 
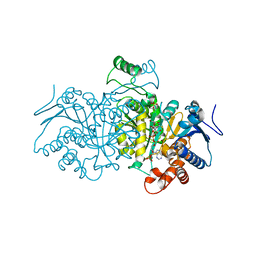 | | 3D structure of E. coli Isocitrate Dehydrogenase K100M mutant in complex with Isocitrate, magnesium(II) and thioNADP | | Descriptor: | 7-THIONICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ISOCITRIC ACID, MAGNESIUM ION, ... | | Authors: | Goncalves, S, Miller, S.P, Carrondo, M.A, Dean, A.M, Matias, P.M. | | Deposit date: | 2012-02-16 | | Release date: | 2012-10-31 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Induced Fit and the Catalytic Mechanism of Isocitrate Dehydrogenase.
Biochemistry, 51, 2012
|
|
4AJC
 
 | | 3D structure of E. coli Isocitrate Dehydrogenase K100M mutant in complex with alpha-ketoglutarate, calcium(II) and adenine nucleotide phosphate | | Descriptor: | 2-OXOGLUTARIC ACID, ADENOSINE-2'-5'-DIPHOSPHATE, CALCIUM ION, ... | | Authors: | Goncalves, S, Miller, S.P, Carrondo, M.A, Dean, A.M, Matias, P.M. | | Deposit date: | 2012-02-16 | | Release date: | 2012-10-31 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Induced Fit and the Catalytic Mechanism of Isocitrate Dehydrogenase.
Biochemistry, 51, 2012
|
|
4AJ3
 
 | | 3D structure of E. coli Isocitrate Dehydrogenase in complex with Isocitrate, calcium(II) and NADP - The pseudo-Michaelis complex | | Descriptor: | CALCIUM ION, ISOCITRIC ACID, NADP ISOCITRATE DEHYDROGENASE, ... | | Authors: | Goncalves, S, Miller, S.P, Carrondo, M.A, Dean, A.M, Matias, P.M. | | Deposit date: | 2012-02-15 | | Release date: | 2012-10-31 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Induced Fit and the Catalytic Mechanism of Isocitrate Dehydrogenase.
Biochemistry, 51, 2012
|
|
4AJR
 
 | | 3D structure of E. coli Isocitrate Dehydrogenase K100M mutant in complex with alpha-ketoglutarate, magnesium(II) and NADPH - The product complex | | Descriptor: | 2-OXOGLUTARIC ACID, BETA-NICOTINAMIDE RIBOSE MONOPHOSPHATE, ISOCITRATE DEHYDROGENASE [NADP], ... | | Authors: | Goncalves, S, Miller, S.P, Carrondo, M.A, Dean, A.M, Matias, P.M. | | Deposit date: | 2012-02-17 | | Release date: | 2012-10-31 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.687 Å) | | Cite: | Induced Fit and the Catalytic Mechanism of Isocitrate Dehydrogenase.
Biochemistry, 51, 2012
|
|
4AJA
 
 | | 3D structure of E. coli Isocitrate Dehydrogenase in complex with Isocitrate, calcium(II) and thioNADP | | Descriptor: | 7-THIONICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, CALCIUM ION, ISOCITRIC ACID, ... | | Authors: | Goncalves, S, Miller, S.P, Carrondo, M.A, Dean, A.M, Matias, P.M. | | Deposit date: | 2012-02-16 | | Release date: | 2012-10-31 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.799 Å) | | Cite: | Induced Fit and the Catalytic Mechanism of Isocitrate Dehydrogenase.
Biochemistry, 51, 2012
|
|
4AJS
 
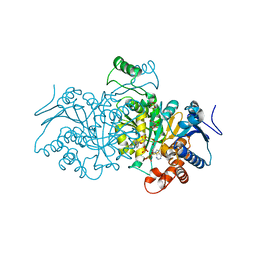 | | 3D structure of E. coli Isocitrate Dehydrogenase K100M mutant in complex with isocitrate, magnesium(II), Adenosine 2',5'-biphosphate and ribosylnicotinamide-5'-phosphate | | Descriptor: | ADENOSINE-2'-5'-DIPHOSPHATE, BETA-NICOTINAMIDE RIBOSE MONOPHOSPHATE, ISOCITRATE DEHYDROGENASE [NADP], ... | | Authors: | Goncalves, S, Miller, S.P, Carrondo, M.A, Dean, A.M, Matias, P.M. | | Deposit date: | 2012-02-17 | | Release date: | 2012-10-31 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.802 Å) | | Cite: | Induced Fit and the Catalytic Mechanism of Isocitrate Dehydrogenase.
Biochemistry, 51, 2012
|
|
4BLG
 
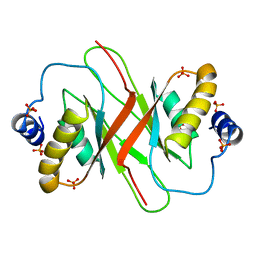 | | Crystal structure of MHV-68 Latency-associated nuclear antigen (LANA) C-terminal DNA binding domain | | Descriptor: | LATENCY-ASSOCIATED NUCLEAR ANTIGEN, PHOSPHATE ION | | Authors: | Correia, B, Cerqueira, S.A, Beauchemin, C, Pires De Miranda, M, Li, S, Ponnusamy, R, Rodrigues, L, Schneider, T.R, Carrondo, M.A, Kaye, K.M, Simas, J.P, McVey, C.E. | | Deposit date: | 2013-05-02 | | Release date: | 2013-10-30 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of the Gamma-2 Herpesvirus Lana DNA Binding Domain Identifies Charged Surface Residues which Impact Viral Latency
Plos Pathog., 9, 2013
|
|
19HC
 
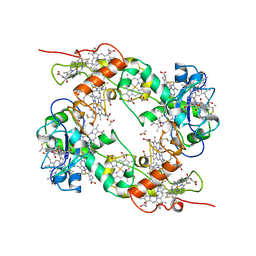 | | NINE-HAEM CYTOCHROME C FROM DESULFOVIBRIO DESULFURICANS ATCC 27774 | | Descriptor: | ACETATE ION, PROTEIN (NINE-HAEM CYTOCHROME C), PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Matias, P.M, Coelho, R, Pereira, I.A.C, Coelho, A.V, Thompson, A.W, Sieker, L, Gall, J.L, Carrondo, M.A. | | Deposit date: | 1998-12-01 | | Release date: | 1999-12-01 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The primary and three-dimensional structures of a nine-haem cytochrome c from Desulfovibrio desulfuricans ATCC 27774 reveal a new member of the Hmc family.
Structure Fold.Des., 7, 1999
|
|
