7Y3K
 
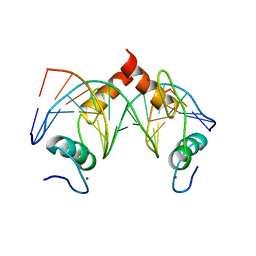 | | Structure of SALL4 ZFC4 bound with 16 bp AT-rich dsDNA | | Descriptor: | DNA (16-mer), Sal-like protein 4, ZINC ION | | Authors: | Ru, W, Xu, C. | | Deposit date: | 2022-06-11 | | Release date: | 2022-10-26 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.501 Å) | | Cite: | Structural studies of SALL family protein zinc finger cluster domains in complex with DNA reveal preferential binding to an AATA tetranucleotide motif.
J.Biol.Chem., 298, 2022
|
|
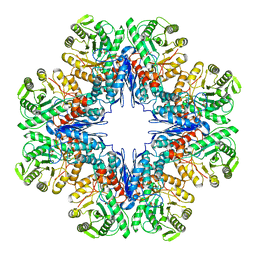
7E51
 | |
7E4F
 
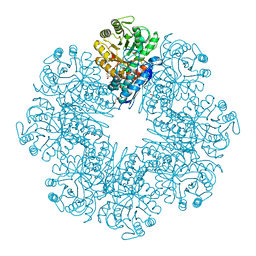 | | Mycobacterium tuberculosis enolase mutant - E204A complex with phosphoenolpyruvate | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Ahmad, M, Pal, R.K, Biswal, B.K. | | Deposit date: | 2021-02-11 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural snapshots of Mycobacterium tuberculosis enolase reveal dual mode of 2PG binding and its implication in enzyme catalysis.
Iucrj, 10, 2023
|
|
7E4X
 
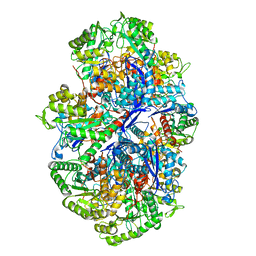 | |
5T7F
 
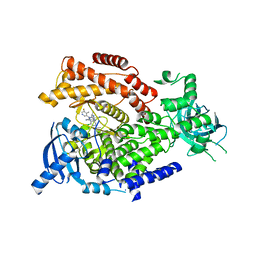 | | PI3Kdelta in complex with the inhibitor GS-643624 | | Descriptor: | 2,4-bis(azanyl)-6-[[(1~{S})-1-[5-chloranyl-3-(5-fluoranyl-4-methyl-pyridin-3-yl)-4-oxidanylidene-quinazolin-2-yl]ethyl]amino]pyrimidine-5-carbonitrile, Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform | | Authors: | Somoza, J.R, Villasenor, A. | | Deposit date: | 2016-09-04 | | Release date: | 2016-12-28 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Discovery of GS-9901: A Potent, Selective and Metabolically Stable Inhibitor of PI3Kd
To Be Published
|
|
5T8I
 
 | | PI3Kdelta in complex with the inhibitor GS-9901 | | Descriptor: | 2,4-diamino-6-{[(S)-[5-chloro-8-fluoro-4-oxo-3-(pyridin-3-yl)-3,4-dihydroquinazolin-2-yl](cyclopropyl)methyl]amino}pyrimidine-5-carbonitrile, Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform | | Authors: | Somoza, J.R, Villasenor, A. | | Deposit date: | 2016-09-07 | | Release date: | 2016-12-28 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Discovery of GS-9901: A Potent, Selective and Metabolically Stable Inhibitor of PI3Kd
To Be Published
|
|
2JB6
 
 | | Fab fragment in complex with small molecule hapten, crystal form-2 | | Descriptor: | 2-{(1E,3Z,5E,7E)-7-[3,3-DIMETHYL-5-SULFO-1-(2-SULFOETHYL)-1,3-DIHYDRO-2H-INDOL-2-YLIDENE]-4-METHYLHEPTA-1,3,5-TRIEN-1-YL}-3,3-DIMETHYL-5-SULFO-1-(2-SULFOETHYL)-3H-INDOLIUM, FAB FRAGMENT MOR03268 HEAVY CHAIN, FAB FRAGMENT MOR03268 LIGHT CHAIN | | Authors: | Hillig, R.C, Baesler, S, Malawski, G, Badock, V, Bahr, I, Schirner, M, Licha, K. | | Deposit date: | 2006-12-03 | | Release date: | 2008-01-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Fab Mor03268 Triggers Absorption Shift of a Diagnostic Dye Via Packaging in a Solvent-Shielded Fab Dimer Interface
J.Mol.Biol., 377, 2008
|
|
2JB5
 
