4JMW
 
 | |
4JM6
 
 | | Crystal structure of Cytochrome C Peroxidase W191G-Gateless in complex with 2,4-diaminopyrimidine | | 分子名称: | Cytochrome c peroxidase, PHOSPHATE ION, PROTOPORPHYRIN IX CONTAINING FE, ... | | 著者 | Boyce, S.E, Fischer, M, Fish, I. | | 登録日 | 2013-03-13 | | 公開日 | 2013-05-01 | | 最終更新日 | 2024-02-28 | | 実験手法 | X-RAY DIFFRACTION (1.45 Å) | | 主引用文献 | Blind prediction of charged ligand binding affinities in a model binding site.
J.Mol.Biol., 425, 2013
|
|
7UM7
 
 | | CryoEM structure of Go-coupled 5-HT5AR in complex with Methylergometrine | | 分子名称: | (8beta)-N-[(2S)-1-hydroxybutan-2-yl]-6-methyl-9,10-didehydroergoline-8-carboxamide, 5-hydroxytryptamine receptor 5A, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | 著者 | Zhang, S, Fay, J.F, Roth, B.L. | | 登録日 | 2022-04-06 | | 公開日 | 2022-07-20 | | 最終更新日 | 2022-07-27 | | 実験手法 | ELECTRON MICROSCOPY (2.75 Å) | | 主引用文献 | Inactive and active state structures template selective tools for the human 5-HT 5A receptor.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7UM5
 
 | | CryoEM structure of Go-coupled 5-HT5AR in complex with 5-CT | | 分子名称: | 3-(2-azanylethyl)-1H-indole-5-carboxamide, 5-hydroxytryptamine receptor 5A, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | 著者 | Zhang, S, Fay, J.F, Roth, B.L. | | 登録日 | 2022-04-06 | | 公開日 | 2022-07-20 | | 最終更新日 | 2022-07-27 | | 実験手法 | ELECTRON MICROSCOPY (2.73 Å) | | 主引用文献 | Inactive and active state structures template selective tools for the human 5-HT 5A receptor.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7UM4
 
 | |
7UM6
 
 | | CryoEM structure of Go-coupled 5-HT5AR in complex with Lisuride | | 分子名称: | 5-hydroxytryptamine receptor 5A, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | 著者 | Zhang, S, Fay, J.F, Roth, B.L. | | 登録日 | 2022-04-06 | | 公開日 | 2022-07-20 | | 最終更新日 | 2022-07-27 | | 実験手法 | ELECTRON MICROSCOPY (2.79 Å) | | 主引用文献 | Inactive and active state structures template selective tools for the human 5-HT 5A receptor.
Nat.Struct.Mol.Biol., 29, 2022
|
|
4JMA
 
 | | Crystal structure of Cytochrome C Peroxidase W191G-Gateless in complex with 3-Fluorocatechol | | 分子名称: | 3-FLUOROBENZENE-1,2-DIOL, Cytochrome c peroxidase, PROTOPORPHYRIN IX CONTAINING FE | | 著者 | Boyce, S.E, Fischer, M, Fish, I. | | 登録日 | 2013-03-13 | | 公開日 | 2013-05-01 | | 最終更新日 | 2024-02-28 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Blind prediction of charged ligand binding affinities in a model binding site.
J.Mol.Biol., 425, 2013
|
|
4JM9
 
 | | Crystal structure of Cytochrome C Peroxidase W191G-Gateless in complex with 3-amino-1-methylpyridinium | | 分子名称: | 1-METHYL-1,6-DIHYDROPYRIDIN-3-AMINE, Cytochrome c peroxidase, IODIDE ION, ... | | 著者 | Boyce, S.E, Fischer, M, Fish, I. | | 登録日 | 2013-03-13 | | 公開日 | 2013-05-01 | | 最終更新日 | 2024-02-28 | | 実験手法 | X-RAY DIFFRACTION (1.41 Å) | | 主引用文献 | Blind prediction of charged ligand binding affinities in a model binding site.
J.Mol.Biol., 425, 2013
|
|
4JM8
 
