6BTP
 
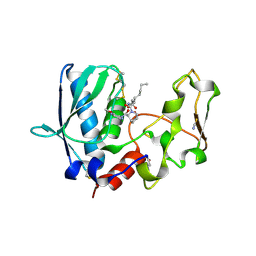 | | BMP1 complexed with a hydroxamate | | Descriptor: | (1R,3S,4S)-2-{(2R)-2-[2-(hydroxyamino)-2-oxoethyl]heptanoyl}-N-(3-methoxypyrazin-2-yl)-2-azabicyclo[2.2.1]heptane-3-carboxamide, Bone morphogenetic protein 1, THIOCYANATE ION, ... | | Authors: | Gampe, R, Shewchuk, L. | | Deposit date: | 2017-12-07 | | Release date: | 2018-08-08 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Reverse Hydroxamate Inhibitors of Bone Morphogenetic Protein 1.
ACS Med Chem Lett, 9, 2018
|
|
3N45
 
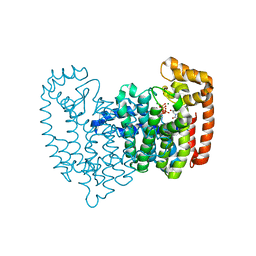 | | Human FPPS complex with FBS_04 and zoledronic acid/MG2+ | | Descriptor: | (2S)-1-[(benzyloxy)carbonyl]-2,3-dihydro-1H-indole-2-carboxylic acid, FARNESYL PYROPHOSPHATE SYNTHASE, MAGNESIUM ION, ... | | Authors: | Rondeau, J.-M. | | Deposit date: | 2010-05-21 | | Release date: | 2010-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Allosteric non-bisphosphonate FPPS inhibitors identified by fragment-based discovery.
Nat.Chem.Biol., 6, 2010
|
|
3N5H
 
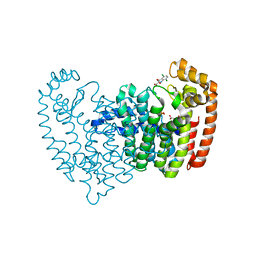 | | Human fpps complex with NOV_304 | | Descriptor: | 3-(carboxymethyl)-4,7-dichloro-1H-indole-2-carboxylic acid, FARNESYL PYROPHOSPHATE SYNTHASE, PHOSPHATE ION | | Authors: | Rondeau, J.-M. | | Deposit date: | 2010-05-25 | | Release date: | 2010-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Allosteric non-bisphosphonate FPPS inhibitors identified by fragment-based discovery.
Nat.Chem.Biol., 6, 2010
|
|
3N6K
 
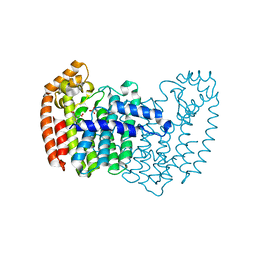 | | Human FPPS complex with NOV_823 | | Descriptor: | 1-(carboxymethyl)-1H-benzo[g]indole-2-carboxylic acid, FARNESYL PYROPHOSPHATE SYNTHASE, PHOSPHATE ION | | Authors: | Rondeau, J.-M. | | Deposit date: | 2010-05-26 | | Release date: | 2010-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Allosteric non-bisphosphonate FPPS inhibitors identified by fragment-based discovery.
Nat.Chem.Biol., 6, 2010
|
|
3N5J
 
 | | Human fpps complex with NOV_311 | | Descriptor: | 3-(carboxymethyl)-5,7-dichloro-1H-indole-2-carboxylic acid, FARNESYL PYROPHOSPHATE SYNTHASE, PHOSPHATE ION | | Authors: | Rondeau, J.-M. | | Deposit date: | 2010-05-25 | | Release date: | 2010-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Allosteric non-bisphosphonate FPPS inhibitors identified by fragment-based discovery.
Nat.Chem.Biol., 6, 2010
|
|
6BSL
 
 | | BMP1 complexed with a reverse hydroxymate - compound 22 | | Descriptor: | 1,2-ETHANEDIOL, Bone morphogenetic protein 1, N-(2-ethoxy-4-{5-[({[(2R)-2-{(1R)-1-[formyl(hydroxy)amino]propyl}heptanoyl]amino}methyl)carbamoyl]furan-2-yl}benzene-1-carbonyl)-L-aspartic acid, ... | | Authors: | Gampe, R, Shewchuk, L. | | Deposit date: | 2017-12-04 | | Release date: | 2018-08-08 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Reverse Hydroxamate Inhibitors of Bone Morphogenetic Protein 1.
ACS Med Chem Lett, 9, 2018
|
|
3N1V
 