 | | Fab fragment in complex with small molecule hapten, crystal form-1 | | Descriptor: | 2-{(1E,3Z,5E,7E)-7-[3,3-DIMETHYL-5-SULFO-1-(2-SULFOETHYL)-1,3-DIHYDRO-2H-INDOL-2-YLIDENE]-4-METHYLHEPTA-1,3,5-TRIEN-1-YL}-3,3-DIMETHYL-5-SULFO-1-(2-SULFOETHYL)-3H-INDOLIUM, FAB FRAGMENT MOR03268 HEAVY CHAIN, FAB FRAGMENT MOR03268 LIGHT CHAIN | | Authors: | Hillig, R.C, Baesler, S, Malawski, G, Badock, V, Bahr, I, Schirner, M, Licha, K. | | Deposit date: | 2006-12-03 | | Release date: | 2008-01-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Fab Mor03268 Triggers Absorption Shift of a Diagnostic Dye Via Packaging in a Solvent-Shielded Fab Dimer Interface
J.Mol.Biol., 377, 2008
|
|
1QND
 
 | | STEROL CARRIER PROTEIN-2, NMR, 20 STRUCTURES | | Descriptor: | NONSPECIFIC LIPID-TRANSFER PROTEIN | | Authors: | Lopez-Garcia, F, Szyperski, T, Dyer, J.H, Choinowski, T, Seedorf, U, Hauser, H, Wuthrich, K. | | Deposit date: | 1999-10-14 | | Release date: | 2000-07-03 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of the Sterol Carrier Protein-2: Implications for the Biological Role
J.Mol.Biol., 295, 2000
|
|
3BT1
 
 | | Structure of urokinase receptor, urokinase and vitronectin complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Urokinase plasminogen activator surface receptor, Urokinase-type plasminogen activator, ... | | Authors: | Huang, M. | | Deposit date: | 2007-12-27 | | Release date: | 2008-03-25 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structures of two human vitronectin, urokinase and urokinase receptor complexes
Nat.Struct.Mol.Biol., 15, 2008
|
|
5XNX
 
 | | Crystallographic structure of the enzymatically active N-terminal domain of the Rel protein from Mycobacterium tuberculosis | | Descriptor: | Bifunctional (p)ppGpp synthase/hydrolase RelA, MAGNESIUM ION | | Authors: | Singal, B, Balakrishna, A.M, Manimekalai, M.S.S, Nartey, W, Gruber, G. | | Deposit date: | 2017-05-24 | | Release date: | 2017-07-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | Crystallographic and solution structure of the N-terminal domain of the Rel protein from Mycobacterium tuberculosis
FEBS Lett., 591, 2017
|
|
3BT2
 
 | | Structure of urokinase receptor, urokinase and vitronectin complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Urokinase plasminogen activator surface receptor, ... | | Authors: | Huang, M. | | Deposit date: | 2007-12-27 | | Release date: | 2008-03-25 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structures of two human vitronectin, urokinase and urokinase receptor complexes
Nat.Struct.Mol.Biol., 15, 2008
|
|
7VXX
 
 | | Zika virus NS2B/NS3 protease bZipro(C143S) in complex with 4-amino benzamidine | | Descriptor: | P-AMINO BENZAMIDINE, Serine protease NS3, Serine protease subunit NS2B | | Authors: | Xiong, Y.C, Cheng, F, Zhang, J.Y, Su, H.X, Hu, H.C, Zou, Y, Li, M.J, Xu, Y.C. | | Deposit date: | 2021-11-13 | | Release date: | 2022-09-14 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure-based design of a novel inhibitor of the ZIKA virus NS2B/NS3 protease.
Bioorg.Chem., 128, 2022
|
|
7VXY
 