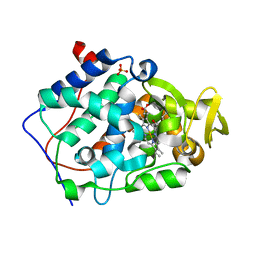 | | Crystal structure of Cytochrome C Peroxidase W191G-Gateless in complex with 2,6-diaminopyridine | | 分子名称: | Cytochrome c peroxidase, PHOSPHATE ION, PROTOPORPHYRIN IX CONTAINING FE, ... | | 著者 | Boyce, S.E, Fischer, M, Fish, I. | | 登録日 | 2013-03-13 | | 公開日 | 2013-05-01 | | 最終更新日 | 2024-02-28 | | 実験手法 | X-RAY DIFFRACTION (1.3 Å) | | 主引用文献 | Blind prediction of charged ligand binding affinities in a model binding site.
J.Mol.Biol., 425, 2013
|
|
7RAN
 
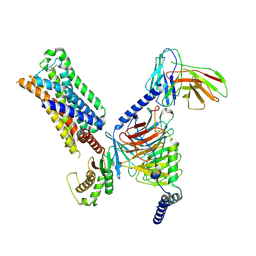 | | 5-HT2AR bound to a novel agonist in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM) | | 分子名称: | (3R)-3-methyl-5-(1H-pyrrolo[2,3-b]pyridin-3-yl)-1,2,3,6-tetrahydropyridin-1-ium, 5-hydroxytryptamine receptor 2A, G protein subunit q (Gi2-mini-Gq chimera), ... | | 著者 | Barros-Alvarez, X, Kim, K, Panova, O, Roth, B.L, Skiniotis, G. | | 登録日 | 2021-07-02 | | 公開日 | 2022-07-06 | | 最終更新日 | 2022-11-02 | | 実験手法 | ELECTRON MICROSCOPY (3.45 Å) | | 主引用文献 | Bespoke library docking for 5-HT 2A receptor agonists with antidepressant activity.
Nature, 610, 2022
|
|
7S8O
 
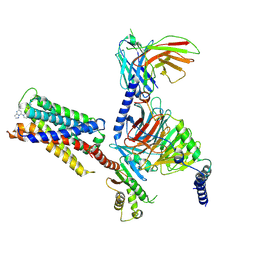 | | CryoEM structure of Gi-coupled MRGPRX2 with small molecule agonist (R)-Zinc-3573 | | 分子名称: | (3R)-N,N-dimethyl-1-[(8S)-5-phenylpyrazolo[1,5-a]pyrimidin-7-yl]pyrrolidin-3-amine, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | 著者 | Cao, C, Fay, J.F, Gumpper, R.H, Roth, B.L. | | 登録日 | 2021-09-18 | | 公開日 | 2021-11-17 | | 最終更新日 | 2021-12-15 | | 実験手法 | ELECTRON MICROSCOPY (2.58 Å) | | 主引用文献 | Structure, function and pharmacology of human itch GPCRs.
Nature, 600, 2021
|
|
7S8L
 
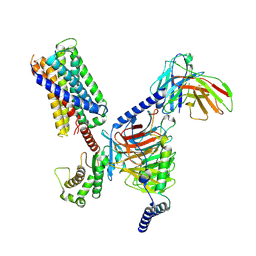 | | CryoEM structure of Gq-coupled MRGPRX2 with peptide agonist Cortistatin-14 | | 分子名称: | Cortistatin 14, Gs-mini-Gq chimera, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | 著者 | Cao, C, Fay, J.F, Gumpper, R.H, Roth, B.L. | | 登録日 | 2021-09-18 | | 公開日 | 2021-11-17 | | 最終更新日 | 2021-12-15 | | 実験手法 | ELECTRON MICROSCOPY (2.45 Å) | | 主引用文献 | Structure, function and pharmacology of human itch GPCRs.
Nature, 600, 2021
|
|
7S8P
 
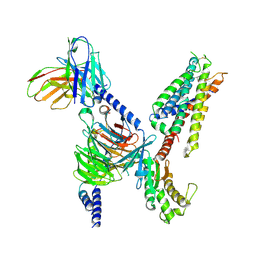 | | CryoEM structure of Gq-coupled MRGPRX4 with small molecule agonist MS47134 | | 分子名称: | Gs-mini-Gq chimera, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | 著者 | Cao, C, Fay, J.F, Gumpper, R.H, Roth, B.L. | | 登録日 | 2021-09-18 | | 公開日 | 2021-11-17 | | 最終更新日 | 2021-12-15 | | 実験手法 | ELECTRON MICROSCOPY (2.6 Å) | | 主引用文献 | Structure, function and pharmacology of human itch GPCRs.
Nature, 600, 2021
|
|
7S8M
 