 | | Human FPPS COMPLEX WITH FBS_01 | | Descriptor: | (5-chloro-3-methyl-1-benzothiophen-2-yl)acetic acid, FARNESYL PYROPHOSPHATE SYNTHASE, PHOSPHATE ION | | Authors: | Rondeau, J.-M. | | Deposit date: | 2010-05-17 | | Release date: | 2010-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Allosteric non-bisphosphonate FPPS inhibitors identified by fragment-based discovery.
Nat.Chem.Biol., 6, 2010
|
|
3N49
 
 | | Human FPPS COMPLEX WITH NOV_292 | | Descriptor: | FARNESYL PYROPHOSPHATE SYNTHASE, PHOSPHATE ION, naphtho[2,1-b]thiophen-1-ylacetic acid | | Authors: | Rondeau, J.-M. | | Deposit date: | 2010-05-21 | | Release date: | 2010-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Allosteric non-bisphosphonate FPPS inhibitors identified by fragment-based discovery.
Nat.Chem.Biol., 6, 2010
|
|
3N3L
 
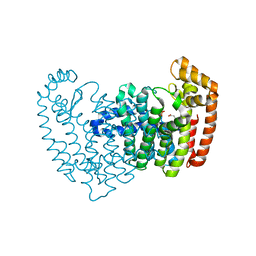 | | Human FPPS complex with FBS_03 | | Descriptor: | (6-methoxy-1-benzofuran-3-yl)acetic acid, FARNESYL PYROPHOSPHATE SYNTHASE, PHOSPHATE ION | | Authors: | Rondeau, J.-M. | | Deposit date: | 2010-05-20 | | Release date: | 2010-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.74 Å) | | Cite: | Allosteric non-bisphosphonate FPPS inhibitors identified by fragment-based discovery.
Nat.Chem.Biol., 6, 2010
|
|
3N46
 
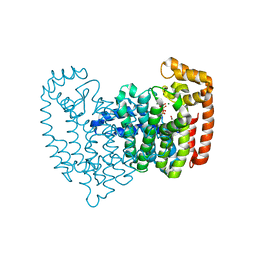 | | Human FPPS complex with NOV_980 and zoledronic acid/MG2+ | | Descriptor: | (2R)-1-[(benzyloxy)carbonyl]-2,3-dihydro-1H-indole-2-carboxylic acid, FARNESYL PYROPHOSPHATE SYNTHASE, MAGNESIUM ION, ... | | Authors: | Rondeau, J.-M. | | Deposit date: | 2010-05-21 | | Release date: | 2010-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Allosteric non-bisphosphonate FPPS inhibitors identified by fragment-based discovery.
Nat.Chem.Biol., 6, 2010
|
|
3N1W
 
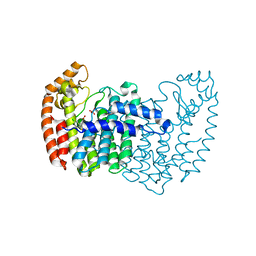 | | Human FPPS COMPLEX WITH FBS_02 | | Descriptor: | (5-chloro-1-benzothiophen-3-yl)acetic acid, FARNESYL PYROPHOSPHATE SYNTHASE, PHOSPHATE ION | | Authors: | Rondeau, J.-M. | | Deposit date: | 2010-05-17 | | Release date: | 2010-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.56 Å) | | Cite: | Allosteric non-bisphosphonate FPPS inhibitors identified by fragment-based discovery.
Nat.Chem.Biol., 6, 2010
|
|
1HPH
 
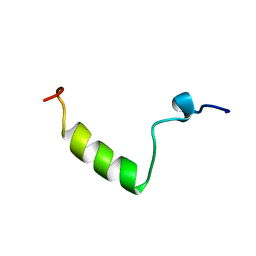 | |
7B17
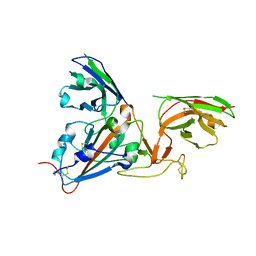 
 | | SARS-CoV-spike RBD bound to two neutralising nanobodies. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, SARS-CoV-2 neutralizing biparatopic nanobody VE,nanobody E from Lama glama,SARS-CoV-2 neutralizing biparatopic nanobody VE,nanobody E from Lama glama, Spike protein S1 | | Authors: | Hallberg, B.M, Das, H. | | Deposit date: | 2020-11-23 | | Release date: | 2021-02-10 | | Last modified: | 2022-12-21 | | Method: | ELECTRON MICROSCOPY (4.01 Å) | | Cite: | Structure-guided multivalent nanobodies block SARS-CoV-2 infection and suppress mutational escape
Science, 371, 2021
|
|
7B14
 
