4MJR
 
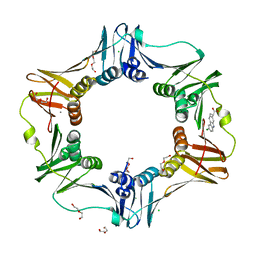 | | E. coli sliding clamp in complex with (S)-Carprofen | | Descriptor: | (2S)-2-(6-chloro-9H-carbazol-2-yl)propanoic acid, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Yin, Z, Oakley, A.J. | | Deposit date: | 2013-09-04 | | Release date: | 2013-09-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | DNA replication is the target for the antibacterial effects of nonsteroidal anti-inflammatory drugs.
Chem.Biol., 21, 2014
|
|
4MJP
 
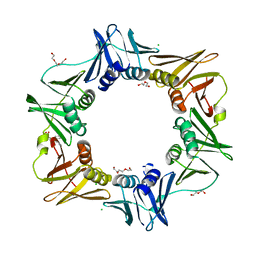 | | E. coli sliding clamp in complex with (R)-Vedaprofen | | Descriptor: | (2R)-2-(4-cyclohexylnaphthalen-1-yl)propanoic acid, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Yin, Z, Oakley, A.J. | | Deposit date: | 2013-09-04 | | Release date: | 2013-09-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.855 Å) | | Cite: | DNA replication is the target for the antibacterial effects of nonsteroidal anti-inflammatory drugs.
Chem.Biol., 21, 2014
|
|
4MJQ
 
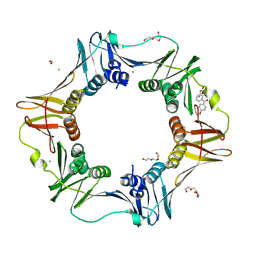 | | E. coli sliding clamp in complex with Bromfenac | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Yin, Z, Oakley, A.J. | | Deposit date: | 2013-09-04 | | Release date: | 2013-09-18 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | DNA replication is the target for the antibacterial effects of nonsteroidal anti-inflammatory drugs.
Chem.Biol., 21, 2014
|
|
4O04
 
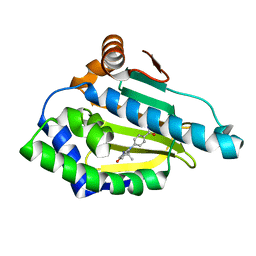 | | Identification of novel HSP90/isoform selective inhibitors using structure-based drug design. Demonstration of potential utility in treating CNS disorders such as Huntington's disease | | Descriptor: | 4-(2,7,7-trimethyl-5-oxo-1,2,3,4,5,6,7,8-octahydro-9H-beta-carbolin-9-yl)benzamide, Heat shock protein HSP 90-alpha | | Authors: | Zuccola, H.J, Ernst, J. | | Deposit date: | 2013-12-13 | | Release date: | 2014-12-24 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Identification of Novel HSP90 alpha / beta Isoform Selective Inhibitors Using Structure-Based Drug Design. Demonstration of Potential Utility in Treating CNS Disorders such as Huntington's Disease.
J.Med.Chem., 57, 2014
|
|
4O0B
 
 | | Identification of novel HSP90/isoform selective inhibitors using structure-based drug design. Demonstration of potential utility in treating CNS disorders such as Huntington's disease | | Descriptor: | 8-cyclopentyl-6-(3,6,6-trimethyl-4-oxo-4,5,6,7-tetrahydro-1H-indol-1-yl)-3,4-dihydroisoquinolin-1(2H)-one, Heat shock protein HSP 90-alpha | | Authors: | Zuccola, H.J, Ernst, J.T. | | Deposit date: | 2013-12-13 | | Release date: | 2014-04-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Identification of Novel HSP90 alpha / beta Isoform Selective Inhibitors Using Structure-Based Drug Design. Demonstration of Potential Utility in Treating CNS Disorders such as Huntington's Disease.
J.Med.Chem., 57, 2014
|
|
4O05
 