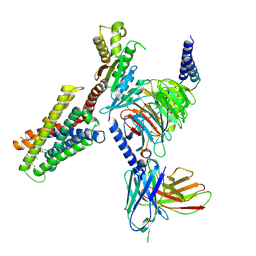 | | CryoEM structure of Gi-coupled MRGPRX2 with peptide agonist Cortistatin-14 | | 分子名称: | Cortistatin 14, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | 著者 | Cao, C, Fay, J.F, Gumpper, R.H, Roth, B.L. | | 登録日 | 2021-09-18 | | 公開日 | 2021-11-17 | | 最終更新日 | 2021-12-15 | | 実験手法 | ELECTRON MICROSCOPY (2.54 Å) | | 主引用文献 | Structure, function and pharmacology of human itch GPCRs.
Nature, 600, 2021
|
|
7S8N
 
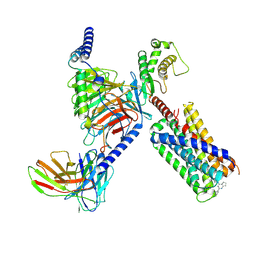 | | CryoEM structure of Gq-coupled MRGPRX2 with small molecule agonist (R)-Zinc-3573 | | 分子名称: | (3R)-N,N-dimethyl-1-[(8S)-5-phenylpyrazolo[1,5-a]pyrimidin-7-yl]pyrrolidin-3-amine, Gs-mini-Gq chimera, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | 著者 | Cao, C, Fay, J.F, Gumpper, R.H, Roth, B.L. | | 登録日 | 2021-09-18 | | 公開日 | 2021-11-17 | | 最終更新日 | 2021-12-15 | | 実験手法 | ELECTRON MICROSCOPY (2.9 Å) | | 主引用文献 | Structure, function and pharmacology of human itch GPCRs.
Nature, 600, 2021
|
|
8DWH
 
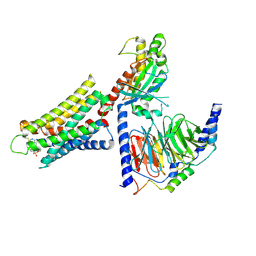 | | CryoEM structure of Gq-coupled MRGPRX1 with ligand Compound-16 | | 分子名称: | Gs-mini-Gq chimera, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | 著者 | Liu, Y, Cao, C, Fay, J.F, Roth, B.L. | | 登録日 | 2022-08-01 | | 公開日 | 2022-11-02 | | 最終更新日 | 2023-04-12 | | 実験手法 | ELECTRON MICROSCOPY (3.25 Å) | | 主引用文献 | Ligand recognition and allosteric modulation of the human MRGPRX1 receptor.
Nat.Chem.Biol., 19, 2023
|
|
8DWG
 
 | | CryoEM structure of Gq-coupled MRGPRX1 with peptide ligand BAM8-22 and positive allosteric modulator ML382 | | 分子名称: | 2-[(cyclopropanesulfonyl)amino]-N-(2-ethoxyphenyl)benzamide, Gs-mini-Gq chimera, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | 著者 | Liu, Y, Cao, C, Fay, J.F, Roth, B.L. | | 登録日 | 2022-08-01 | | 公開日 | 2022-11-02 | | 最終更新日 | 2023-04-12 | | 実験手法 | ELECTRON MICROSCOPY (2.71 Å) | | 主引用文献 | Ligand recognition and allosteric modulation of the human MRGPRX1 receptor.
Nat.Chem.Biol., 19, 2023
|
|
8DWC
 
 | | CryoEM structure of Gq-coupled MRGPRX1 with peptide agonist BAM8-22 | | 分子名称: | Gs-mini-Gq chimera, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | 著者 | Liu, Y, Cao, C, Fay, J.F, Roth, B.L. | | 登録日 | 2022-08-01 | | 公開日 | 2022-11-02 | | 最終更新日 | 2023-04-12 | | 実験手法 | ELECTRON MICROSCOPY (2.87 Å) | | 主引用文献 | Ligand recognition and allosteric modulation of the human MRGPRX1 receptor.
Nat.Chem.Biol., 19, 2023
|
|
4XGK
 