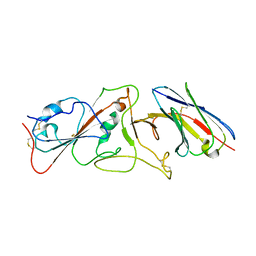 | | Nanobody E bound to Spike-RBD in a localized reconstruction | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Nanobody against SARS-CoV-2, Spike protein S1 | | Authors: | Hallberg, B.M, Das, H. | | Deposit date: | 2020-11-23 | | Release date: | 2021-04-28 | | Last modified: | 2022-12-21 | | Method: | ELECTRON MICROSCOPY (3.79 Å) | | Cite: | Structure-guided multivalent nanobodies block SARS-CoV-2 infection and suppress mutational escape
Science, 371, 2021
|
|
7B18
 
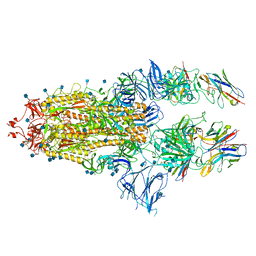 | | SARS-CoV-spike bound to two neutralising nanobodies | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Nanobody against SARS-CoV-2 VHH E, ... | | Authors: | Hallberg, B.M, Das, H. | | Deposit date: | 2020-11-24 | | Release date: | 2021-04-28 | | Method: | ELECTRON MICROSCOPY (2.62 Å) | | Cite: | Structure-guided multivalent nanobodies block SARS-CoV-2 infection and suppress mutational escape.
Science, 371, 2021
|
|
3SD5
 
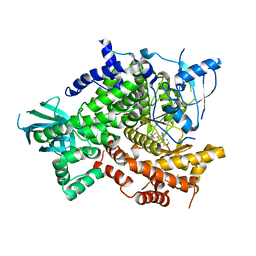 | | Crystal Structure of PI3K gamma with 5-(2,4-dimorpholinopyrimidin-6-yl)-4-(trifluoromethyl)pyridin-2-amine | | Descriptor: | 5-[2,6-di(morpholin-4-yl)pyrimidin-4-yl]-4-(trifluoromethyl)pyridin-2-amine, Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform | | Authors: | Knapp, M.S, Elling, R.A. | | Deposit date: | 2011-06-08 | | Release date: | 2012-01-04 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Identification and Characterization of NVP-BKM120, an Orally Available Pan-Class I PI3-Kinase Inhibitor.
Mol.Cancer Ther., 11, 2012
|
|
5D10
 
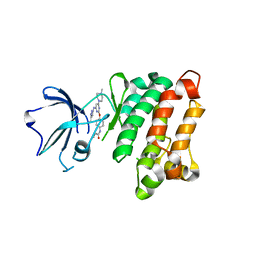 | | Kinase domain of cSrc in complex with RL236 | | Descriptor: | N-[4-({4-(4-methylpiperazin-1-yl)-6-[(5-methyl-1H-pyrazol-3-yl)amino]pyrimidin-2-yl}oxy)phenyl]prop-2-enamide, Proto-oncogene tyrosine-protein kinase Src | | Authors: | Becker, C, Mayer-Wrangowski, S.C, Julian, E, Rauh, D. | | Deposit date: | 2015-08-03 | | Release date: | 2015-09-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Targeting Drug Resistance in EGFR with Covalent Inhibitors: A Structure-Based Design Approach.
J.Med.Chem., 58, 2015
|
|
5D11
 
 | | Kinase domain of cSrc in complex with RL235 | | Descriptor: | GLYCEROL, N-[3-({4-(4-methylpiperazin-1-yl)-6-[(5-methyl-1H-pyrazol-3-yl)amino]pyrimidin-2-yl}oxy)phenyl]prop-2-enamide, Proto-oncogene tyrosine-protein kinase Src | | Authors: | Becker, C, Gruetter, C, Engel, J, Rauh, D. | | Deposit date: | 2015-08-03 | | Release date: | 2015-09-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Targeting Drug Resistance in EGFR with Covalent Inhibitors: A Structure-Based Design Approach.
J.Med.Chem., 58, 2015
|
|
5D12
 