 | | Zika virus NS2B/NS3 protease bZipro(C143S) in complex with D-RKOR | | Descriptor: | Peptide inhibitor, Serine protease NS3, Serine protease subunit NS2B | | Authors: | Xiong, Y.C, Cheng, F, Zhang, J.Y, Su, H.X, Hu, H.C, Zou, Y, Li, M.J, Xu, Y.C. | | Deposit date: | 2021-11-13 | | Release date: | 2022-09-14 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.901 Å) | | Cite: | Structure-based design of a novel inhibitor of the ZIKA virus NS2B/NS3 protease.
Bioorg.Chem., 128, 2022
|
|
3I73
 
 | | Structural characterization for the nucleotide binding ability of subunit A with ADP of the A1AO ATP synthase | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, A-TYPE ATP SYNTHASE CATALYTIC SUBUNIT A, ... | | Authors: | Manimekalai, S.M.S, Kumar, A, Balakrishna, A.M, Gruber, G. | | Deposit date: | 2009-07-08 | | Release date: | 2010-01-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Nucleotide binding states of subunit A of the A-ATP synthase and the implication of P-loop switch in evolution.
J.Mol.Biol., 396, 2010
|
|
4AIU
 
 | | A complex structure of BtGH84 | | Descriptor: | (3AR,5R,6S,7R,7AR)-2,5-BIS(HYDROXYMETHYL)-5,6,7,7A-TETRAHYDRO-3AH-PYRANO[3,2-D][1,3]OXAZOLE-6,7-DIOL, CALCIUM ION, O-GLCNACASE BT_4395 | | Authors: | He, Y, Davies, G.J. | | Deposit date: | 2012-02-13 | | Release date: | 2012-06-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Metabolism of Vertebrate Amino Sugars with N-Glycolyl Groups: Intracellular Beta-O-Linked N-Glycolylglucosamine (Glcngc), Udp-Glcngc, and the Biochemical and Structural Rationale for the Substrate Tolerance of Beta-O-Linked Beta-N-Acetylglucosaminidase.
J.Biol.Chem., 287, 2012
|
|
3I72
 
 | | Structural characterization for the nucleotide binding ability of subunit A with SO4 of the A1AO ATP synthase | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, A-TYPE ATP SYNTHASE CATALYTIC SUBUNIT A, ... | | Authors: | Manimekalai, S.M.S, Kumar, A, Balakrishna, A.M, Gruber, G. | | Deposit date: | 2009-07-07 | | Release date: | 2010-01-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.47 Å) | | Cite: | Nucleotide binding states of subunit A of the A-ATP synthase and the implication of P-loop switch in evolution.
J.Mol.Biol., 396, 2010
|
|
7X4Y
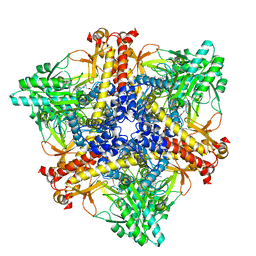 
 | | Crystal structure of Bacteroides thetaiotaomicron glutamate decarboxylase BTGAD-PLP-GABA complex | | Descriptor: | GAMMA-AMINO-BUTANOIC ACID, Glutamate decarboxylase, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Liu, S, Du, G, Wang, Y, Wen, B, Xin, F. | | Deposit date: | 2022-03-03 | | Release date: | 2023-04-05 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Coordinated regulation of Bacteroides thetaiotaomicron glutamate decarboxylase activity by multiple elements under different pH.
Food Chem, 403, 2023
|
|
7X52
 
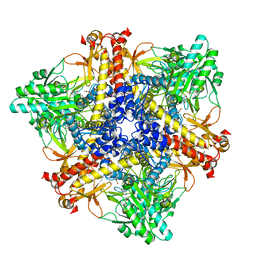 | | Crystal structure of Bacteroides thetaiotaomicron glutamate decarboxylase BTGAD-PLP complex | | Descriptor: | ACETATE ION, Glutamate decarboxylase, MALONATE ION, ... | | Authors: | Liu, S, Du, G, Wang, Y, Wen, B, Xin, F. | | Deposit date: | 2022-03-03 | | Release date: | 2023-04-05 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Coordinated regulation of Bacteroides thetaiotaomicron glutamate decarboxylase activity by multiple elements under different pH.
Food Chem, 403, 2023
|
|
7X4L
 
