6LLG
 
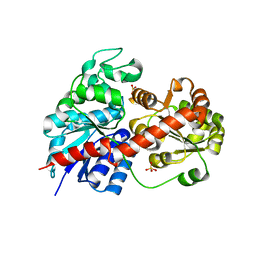 | | Crystal Structure of Fagopyrum esculentum M UGT708C1 | | Descriptor: | BENZAMIDINE, SULFATE ION, UDP-glycosyltransferase 708C1 | | Authors: | Wang, X, Liu, M. | | Deposit date: | 2019-12-23 | | Release date: | 2020-09-09 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structures of theC-Glycosyltransferase UGT708C1 from Buckwheat Provide Insights into the Mechanism ofC-Glycosylation.
Plant Cell, 32, 2020
|
|
6LLZ
 
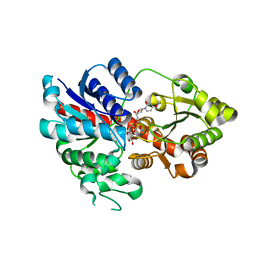 | |
6LLW
 
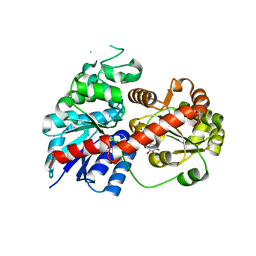 | |
8I3Q
 
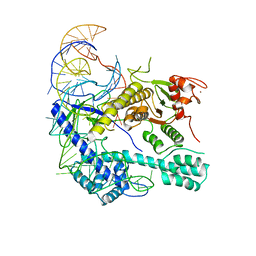 | |
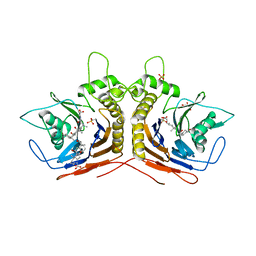
5Y7P
 | |
6UQE
 
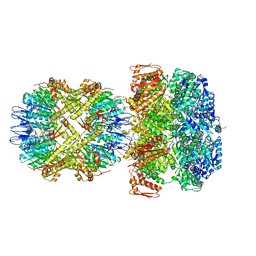 | | ClpA/ClpP Disengaged State bound to RepA-GFP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ATP-dependent Clp protease ATP-binding subunit ClpA, ATP-dependent Clp protease proteolytic subunit, ... | | Authors: | Lopez, K.L, Rizo, A.R, Southworth, D.R. | | Deposit date: | 2019-10-18 | | Release date: | 2020-04-22 | | Last modified: | 2022-03-16 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Conformational plasticity of the ClpAP AAA+ protease couples protein unfolding and proteolysis.
Nat.Struct.Mol.Biol., 27, 2020
|
|
6UQO
 
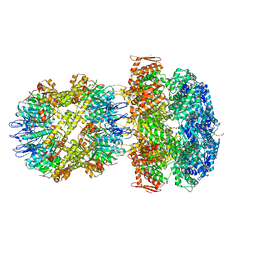 | | ClpA/ClpP Engaged State bound to RepA-GFP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ATP-dependent Clp endopeptidase proteolytic subunit ClpP, ATP-dependent Clp protease ATP-binding subunit ClpA, ... | | Authors: | Lopez, K.L, Rizo, A.N, Southworth, D.R. | | Deposit date: | 2019-10-21 | | Release date: | 2020-04-22 | | Last modified: | 2022-03-16 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Conformational plasticity of the ClpAP AAA+ protease couples protein unfolding and proteolysis.
Nat.Struct.Mol.Biol., 27, 2020
|
|
3CCN
 
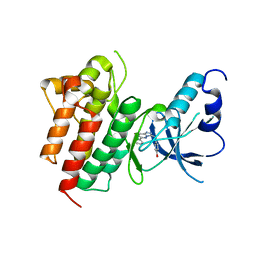 | | X-ray structure of c-Met with triazolopyridazine inhibitor. | | Descriptor: | 4-[(6-phenyl[1,2,4]triazolo[4,3-b]pyridazin-3-yl)methyl]phenol, Hepatocyte growth factor receptor | | Authors: | Abrecht, B.K, Harmange, J.-C, Bauer, D, Dussault, I, long, A, Bellon, S.F. | | Deposit date: | 2008-02-26 | | Release date: | 2008-04-29 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Discovery and Optimization of Triazolopyridazines as Potent and Selective Inhibitors of the c-Met Kinase.
J.Med.Chem., 51, 2008
|
|
3CD8
 
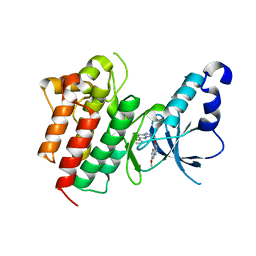 | | X-ray Structure of c-Met with triazolopyridazine Inhibitor. | | Descriptor: | 7-methoxy-4-[(6-phenyl[1,2,4]triazolo[4,3-b]pyridazin-3-yl)methoxy]quinoline, Hepatocyte growth factor receptor | | Authors: | Bellon, S.F, Albrecht, B.K, Harmange, J.-C, Bauer, D, Choquette, D, Dussault, I. | | Deposit date: | 2008-02-26 | | Release date: | 2008-04-29 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery and Optimization of Triazolopyridazines as Potent and Selective Inhibitors of the c-Met Kinase.
J.Med.Chem., 51, 2008
|
|
5E06
 