 | | Kinase domain of cSrc in complex with RL40 | | Descriptor: | N-[2-phenyl-4-(1H-pyrazol-3-ylamino)quinazolin-7-yl]prop-2-enamide, Proto-oncogene tyrosine-protein kinase Src | | Authors: | Becker, C, Richters, A, Engel, J, Rauh, D. | | Deposit date: | 2015-08-03 | | Release date: | 2015-09-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Targeting Drug Resistance in EGFR with Covalent Inhibitors: A Structure-Based Design Approach.
J.Med.Chem., 58, 2015
|
|
2HAL
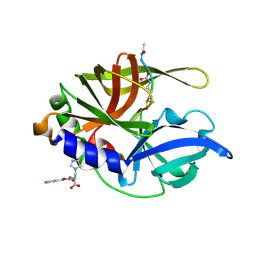 
 | | An episulfide cation (thiiranium ring) trapped in the active site of HAV 3C proteinase inactivated by peptide-based ketone inhibitors | | Descriptor: | Hepatitis A Protease 3C, N-ACETYL-LEUCYL-PHENYLALANYL-PHENYLALANYL-GLUTAMATE-FLUOROMETHYLKETONE INHIBITOR, N-[(BENZYLOXY)CARBONYL]-L-ALANINE | | Authors: | Yin, J, Cherney, M.M, Bergmann, E.M, James, M.N. | | Deposit date: | 2006-06-13 | | Release date: | 2006-08-08 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | An Episulfide Cation (Thiiranium Ring) Trapped in the Active Site of HAV 3C Proteinase Inactivated by Peptide-based Ketone Inhibitors.
J.Mol.Biol., 361, 2006
|
|
3M0E
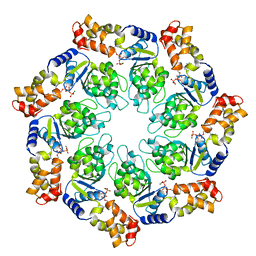 
 | | Crystal structure of the ATP-bound state of Walker B mutant of NtrC1 ATPase domain | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, Transcriptional regulator (NtrC family) | | Authors: | Chen, B, Sysoeva, T.A, Chowdhury, S, Rusu, M, Birmanns, S, Guo, L, Hanson, J, Yang, H, Nixon, B.T. | | Deposit date: | 2010-03-02 | | Release date: | 2010-11-03 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Engagement of Arginine Finger to ATP Triggers Large Conformational Changes in NtrC1 AAA+ ATPase for Remodeling Bacterial RNA Polymerase.
Structure, 18, 2010
|
|
2HV9
 
 | | Encephalitozoon cuniculi mRNA Cap (Guanine-N7) Methyltransferase in complex with sinefungin | | Descriptor: | SINEFUNGIN, mRNA cap guanine-N7 methyltransferase | | Authors: | Lima, C.D. | | Deposit date: | 2006-07-27 | | Release date: | 2006-09-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Mutational Analysis of Encephalitozoon cuniculi mRNA Cap (Guanine-N7) Methyltransferase, Structure of the Enzyme Bound to Sinefungin, and Evidence That Cap Methyltransferase Is the Target of Sinefungin's Antifungal Activity
J.Biol.Chem., 281, 2006
|
|
3KTT
 
 | | Atomic model of bovine TRiC CCT2(beta) subunit derived from a 4.0 Angstrom cryo-EM map | | Descriptor: | T-complex protein 1 subunit beta | | Authors: | Cong, Y, Baker, M.L, Ludtke, S.J, Frydman, J, Chiu, W. | | Deposit date: | 2009-11-25 | | Release date: | 2010-03-16 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | 4.0-A resolution cryo-EM structure of the mammalian chaperonin TRiC/CCT reveals its unique subunit arrangement.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
6QET
 
 | | [41-82]Gga-AvBD11 | | Descriptor: | Gallinacin-11 | | Authors: | Meudal, H, Loth, K, Delmas, A.F, Landon, C. | | Deposit date: | 2019-01-08 | | Release date: | 2019-12-18 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structure, function, and evolution ofGga-AvBD11, the archetype of the structural avian-double-beta-defensin family.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
6QEU
 
 | | Gga-AvBD11 (Avian beta-defensin 11 from Gallus gallus) | | Descriptor: | Gallinacin-11 | | Authors: | Meudal, H, Loth, K, Delmas, A.F, Landon, C. | | Deposit date: | 2019-01-08 | | Release date: | 2019-12-18 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structure, function, and evolution ofGga-AvBD11, the archetype of the structural avian-double-beta-defensin family.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
