8ASN
 
 | |
5HKG
 
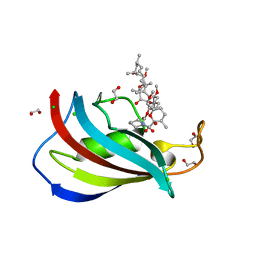 | | Total chemical synthesis, refolding and crystallographic structure of a fully active immunophilin: calstabin 2 (FKBP12.6). | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Peptidyl-prolyl cis-trans isomerase FKBP1B, ... | | Authors: | Sirigu, S, Huet, T, Bacchi, M, Jullian, M, Fould, B, Ferry, G, Vuillard, L, Chavas, L, Puget, K, Nosjean, O, Boutin, J.A. | | Deposit date: | 2016-01-14 | | Release date: | 2016-10-05 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Total chemical synthesis, refolding, and crystallographic structure of fully active immunophilin calstabin 2 (FKBP12.6).
Protein Sci., 25, 2016
|
|
3DEL
 
 | |
8QEL
 
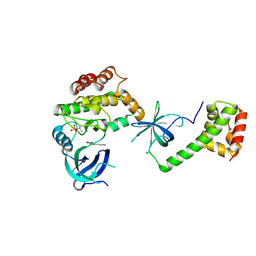 | | PKR kinase domain- eIF2alpha in complex with compound | | Descriptor: | (3~{Z})-3-[(4-methyl-1~{H}-imidazol-5-yl)methylidene]-2-oxidanylidene-1~{H}-indole-5-carboxamide, Eukaryotic translation initiation factor 2 subunit alpha, Interferon-induced, ... | | Authors: | Nawrotek, A, Vuillard, L, Miallau, L. | | Deposit date: | 2023-08-31 | | Release date: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.451 Å) | | Cite: | PKR kinase domain- eIF2alpha in complex with compound
To Be Published
|
|
4DM5
 
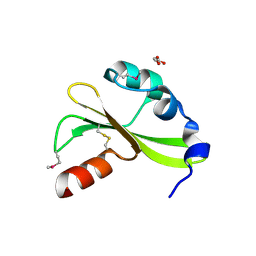 | |
8A5J
 
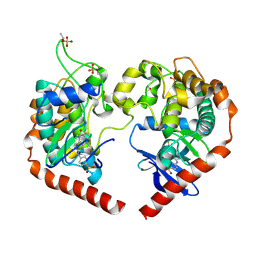 | | Crystal structure of Human STE20-like kinase 1, MST1 in complex with compound XMU-MP-1 | | Descriptor: | 4-[(5,10-dimethyl-6-oxo-6,10-dihydro-5H-pyrimido[5,4-b]thieno[3,2-e][1,4]diazepin-2-yl)amino]benzenesulfonamide, Serine/threonine-protein kinase 4 37kDa subunit | | Authors: | Nawrotek, A, Vuillard, L, Miallau, L. | | Deposit date: | 2022-06-15 | | Release date: | 2022-07-13 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.123 Å) | | Cite: | Crystal structure of the Kelch domain of human Keap1in complex with ligand S217879
To Be Published
|
|
8A66
 
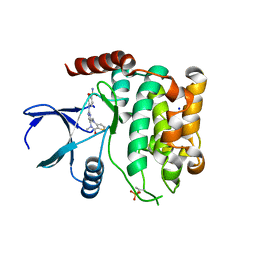 | | Crystal structure of MST2 in complex with XMU-MP-1 | | Descriptor: | 4-[(5,10-dimethyl-6-oxo-6,10-dihydro-5H-pyrimido[5,4-b]thieno[3,2-e][1,4]diazepin-2-yl)amino]benzenesulfonamide, SODIUM ION, Serine/threonine-protein kinase 3 36kDa subunit | | Authors: | Nawrotek, A, Vuillard, L, Miallau, L, Weber, C. | | Deposit date: | 2022-06-16 | | Release date: | 2022-07-20 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.901 Å) | | Cite: | Crystal structure of the Kelch domain of human Keap1in complex with ligand S217879
To Be Published
|
|
7OTS
 