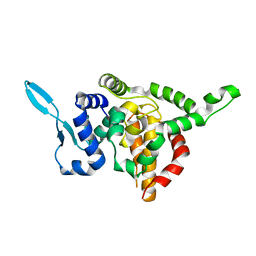 | |
3BDW
 
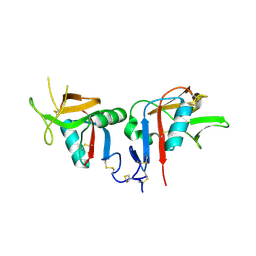 | | Human CD94/NKG2A | | Descriptor: | NKG2-A/NKG2-B type II integral membrane protein, Natural killer cells antigen CD94 | | Authors: | Sullivan, L.C, Clements, C.S. | | Deposit date: | 2007-11-15 | | Release date: | 2008-01-01 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The Heterodimeric Assembly of the CD94-NKG2 Receptor Family and Implications for Human Leukocyte Antigen-E Recognition
Immunity, 27, 2007
|
|
6E3Z
 
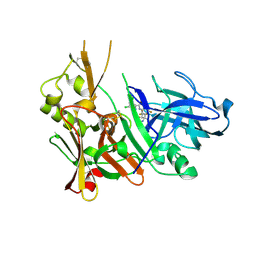 | | Structure of Bace-1 in complex with Ligand 8 | | Descriptor: | Beta-secretase 1, N-{3-[(2R,3R)-5-amino-3-methyl-2-(trifluoromethyl)-3,6-dihydro-2H-1,4-oxazin-3-yl]-4-fluorophenyl}-3,5-dichloropyridine-2-carboxamide | | Authors: | Shaffer, P.L. | | Deposit date: | 2018-07-16 | | Release date: | 2019-09-11 | | Last modified: | 2020-03-25 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Discovery and Chemical Development of JNJ-50138803, a Clinical Candidate BACE1 Inhibitor
Acs Symp.Ser., 1307, 2020
|
|
4EKL
 
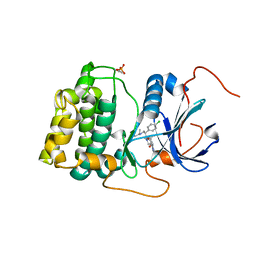 | | Akt1 with GDC0068 | | Descriptor: | (2S)-2-(4-chlorophenyl)-1-{4-[(5R,7R)-7-hydroxy-5-methyl-6,7-dihydro-5H-cyclopenta[d]pyrimidin-4-yl]piperazin-1-yl}-3-(propan-2-ylamino)propan-1-one, RAC-alpha serine/threonine-protein kinase | | Authors: | Wu, W.-I, Vigers, G.P.A, Morales, T.H, Brandhuber, B.J. | | Deposit date: | 2012-04-09 | | Release date: | 2012-05-23 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | An ATP-Site On-Off Switch That Restricts Phosphatase Accessibility of Akt.
Sci.Signal., 5, 2012
|
|
8IZL
 
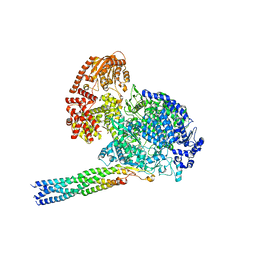 | |
3C5U
 
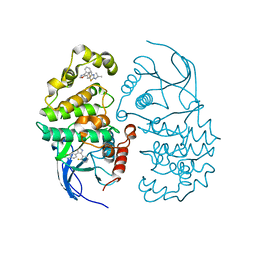 | | P38 ALPHA map kinase complexed with a benzothiazole based inhibitor | | Descriptor: | 6-[4-(2-fluorophenyl)-1,3-oxazol-5-yl]-N-(1-methylethyl)-1,3-benzothiazol-2-amine, Mitogen-activated protein kinase 14 | | Authors: | Sack, J.S. | | Deposit date: | 2008-02-01 | | Release date: | 2008-03-25 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Benzothiazole based inhibitors of p38alpha MAP kinase.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
4NSQ
 
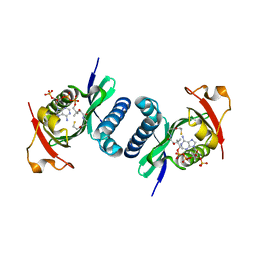 | | Crystal structure of PCAF | | Descriptor: | COENZYME A, Histone acetyltransferase KAT2B | | Authors: | Lin, J.Y, Cai, Y.F. | | Deposit date: | 2013-11-28 | | Release date: | 2014-03-05 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.3108 Å) | | Cite: | Dimeric structure of p300/CBP associated factor.
Bmc Struct.Biol., 14, 2014
|
|
6CF9
 