 | | Identification of novel HSP90/isoform selective inhibitors using structure-based drug design. Demonstration of potential utility in treating CNS disorders such as Huntington's disease | | Descriptor: | 2,7,7-trimethyl-9-[1-oxo-8-(propan-2-ylamino)-1,2,3,4-tetrahydroisoquinolin-6-yl]-1,2,3,4,6,7,8,9-octahydro-5H-beta-carbolin-5-one, Heat shock protein HSP 90-alpha | | Authors: | Zuccola, H.J, Ernst, J.T. | | Deposit date: | 2013-12-13 | | Release date: | 2014-04-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Identification of Novel HSP90 alpha / beta Isoform Selective Inhibitors Using Structure-Based Drug Design. Demonstration of Potential Utility in Treating CNS Disorders such as Huntington's Disease.
J.Med.Chem., 57, 2014
|
|
4O09
 
 | | Identification of novel HSP90 / isoform selective inhibitors using structure-based drug design. Demonstration of potential utility in treating CNS disorders such as Huntington s disease | | Descriptor: | 8-(2-methylpropyl)-6-(3,6,6-trimethyl-4-oxo-4,5,6,7-tetrahydro-1H-indol-1-yl)-3,4-dihydroisoquinolin-1(2H)-one, Heat shock protein HSP 90-alpha | | Authors: | Zuccola, H.J, Ernst, J.T. | | Deposit date: | 2013-12-13 | | Release date: | 2014-04-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Identification of Novel HSP90 alpha / beta Isoform Selective Inhibitors Using Structure-Based Drug Design. Demonstration of Potential Utility in Treating CNS Disorders such as Huntington's Disease.
J.Med.Chem., 57, 2014
|
|
4ER1
 
 | | THE ACTIVE SITE OF ASPARTIC PROTEINASES | | Descriptor: | ENDOTHIAPEPSIN, N-[(1R,2R,4R)-1-(cyclohexylmethyl)-2-hydroxy-6-methyl-4-{[(2R)-2-methylbutyl]carbamoyl}heptyl]-3-(1H-imidazol-3-ium-4-yl)-N~2~-[3-naphthalen-1-yl-2-(naphthalen-1-ylmethyl)propanoyl]-L-alaninamide | | Authors: | Quail, J.W, Cooper, J.B, Szelke, M, Blundell, T.L. | | Deposit date: | 1990-10-14 | | Release date: | 1991-01-15 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The active site of aspartic proteinases
FEBS Lett., 174, 1984
|
|
4NH9
 
 | | Correlation between chemotype-dependent binding conformations of HSP90 alpha/beta and isoform selectivity | | Descriptor: | 2-fluoro-6-[(3S)-tetrahydrofuran-3-ylamino]-4-(3,6,6-trimethyl-4-oxo-4,5,6,7-tetrahydro-1H-indol-1-yl)benzamide, Endoplasmin | | Authors: | Zuccola, H.J, Ernst, J.T. | | Deposit date: | 2013-11-04 | | Release date: | 2014-01-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.77 Å) | | Cite: | Correlation between chemotype-dependent binding conformations of HSP90 alpha / beta and isoform selectivity-Implications for the structure-based design of HSP90 alpha / beta selective inhibitors for treating neurodegenerative diseases.
Bioorg.Med.Chem.Lett., 24, 2014
|
|
4N94
 
 | | E. coli sliding clamp in complex with 3,4-difluorobenzamide | | Descriptor: | 1,2-ETHANEDIOL, 3,4-difluorobenzamide, CALCIUM ION, ... | | Authors: | Yin, Z, Oakley, A.J. | | Deposit date: | 2013-10-19 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Discovery of lead compounds targeting the bacterial sliding clamp using a fragment-based approach.
J.Med.Chem., 57, 2014
|
|
4NH7
 
 | | Correlation between chemotype-dependent binding conformations of HSP90 alpha/beta and isoform selectivity | | Descriptor: | 4-[6,6-dimethyl-4-oxidanylidene-3-(trifluoromethyl)-5,7-dihydroindazol-1-yl]-2-[(4-oxidanylcyclohexyl)amino]benzamide, GLYCEROL, Heat shock protein HSP 90-alpha | | Authors: | Zuccola, H.J, Ernst, J. | | Deposit date: | 2013-11-04 | | Release date: | 2014-01-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Correlation between chemotype-dependent binding conformations of HSP90 alpha / beta and isoform selectivity-Implications for the structure-based design of HSP90 alpha / beta selective inhibitors for treating neurodegenerative diseases.
Bioorg.Med.Chem.Lett., 24, 2014
|
|
4N98
 