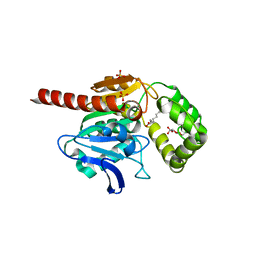 | | Crystal structure of human Monoacylglycerol Lipase ABHD6 in complex with oleic acid and octyl glucoside | | Descriptor: | GLYCEROL, Monoacylglycerol lipase ABHD6, OLEIC ACID, ... | | Authors: | Nawrotek, A, Talagas, A, Vuillard, L, Miallau, L. | | Deposit date: | 2021-06-10 | | Release date: | 2021-06-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.792 Å) | | Cite: | Crystal structure of human Monoacylglycerol Lipase ABHD6 in complex with oleic acid and octyl glucoside
To Be Published
|
|
3N26
 
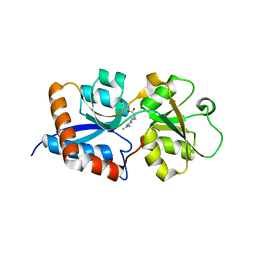 | | Cpn0482 : the arginine binding protein from the periplasm of chlamydia Pneumoniae | | Descriptor: | ARGININE, Amino acid ABC transporter, periplasmic amino acid-binding protein | | Authors: | Petit, P, Garcia, C, Vuillard, L, Soriani, M, Grandi, G, Marseilles Structural Genomics Program AFMB (MSGP), Marseilles Structural Genomics Program @ AFMB (MSGP) | | Deposit date: | 2010-05-17 | | Release date: | 2010-06-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Exploiting antigenic diversity for vaccine design: the Chlamydia ArtJ paradigm.
J.Biol.Chem., 2010
|
|
4NO7
 
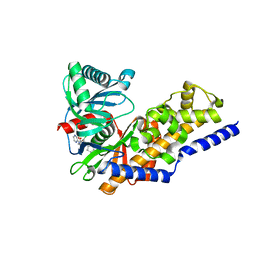 | | Human Glucokinase in complex with a nanomolar activator. | | Descriptor: | (2R)-2-[3-chloro-4-(methylsulfonyl)phenyl]-3-[(1R)-3-oxocyclopentyl]-N-(pyrazin-2-yl)propanamide, Glucokinase, alpha-D-glucopyranose | | Authors: | Petit, P, Ferry, G, Antoine, M, Boutin, J.A, Kotschy, A, Perron-Sierra, F, Mamelli, L, Vuillard, L. | | Deposit date: | 2013-11-19 | | Release date: | 2014-10-29 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The active conformation of human glucokinase is not altered by allosteric activators.
Acta Crystallogr. D Biol. Crystallogr., 67, 2011
|
|
3ID8
 
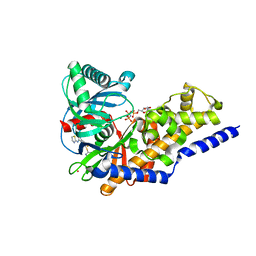 | | Ternary complex of human pancreatic glucokinase crystallized with activator, glucose and AMP-PNP | | Descriptor: | 2-AMINO-4-FLUORO-5-[(1-METHYL-1H-IMIDAZOL-2-YL)SULFANYL]-N-(1,3-THIAZOL-2-YL)BENZAMIDE, Glucokinase, MAGNESIUM ION, ... | | Authors: | Petit, P, Gluais, L, Lagarde, A, Boutin, J.A, Ferry, G, Vuillard, L. | | Deposit date: | 2009-07-20 | | Release date: | 2010-07-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The active conformation of human glucokinase is not altered by allosteric activators
Acta Crystallogr.,Sect.D, 67, 2011
|
|
4NNZ
 
