6JQ8
 
 | |
1DST
 | |
5GLG
 
 | |
6KIA
 
 | |
6KI9
 
 | | Apo structure of FabMG, novel types of Enoyl-acyl carrier protein reductase | | Descriptor: | 1,2-ETHANEDIOL, FabMG, novel types of Enoyl-acyl carrier protein reductase, ... | | Authors: | Kim, S, Rhee, S. | | Deposit date: | 2019-07-17 | | Release date: | 2020-05-20 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | A triclosan-resistance protein from the soil metagenome is a novel enoyl-acyl carrier protein reductase: Structure-guided functional analysis.
Febs J., 287, 2020
|
|
5UUT
 
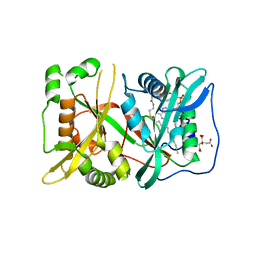 | | N-myristoyltransferase 1 (NMT) bound to myristoyl-CoA | | Descriptor: | CITRIC ACID, Glycylpeptide N-tetradecanoyltransferase 1, TETRADECANOYL-COA | | Authors: | Goodwin, O, Pegan, S. | | Deposit date: | 2017-02-17 | | Release date: | 2018-01-17 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.252 Å) | | Cite: | Blocking Myristoylation of Src Inhibits Its Kinase Activity and Suppresses Prostate Cancer Progression.
Cancer Res., 77, 2017
|
|
5ZYN
 
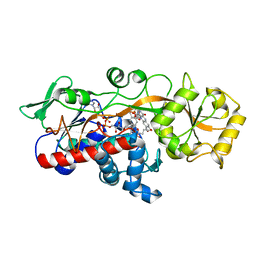 | | Fumarate reductase | | Descriptor: | FLAVIN MONONUCLEOTIDE, FLAVIN-ADENINE DINUCLEOTIDE, Fumarate reductase 2, ... | | Authors: | Park, H.H, Kim, C.M. | | Deposit date: | 2018-05-25 | | Release date: | 2018-10-03 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Molecular basis of maintaining an oxidizing environment under anaerobiosis by soluble fumarate reductase.
Nat Commun, 9, 2018
|
|
4DZO
 
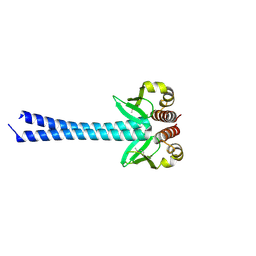 | |
2EIX
 
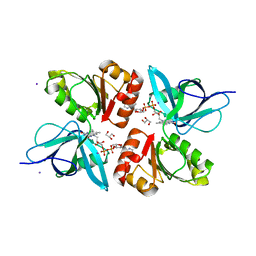 | | The Structure of Physarum polycephalum cytochrome b5 reductase | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, IODIDE ION, ... | | Authors: | Kim, S.W, Suga, M, Ogasahara, K, Ikegami, T, Minami, Y, Yubisui, T, Tsukihara, T. | | Deposit date: | 2007-03-14 | | Release date: | 2007-04-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Structure of Physarum polycephalum cytochrome b5 reductase at 1.56 A resolution.
Acta Crystallogr.,Sect.F, 63, 2007
|
|
1MEC
 
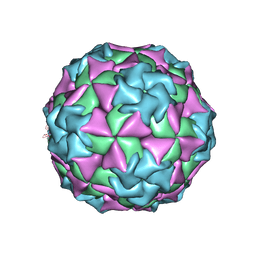 | |
1R1A
 
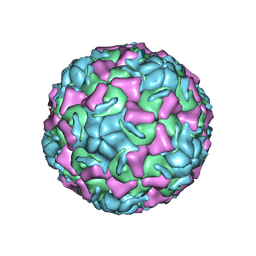 | | CRYSTAL STRUCTURE OF HUMAN RHINOVIRUS SEROTYPE 1A (HRV1A) | | Descriptor: | HUMAN RHINOVIRUS 1A COAT PROTEIN (SUBUNIT VP1), HUMAN RHINOVIRUS 1A COAT PROTEIN (SUBUNIT VP2), HUMAN RHINOVIRUS 1A COAT PROTEIN (SUBUNIT VP3), ... | | Authors: | Kim, S, Rossmann, M.G. | | Deposit date: | 1989-03-15 | | Release date: | 1990-07-15 | | Last modified: | 2023-03-15 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal structure of human rhinovirus serotype 1A (HRV1A).
J.Mol.Biol., 210, 1989
|
|
4UZ0
 | | Crystal Structure of apoptosis repressor with CARD (ARC) | | Descriptor: | GLYCEROL, NUCLEOLAR PROTEIN 3 | | Authors: | Kim, S.H, Jeong, J.H, Jang, T.H, Kim, Y.G, Park, H.H. | | Deposit date: | 2014-09-04 | | Release date: | 2015-07-01 | | Last modified: | 2017-07-12 | | Method: | X-RAY DIFFRACTION (2.399 Å) | | Cite: | Crystal Structure of Caspase Recruiting Domain (Card) of Apoptosis Repressor with Card (Arc) and its Implication in Inhibition of Apoptosis.
Sci.Rep., 5, 2015
|
|
8W77
 