 | |
4N9A
 
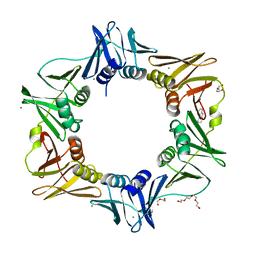 | | E. coli sliding clamp in complex with (R)-6-chloro-2,3,4,9-tetrahydro-1H-carbazole-1-carboxylic acid | | Descriptor: | (1R)-6-chloro-2,3,4,9-tetrahydro-1H-carbazole-1-carboxylic acid, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Yin, Z, Oakley, A.J. | | Deposit date: | 2013-10-19 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Discovery of lead compounds targeting the bacterial sliding clamp using a fragment-based approach.
J.Med.Chem., 57, 2014
|
|
4O07
 
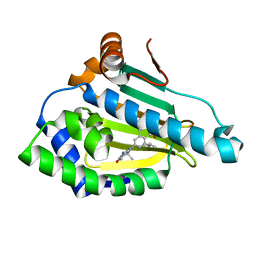 | | Identification of novel HSP90/isoform selective inhibitors using structure-based drug design. Demonstration of potential utility in treating CNS disorders such as Huntington's disease | | Descriptor: | 2,7,7-trimethyl-9-[8-(2-methylpropyl)-1-oxo-1,2,3,4-tetrahydroisoquinolin-6-yl]-1,2,3,4,6,7,8,9-octahydro-5H-beta-carbolin-5-one, Heat shock protein HSP 90-alpha | | Authors: | Zuccola, H.J, Ernst, J.T. | | Deposit date: | 2013-12-13 | | Release date: | 2014-04-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Identification of Novel HSP90 alpha / beta Isoform Selective Inhibitors Using Structure-Based Drug Design. Demonstration of Potential Utility in Treating CNS Disorders such as Huntington's Disease.
J.Med.Chem., 57, 2014
|
|
4N97
 
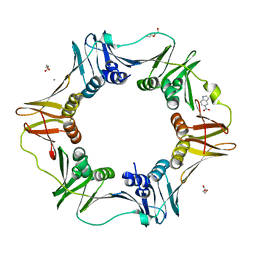 | | E. coli sliding clamp in complex with 5-nitroindole | | Descriptor: | 5-nitro-1H-indole, CALCIUM ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Yin, Z, Oakley, A.J. | | Deposit date: | 2013-10-19 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Discovery of lead compounds targeting the bacterial sliding clamp using a fragment-based approach.
J.Med.Chem., 57, 2014
|
|
4N99
 
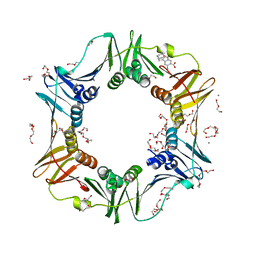 | | E. coli sliding clamp in complex with 6-chloro-2,3,4,9-tetrahydro-1H-carbazole-7-carboxylic acid | | Descriptor: | 6-chloro-2,3,4,9-tetrahydro-1H-carbazole-7-carboxylic acid, CALCIUM ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Yin, Z, Oakley, A.J. | | Deposit date: | 2013-10-19 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Discovery of lead compounds targeting the bacterial sliding clamp using a fragment-based approach.
J.Med.Chem., 57, 2014
|
|
4NH8
 | | Correlation between chemotype-dependent binding conformations of HSP90 alpha/beta and isoform selectivity | | Descriptor: | 2-fluoro-6-[(3S)-tetrahydrofuran-3-ylamino]-4-(3,6,6-trimethyl-4-oxo-4,5,6,7-tetrahydro-1H-indol-1-yl)benzamide, Heat shock protein HSP 90-alpha | | Authors: | Ernst, J.T, Zuccola, H.J. | | Deposit date: | 2013-11-04 | | Release date: | 2014-01-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Correlation between chemotype-dependent binding conformations of HSP90 alpha / beta and isoform selectivity-Implications for the structure-based design of HSP90 alpha / beta selective inhibitors for treating neurodegenerative diseases.
Bioorg.Med.Chem.Lett., 24, 2014
|
|
4N95
 