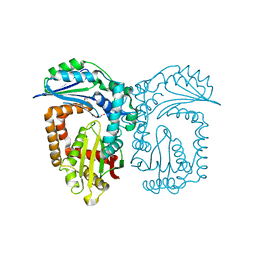 | |
3IDH
 
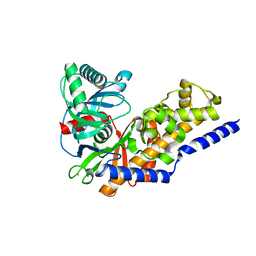 | | Human pancreatic glucokinase in complex with glucose | | Descriptor: | Glucokinase, POTASSIUM ION, alpha-D-glucopyranose | | Authors: | Petit, P, Gluais, L, Lagarde, A, Boutin, J.A, Ferry, G, Vuillard, L. | | Deposit date: | 2009-07-21 | | Release date: | 2010-07-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | The active conformation of human glucokinase is not altered by allosteric activators
Acta Crystallogr.,Sect.D, 67, 2011
|
|
3F9M
 
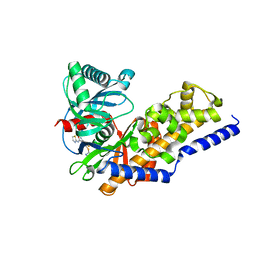 | | Human pancreatic glucokinase in complex with glucose and activator showing a mobile flap | | Descriptor: | 2-AMINO-4-FLUORO-5-[(1-METHYL-1H-IMIDAZOL-2-YL)SULFANYL]-N-(1,3-THIAZOL-2-YL)BENZAMIDE, Glucokinase, alpha-D-glucopyranose | | Authors: | Petit, P, Gluais, L, Lagarde, A, Vuillard, L, Boutin, J.A, Ferry, G. | | Deposit date: | 2008-11-14 | | Release date: | 2008-12-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The active conformation of human glucokinase is not altered by allosteric activators
Acta Crystallogr.,Sect.D, 67, 2011
|
|
3FGU
 
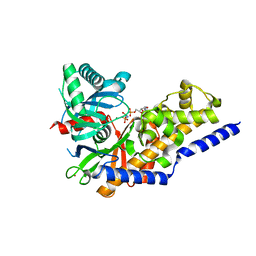 | | Catalytic complex of Human Glucokinase | | Descriptor: | Glucokinase, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Petit, P, Lagarde, A, Boutin, J.A, Ferry, G, Vuillard, L. | | Deposit date: | 2008-12-08 | | Release date: | 2009-12-15 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | The active conformation of human glucokinase is not altered by allosteric activators
Acta Crystallogr.,Sect.D, 67, 2011
|
|
8AXZ
 
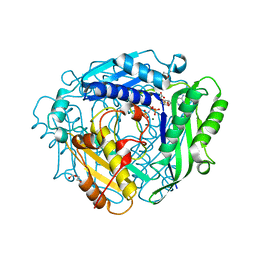 | | Crystal structure of human methionine adenosyltransferase 2A (MAT2A) in complex with S-adenosylmethionine, adenosin and diphosphono-aminophosphonic acid. | | Descriptor: | (DIPHOSPHONO)AMINOPHOSPHONIC ACID, ADENOSINE, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Nawrotek, A, Vuillard, L, Miallau, L. | | Deposit date: | 2022-09-01 | | Release date: | 2022-10-05 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.154 Å) | | Cite: | Crystal structure of human methionine adenosyltransferase 2A (MAT2A) in complex with S-adenosylmethionine, adenosin and diphosphono-aminophosphonic acid.
To Be Published
|
|
8B2J
 