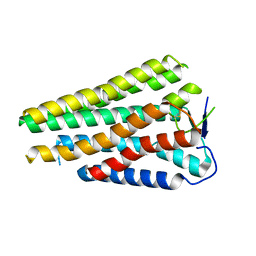 | | Human Consensus Olfactory Receptor OR52c in apo state, OR52c only | | Descriptor: | Human Consensus Olfactory Receptor OR52c in apo state, receptor only,Soluble cytochrome b562 | | Authors: | Choi, C.W, Bae, J, Choi, H.-J, Kim, J. | | Deposit date: | 2023-08-30 | | Release date: | 2023-12-20 | | Method: | ELECTRON MICROSCOPY (3.61 Å) | | Cite: | Understanding the molecular mechanisms of odorant binding and activation of the human OR52 family.
Nat Commun, 14, 2023
|
|
5ITS
 
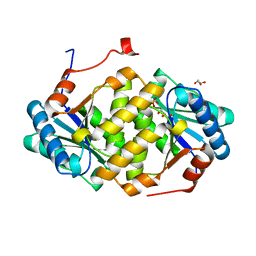 | | Crystal structure of LOG from Corynebacterium glutamicum | | Descriptor: | Cytokinin riboside 5'-monophosphate phosphoribohydrolase, GLYCEROL, PHOSPHATE ION | | Authors: | Seo, H.-G, Kim, K.-J. | | Deposit date: | 2016-03-17 | | Release date: | 2016-08-24 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis for cytokinin production by LOG from Corynebacterium glutamicum
Sci Rep, 6, 2016
|
|
5IZV
 
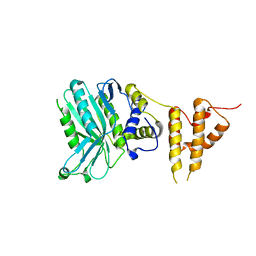 | | Crystal structure of the legionella pneumophila effector protein RavZ - F222 | | Descriptor: | Uncharacterized protein RavZ | | Authors: | Kwon, D.H, Kim, L, Kim, B.-W, Hong, S.B, Song, H.K. | | Deposit date: | 2016-03-26 | | Release date: | 2016-11-09 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.814 Å) | | Cite: | The 1:2 complex between RavZ and LC3 reveals a mechanism for deconjugation of LC3 on the phagophore membrane
Autophagy, 13, 2017
|
|
8HTI
 
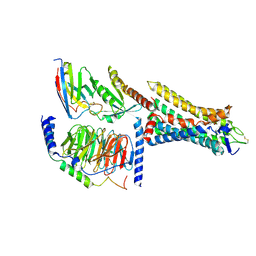 | | Human Consensus Olfactory Receptor OR52c in Complex with Octanoic Acid (OCA) and G Protein | | Descriptor: | Consensus Olfactory Receptor OR52c, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Choi, C.W, Bae, J, Choi, H.-J, Kim, J. | | Deposit date: | 2022-12-21 | | Release date: | 2023-12-20 | | Method: | ELECTRON MICROSCOPY (2.97 Å) | | Cite: | Understanding the molecular mechanisms of odorant binding and activation of the human OR52 family.
Nat Commun, 14, 2023
|
|
2QKY
 