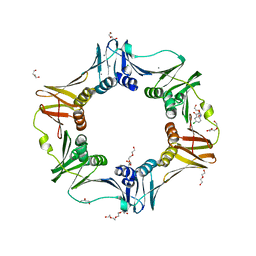 | | E. coli sliding clamp in complex with 5-chloroindoline-2,3-dione | | Descriptor: | 5-chloro-1H-indole-2,3-dione, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Yin, Z, Oakley, A.J. | | Deposit date: | 2013-10-19 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Discovery of lead compounds targeting the bacterial sliding clamp using a fragment-based approach.
J.Med.Chem., 57, 2014
|
|
7CSZ
 
 | | Crystal structure of the N-terminal tandem RRM domains of RBM45 in complex with single-stranded DNA | | Descriptor: | DNA (5'-D(*CP*GP*AP*CP*GP*GP*GP*AP*CP*GP*C)-3'), RNA-binding protein 45 | | Authors: | Chen, X, Yang, Z, Wang, W, Wang, M. | | Deposit date: | 2020-08-17 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for RNA recognition by the N-terminal tandem RRM domains of human RBM45.
Nucleic Acids Res., 49, 2021
|
|
7CSX
 
 | |
4N96
 
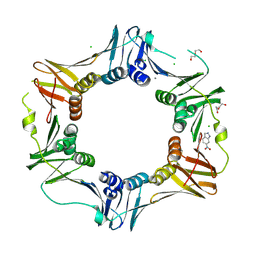 | | E. coli sliding clamp in complex with 6-nitroindazole | | Descriptor: | 6-NITROINDAZOLE, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Yin, Z, Oakley, A.J. | | Deposit date: | 2013-10-19 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Discovery of lead compounds targeting the bacterial sliding clamp using a fragment-based approach.
J.Med.Chem., 57, 2014
|
|
7D4F
 
 | | Structure of COVID-19 RNA-dependent RNA polymerase bound to suramin | | Descriptor: | 8-(3-(3-aminobenzamido)-4-methylbenzamido)naphthalene-1,3,5-trisulfonic acid, Non-structural protein 7, Non-structural protein 8, ... | | Authors: | Li, Z, Yin, W, Zhou, Z, Yu, X, Xu, H. | | Deposit date: | 2020-09-23 | | Release date: | 2020-11-11 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (2.57 Å) | | Cite: | Structural basis for inhibition of the SARS-CoV-2 RNA polymerase by suramin.
Nat.Struct.Mol.Biol., 28, 2021
|
|
7WFC
 
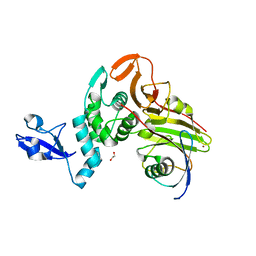 | | X-ray structure of HKU1-PLP2(Cys109Ser) catalytic mutant in complex with free ubiquitin | | Descriptor: | 1,2-ETHANEDIOL, 60S ribosomal protein L40, Papain-like protease, ... | | Authors: | Xiong, Y.X, Fu, Z.Y, Huang, H. | | Deposit date: | 2021-12-26 | | Release date: | 2022-12-28 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The substrate selectivity of papain-like proteases from human-infecting coronaviruses correlates with innate immune suppression.
Sci.Signal., 16, 2023
|
|
5XAF
 
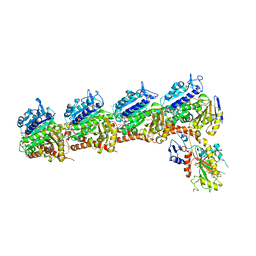 | | Crystal structure of tubulin-stathmin-TTL-Compound Z1 complex | | Descriptor: | (3S,4R)-4-(3-hydroxy-4-methoxyphenyl)-3-methyl-1-(3,4,5-trimethoxyphenyl)azetidin-2-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CALCIUM ION, ... | | Authors: | Zhang, H, Luo, C, Wang, Y. | | Deposit date: | 2017-03-12 | | Release date: | 2017-12-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.551 Å) | | Cite: | Design, synthesis, biological evaluation and cocrystal structures with tubulin of chiral beta-lactam bridged combretastatin A-4 analogues as potent antitumor agents
Eur J Med Chem, 144, 2017
|
|
5XPD
 
 | |