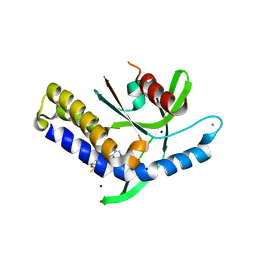 | | Crystal structure of human STING in complex with ADU-S100 | | Descriptor: | (1~{R},3~{S},6~{R},8~{R},9~{R},10~{S},12~{S},15~{R},17~{R},18~{R})-8,17-bis(6-aminopurin-9-yl)-3,12-bis(oxidanylidene)-3,12-bis(sulfanyl)-2,4,7,11,13,16-hexaoxa-3$l^{5},12$l^{5}-diphosphatricyclo[13.2.1.0^{6,10}]octadecane-9,18-diol, CALCIUM ION, SODIUM ION, ... | | Authors: | Nawrotek, A, Vuillard, L, Miallau, L. | | Deposit date: | 2022-09-14 | | Release date: | 2022-11-30 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.174 Å) | | Cite: | Crystal structure of human STING in complex with ADU-S100
To Be Published
|
|
1W50
 
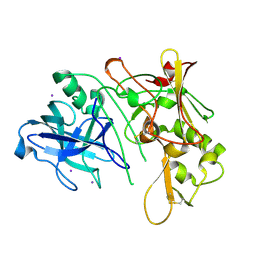 | | Apo Structure of BACE (Beta Secretase) | | Descriptor: | BETA-SECRETASE 1, IODIDE ION | | Authors: | Patel, S, Vuillard, L, Cleasby, A, Murray, C.W, Yon, J. | | Deposit date: | 2004-08-04 | | Release date: | 2004-09-23 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Apo and Inhibitor Complex Structures of Bace (Beta-Secretase)
J.Mol.Biol., 343, 2004
|
|
1W51
 
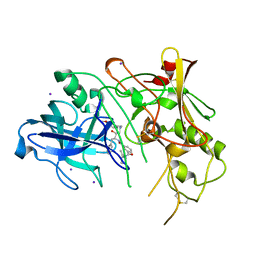 | | BACE (Beta Secretase) in complex with a nanomolar non-peptidic inhibitor | | Descriptor: | 3-[({(1S,2R)-1-BENZYL-2-HYDROXY-3-[(3-METHOXYBENZYL)AMINO]PROPYL}AMINO)(HYDROXY)METHYL]-N,N-DIPROPYLBENZAMIDE, BETA-SECRETASE 1, IODIDE ION | | Authors: | Patel, S, Vuillard, L, Cleasby, A, Murray, C.W, Yon, J. | | Deposit date: | 2004-08-04 | | Release date: | 2004-09-23 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Apo and Inhibitor Complex Structures of Bace (Beta-Secretase)
J.Mol.Biol., 343, 2004
|
|
8A46
 
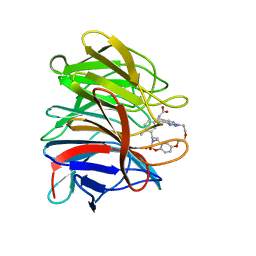 | | Crystal structure of the human Kelch domain of Keap1 in complex with compound S217879 | | Descriptor: | 2-[(1S,2R,8S)-2,4,32-trimethyl-28,28-bis(oxidanylidene)-19,22,27-trioxa-28$l^{6}-thia-1,14,15,16-tetrazahexacyclo[21.5.3.1^{3,7}.1^{9,13}.0^{12,16}.0^{26,30}]tritriaconta-3(33),4,6,9(32),10,12,14,23,25,30-decaen-8-yl]ethanoic acid, Kelch-like ECH-associated protein 1 | | Authors: | Weber, C, Vuillard, L, Delerive, P, Miallau, L. | | Deposit date: | 2022-06-10 | | Release date: | 2022-07-13 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.323 Å) | | Cite: | Selective disruption of NRF2-KEAP1 interaction leads to NASH resolution and reduction of liver fibrosis in mice.
JHEP Rep, 5, 2023
|
|
7Z4V
 