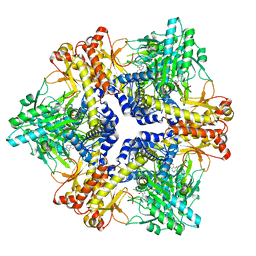 | | Crystal structure of Bacteroides thetaiotaomicron glutamate decarboxylase mutant Y303F-PLP complex | | Descriptor: | Glutamate decarboxylase, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Liu, S, Guoming, D, Yulu, W, Boting, W, Xin, F. | | Deposit date: | 2022-03-02 | | Release date: | 2023-05-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Coordinated regulation of Bacteroides thetaiotaomicron glutamate decarboxylase activity by multiple elements under different pH.
Food Chem, 403, 2023
|
|
7WIA
 
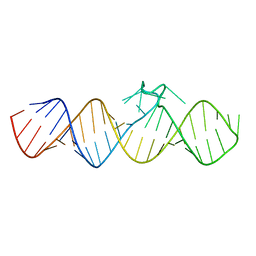 | | The apo-form of THF-II C22G riboswitch | | Descriptor: | RNA (50-MER) | | Authors: | Xu, L, Fang, X, Xiao, Y. | | Deposit date: | 2022-01-03 | | Release date: | 2023-01-18 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (3.22 Å) | | Cite: | Structural insights into translation regulation by the THF-II riboswitch.
Nucleic Acids Res., 51, 2023
|
|
7WII
 
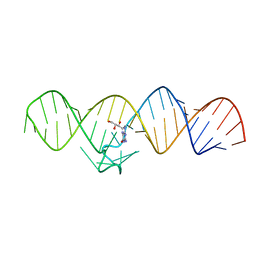 | | The THF-II riboswitch bound to NPR | | Descriptor: | 2-AMINO-7,8-DIHYDRO-6-(1,2,3-TRIHYDROXYPROPYL)-4(1H)-PTERIDINONE, RNA (50-MER) | | Authors: | Xu, L, Fang, X, Xiao, Y. | | Deposit date: | 2022-01-03 | | Release date: | 2023-01-18 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural insights into translation regulation by the THF-II riboswitch.
Nucleic Acids Res., 51, 2023
|
|
7WI9
 
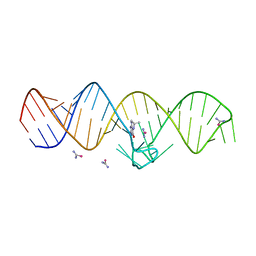 | | The THF-II riboswitch bound to THF and soaking with SeUrea | | Descriptor: | (6S)-5,6,7,8-TETRAHYDROFOLATE, RNA (50-MER), selenourea | | Authors: | Xu, L, Fang, X, Xiao, Y. | | Deposit date: | 2022-01-03 | | Release date: | 2023-01-18 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.98 Å) | | Cite: | Structural insights into translation regulation by the THF-II riboswitch.
Nucleic Acids Res., 51, 2023
|
|
7WIB
 
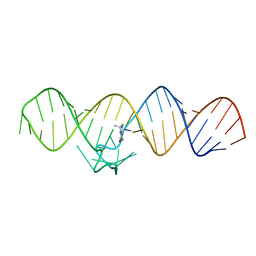 | | The THF-II riboswitch bound to THF | | Descriptor: | (6S)-5,6,7,8-TETRAHYDROFOLATE, RNA (50-MER) | | Authors: | Xu, L, Fang, X, Xiao, Y. | | Deposit date: | 2022-01-03 | | Release date: | 2023-01-18 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.83 Å) | | Cite: | Structural insights into translation regulation by the THF-II riboswitch.
Nucleic Acids Res., 51, 2023
|
|
3IDP
 
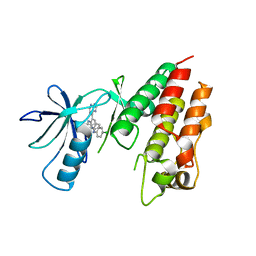 | | B-Raf V600E kinase domain in complex with an aminoisoquinoline inhibitor | | Descriptor: | B-Raf proto-oncogene serine/threonine-protein kinase, N~1~-(4-chlorophenyl)-6-methyl-N~5~-[3-(7H-purin-6-yl)pyridin-2-yl]isoquinoline-1,5-diamine | | Authors: | Whittington, D.A, Epstein, L.F. | | Deposit date: | 2009-07-21 | | Release date: | 2009-10-06 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Selective inhibitors of the mutant B-raf pathway: discovery of a potent and orally bioavailable aminoisoquinoline.
J.Med.Chem., 52, 2009
|
|