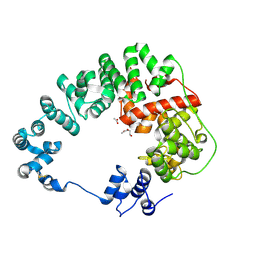 | |
6CFC
 
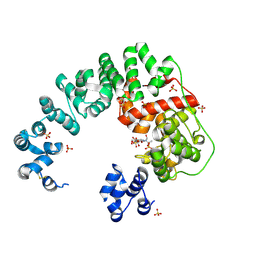 | |
6CF8
 
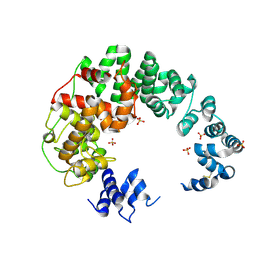 | |
4EKK
 
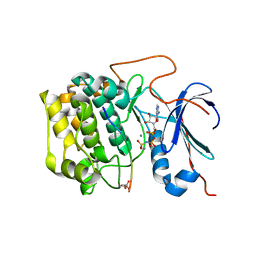 | | Akt1 with AMP-PNP | | Descriptor: | Glycogen synthase kinase-3 beta, MANGANESE (II) ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Wu, W.-I, Vigers, G.P.A, Morales, T.H, Brandhuber, B.J. | | Deposit date: | 2012-04-09 | | Release date: | 2012-05-23 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | An ATP-Site On-Off Switch That Restricts Phosphatase Accessibility of Akt.
Sci.Signal., 5, 2012
|
|
3N1Z
 
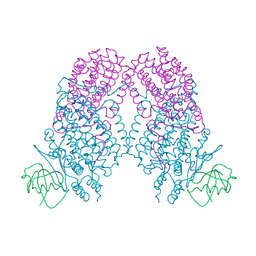 | | X-ray Crystal Structure of Toluene/o-Xylene Monooxygenase Hydroxylase T201S Mutant | | Descriptor: | FE (III) ION, GLYCEROL, HEXAETHYLENE GLYCOL, ... | | Authors: | McCormick, M.S, Sazinsky, M.H, Lippard, S.J. | | Deposit date: | 2010-05-17 | | Release date: | 2010-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Active Site Threonine Facilitates Proton Transfer during Dioxygen Activation at the Diiron Center of Toluene/o-Xylene Monooxygenase Hydroxylase.
J.Am.Chem.Soc., 132, 2010
|
|
3N20
 
 | | X-ray Crystal Structure of Toluene/o-Xylene Monooxygenase Hydroxylase T201V Mutant | | Descriptor: | FE (III) ION, GLYCEROL, HEXAETHYLENE GLYCOL, ... | | Authors: | McCormick, M.S, Sazinsky, M.H, Lippard, S.J. | | Deposit date: | 2010-05-17 | | Release date: | 2010-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Active Site Threonine Facilitates Proton Transfer during Dioxygen Activation at the Diiron Center of Toluene/o-Xylene Monooxygenase Hydroxylase.
J.Am.Chem.Soc., 132, 2010
|
|
6V1N
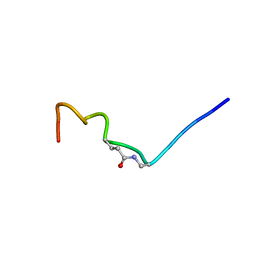 
 | | CSP1-E1A-cyc(Dap6E10) | | Descriptor: | Competence-stimulating peptide type 1 | | Authors: | Yang, Y. | | Deposit date: | 2019-11-20 | | Release date: | 2020-01-08 | | Last modified: | 2020-02-05 | | Method: | SOLUTION NMR | | Cite: | Designing cyclic competence-stimulating peptide (CSP) analogs with pan-group quorum-sensing inhibition activity inStreptococcus pneumoniae.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
3N1X
 
 | | X-ray Crystal Structure of Toluene/o-Xylene Monooxygenase Hydroxylase T201C Mutant | | Descriptor: | 1,2-ETHANEDIOL, FE (III) ION, HYDROXIDE ION, ... | | Authors: | Sazinsky, M.H, McCormick, M.S, Lippard, S.J. | | Deposit date: | 2010-05-17 | | Release date: | 2010-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Active Site Threonine Facilitates Proton Transfer during Dioxygen Activation at the Diiron Center of Toluene/o-Xylene Monooxygenase Hydroxylase.
J.Am.Chem.Soc., 132, 2010
|
|
3N1Y
 
 | | X-ray Crystal Structure of Toluene/o-Xylene Monooxygenase Hydroxylase T201G Mutant | | Descriptor: | FE (III) ION, GLYCEROL, HEXAETHYLENE GLYCOL, ... | | Authors: | McCormick, M.S, Sazinsky, M.H, Lippard, S.J. | | Deposit date: | 2010-05-17 | | Release date: | 2010-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Active Site Threonine Facilitates Proton Transfer during Dioxygen Activation at the Diiron Center of Toluene/o-Xylene Monooxygenase Hydroxylase.
J.Am.Chem.Soc., 132, 2010
|
|