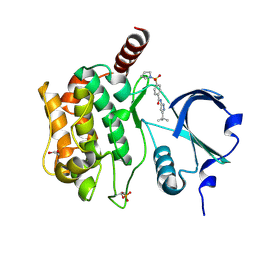 | | Structure of Serine-Threonine kinase STK25 in complex with compound | | Descriptor: | 1,2-ETHANEDIOL, Serine/threonine-protein kinase 25, ~{N}-(5-~{tert}-butyl-1~{H}-pyrazol-3-yl)-4-pyrrolidin-1-ylsulfonyl-benzamide | | Authors: | Nawrotek, A, Vuillard, L, Miallau, L. | | Deposit date: | 2022-03-04 | | Release date: | 2022-04-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.644 Å) | | Cite: | Targeting non-alcoholic fatty liver disease: Design, X-ray co-crystal structure and synthesis of 'first-in-kind' inhibitors of serine/threonine kinase25.
Bioorg.Med.Chem.Lett., 75, 2022
|
|
4XH9
 
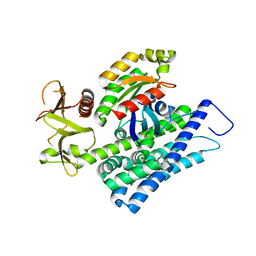 | | CRYSTAL STRUCTURE OF HUMAN RHOA IN COMPLEX WITH DH/PH FRAGMENT OF THE GUANINE NUCLEOTIDE EXCHANGE FACTOR NET1 | | Descriptor: | Neuroepithelial cell-transforming gene 1 protein, Transforming protein RhoA | | Authors: | Garcia, C, Petit, P, Boutin, J.A, Ferry, G, Vuillard, L. | | Deposit date: | 2015-01-05 | | Release date: | 2015-01-14 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A structural study of the complex between neuroepithelial cell transforming gene 1 (Net1) and RhoA reveals a potential anticancer drug hot spot.
J. Biol. Chem., 293, 2018
|
|
6DO6
 | | NMR solution structure of wild type apo hFABP1 at 308 K | | Descriptor: | Fatty acid-binding protein, liver | | Authors: | Scanlon, M.J, Mohanty, B, Doak, B.C, Patil, R. | | Deposit date: | 2018-06-09 | | Release date: | 2018-12-26 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A ligand-induced structural change in fatty acid-binding protein 1 is associated with potentiation of peroxisome proliferator-activated receptor alpha agonists.
J. Biol. Chem., 294, 2019
|
|
6DRG
 | | NMR solution structure of wild type hFABP1 with GW7647 | | Descriptor: | 2-[(4-{2-[(4-cyclohexylbutyl)(cyclohexylcarbamoyl)amino]ethyl}phenyl)sulfanyl]-2-methylpropanoic acid, Fatty acid-binding protein, liver | | Authors: | Scanlon, M.J, Mohanty, B, Doak, B.C, Patil, R. | | Deposit date: | 2018-06-11 | | Release date: | 2018-12-26 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A ligand-induced structural change in fatty acid-binding protein 1 is associated with potentiation of peroxisome proliferator-activated receptor alpha agonists.
J. Biol. Chem., 294, 2019
|
|
6DO7
 
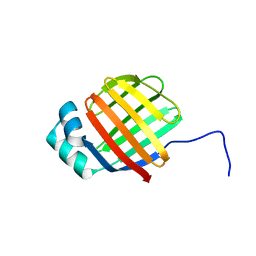 | | NMR solution structure of wild type hFABP1 with GW7647 | | Descriptor: | Fatty acid-binding protein, liver | | Authors: | Scanlon, M.J, Mohanty, B, Doak, B.C, Patil, R. | | Deposit date: | 2018-06-09 | | Release date: | 2019-01-02 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A ligand-induced structural change in fatty acid-binding protein 1 is associated with potentiation of peroxisome proliferator-activated receptor alpha agonists.
J. Biol. Chem., 294, 2019
|
|