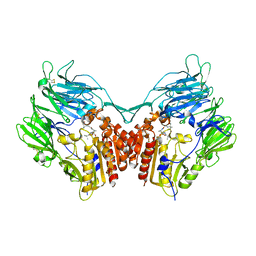 | | complex structure of dipeptidyl peptidase IV and a oxadiazolyl ketone | | Descriptor: | 2-[(2-{(2S,4S)-2-[(R)-(5-tert-butyl-1,3,4-oxadiazol-2-yl)(hydroxy)methyl]-4-fluoropyrrolidin-1-yl}-2-oxoethyl)amino]-2-methylpropan-1-ol, Dipeptidyl peptidase 4 (EC 3.4.14.5) (Dipeptidyl peptidase IV) (DPP IV) (T-cell activation antigen CD26) (TP103) (Adenosine deaminase complexing protein 2) (ADABP) (Dipeptidyl peptidase 4 soluble form) (Dipeptidyl peptidase IV soluble form) | | Authors: | Kim, K.-H, Hong, S.Y, Koo, K.D, Lee, C.-S, Kim, G.T, Han, H.O. | | Deposit date: | 2007-07-12 | | Release date: | 2008-07-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Synthesis, SAR, and X-ray structure of novel potent DPPIV inhibitors: oxadiazolyl ketones.
Bioorg.Med.Chem.Lett., 17, 2007
|
|
8HTG
 
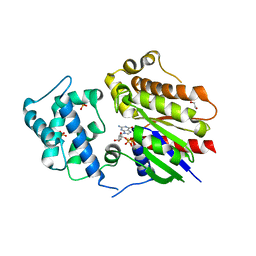 | | Crystal structure of Golf in complex with GTP-gamma S and Mg | | Descriptor: | 1,2-ETHANEDIOL, 5'-GUANOSINE-DIPHOSPHATE-MONOTHIOPHOSPHATE, Guanine nucleotide-binding protein G(olf) subunit alpha, ... | | Authors: | Kang, H, Choi, H.-J. | | Deposit date: | 2022-12-21 | | Release date: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | Understanding the molecular mechanisms of odorant binding and activation of the human OR52 family.
Nat Commun, 14, 2023
|
|
8UV0
 
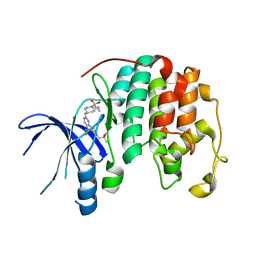 | |
5X42
 
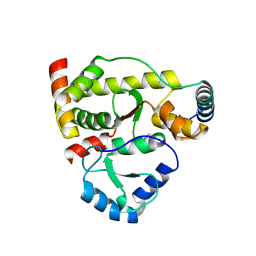 | | Structure of DotL(590-659)-DotN derived from Legionella pneumophila | | Descriptor: | DotN, IcmJ (DotN), IcmO (DotL), ... | | Authors: | Kwak, M.J, Kim, J.D, Oh, B.H. | | Deposit date: | 2017-02-09 | | Release date: | 2017-06-14 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Architecture of the type IV coupling protein complex of Legionella pneumophila
Nat Microbiol, 2, 2017
|
|
1VJS
 
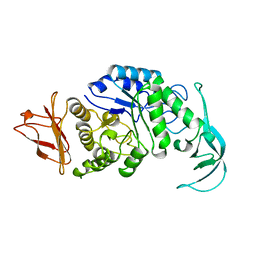 | |
7BYO
 
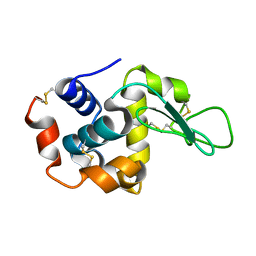 | | Lysozyme structure SS1 from SS mode | | Descriptor: | Lysozyme C | | Authors: | Kang, H.S, Lee, S.J. | | Deposit date: | 2020-04-24 | | Release date: | 2021-03-31 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | High-brightness self-seeded X-ray free-electron laser covering the 3.5 keV to 14.6 keV range
Nat Photonics, 2021
|
|
7BYP
 
 | | Lysozyme structure SASE1 from SASE mode | | Descriptor: | Lysozyme C | | Authors: | Kang, H.S, Lee, S.J. | | Deposit date: | 2020-04-24 | | Release date: | 2021-03-31 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | High-brightness self-seeded X-ray free-electron laser covering the 3.5 keV to 14.6 keV range
Nat Photonics, 2021
|
|
4EZ5
 
 | | CDK6 (monomeric) in complex with inhibitor | | Descriptor: | Cyclin-dependent kinase 6, {5-[4-(dimethylamino)piperidin-1-yl]-1H-imidazo[4,5-b]pyridin-2-yl}[2-(isoquinolin-4-yl)pyridin-4-yl]methanone | | Authors: | Chopra, R, Xu, M. | | Deposit date: | 2012-05-02 | | Release date: | 2013-02-06 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Fragment-Based Discovery of 7-Azabenzimidazoles as Potent, Highly Selective, and Orally Active CDK4/6 Inhibitors.
ACS Med Chem Lett, 3, 2012
|
|
8V4F
 | |