 | | Crystal structure of UDP-galactopyranose mutase from Corynebacterium diphtheriae in complex with 2-[4-(4-chlorophenyl)-7-(2-thienyl)-2-thia-5,6,8,9-tetrazabicyclo[4.3.0]nona-4,7,9-trien-3-yl]acetic | | 分子名称: | DIHYDROFLAVINE-ADENINE DINUCLEOTIDE, SULFATE ION, UDP-galactopyranose mutase, ... | | 著者 | Wangkanont, K, Heroux, A, Forest, K.T, Kiessling, L.L. | | 登録日 | 2014-12-31 | | 公開日 | 2015-08-12 | | 最終更新日 | 2023-09-27 | | 実験手法 | X-RAY DIFFRACTION (2.652 Å) | | 主引用文献 | Virtual Screening for UDP-Galactopyranose Mutase Ligands Identifies a New Class of Antimycobacterial Agents.
Acs Chem.Biol., 10, 2015
|
|
4XX9
 
 | |
7M95
 
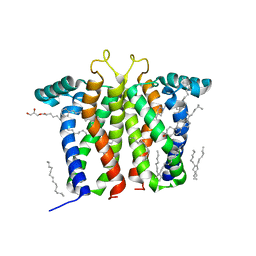 | | Bovine sigma-2 receptor bound to Z1241145220 | | 分子名称: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, 3-[1-(3-phenylpropyl)-1,2,3,6-tetrahydropyridin-4-yl]-1H-pyrrolo[2,3-b]pyridine, CHOLESTEROL, ... | | 著者 | Alon, A, Kruse, A.C. | | 登録日 | 2021-03-30 | | 公開日 | 2021-12-15 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.41 Å) | | 主引用文献 | Structures of the sigma 2 receptor enable docking for bioactive ligand discovery.
Nature, 600, 2021
|
|
7MFI
 
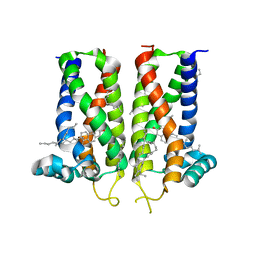 | | Bovine sigma-2 receptor bound to cholesterol | | 分子名称: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CHOLESTEROL, ... | | 著者 | Alon, A, Kruse, A.C. | | 登録日 | 2021-04-09 | | 公開日 | 2021-12-15 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.81 Å) | | 主引用文献 | Structures of the sigma 2 receptor enable docking for bioactive ligand discovery.
Nature, 600, 2021
|
|
7M94
 
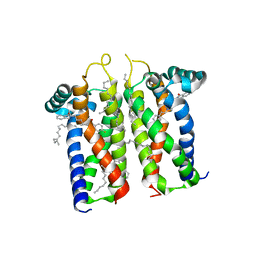 | | Bovine sigma-2 receptor bound to Roluperidone | | 分子名称: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, Roluperidone, Sigma intracellular receptor 2 | | 著者 | Alon, A, Kruse, A.C. | | 登録日 | 2021-03-30 | | 公開日 | 2021-12-15 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.71 Å) | | 主引用文献 | Structures of the sigma 2 receptor enable docking for bioactive ligand discovery.
Nature, 600, 2021
|
|
7M96
 
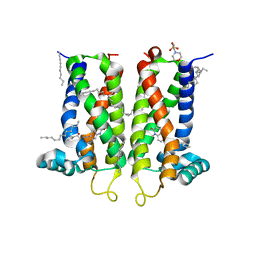 | | Bovine sigma-2 receptor bound to Z4857158944 | | 分子名称: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 6-({[(1S)-1-hydroxy-2-methyl-1-phenylpropan-2-yl]amino}methyl)-1-methyl-3,4-dihydroquinolin-2(1H)-one, ... | | 著者 | Alon, A, Kruse, A.C. | | 登録日 | 2021-03-30 | | 公開日 | 2021-12-15 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.41 Å) | | 主引用文献 | Structures of the sigma 2 receptor enable docking for bioactive ligand discovery.
Nature, 600, 2021
|
|
7M93
 
 | | Bovine sigma-2 receptor bound to PB28 | | 分子名称: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, PB28, Sigma intracellular receptor 2 | | 著者 | Alon, A, Kruse, A.C. | | 登録日 | 2021-03-30 | | 公開日 | 2021-12-15 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.94 Å) | | 主引用文献 | Structures of the sigma 2 receptor enable docking for bioactive ligand discovery.
Nature, 600, 2021
|
|
