5YET
 
 | | Structure of R354_WT | | Descriptor: | Uncharacterized protein R354 | | Authors: | Dou, C, Yu, M.J, Gu, Y.J, Cheng, W. | | Deposit date: | 2017-09-19 | | Release date: | 2018-06-20 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.806 Å) | | Cite: | Structural and Mechanistic Analyses Reveal a Unique Cas4-like Protein in the Mimivirus Virophage Resistance Element System.
Iscience, 3, 2018
|
|
5Y4Z
 
 | | Crystal structure of the Zika virus NS3 helicase complex with AMPPNP | | Descriptor: | MANGANESE (II) ION, NS3 helicase, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Li, L. | | Deposit date: | 2017-08-06 | | Release date: | 2017-08-30 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Structural view of the helicase reveals that Zika virus uses a conserved mechanism for unwinding RNA
Acta Crystallogr.,Sect.F, 74, 2018
|
|
5YSM
 
 | | Crystal Structure Analysis of Rif16 | | Descriptor: | Cytochrome P450, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Li, F.W, Qi, F.F, Xiao, Y.L, Zhao, G.P, Li, S.Y. | | Deposit date: | 2017-11-14 | | Release date: | 2018-07-04 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Deciphering the late steps of rifamycin biosynthesis.
Nat Commun, 9, 2018
|
|
7UFI
 
 | | VchTnsC AAA+ ATPase with DNA, single heptamer | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, DNA (5'-D(P*CP*TP*CP*CP*AP*GP*TP*AP*CP*AP*GP*CP*GP*CP*GP*GP*CP*TP*GP*AP*A)-3'), DNA (5'-D(P*TP*TP*CP*AP*GP*CP*CP*GP*CP*GP*CP*TP*GP*TP*AP*CP*TP*GP*GP*AP*G)-3'), ... | | Authors: | Fernandez, I.S, Sternberg, S.H. | | Deposit date: | 2022-03-22 | | Release date: | 2022-06-08 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Selective TnsC recruitment enhances the fidelity of RNA-guided transposition.
Nature, 609, 2022
|
|
7UFM
 
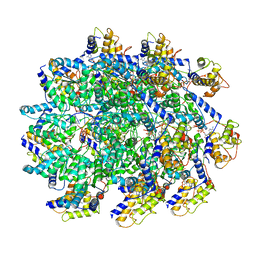 | | VchTnsC AAA+ with DNA (double heptamer) | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, DNA (36-MER), VchTnsC | | Authors: | Fernandez, I.S, Sternberg, S.H. | | Deposit date: | 2022-03-22 | | Release date: | 2022-06-08 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Selective TnsC recruitment enhances the fidelity of RNA-guided transposition.
Nature, 609, 2022
|
|
5YT7
 
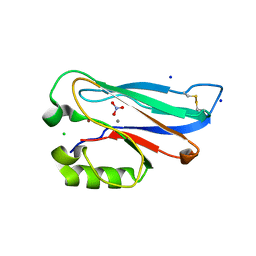 | |
5WV8
 
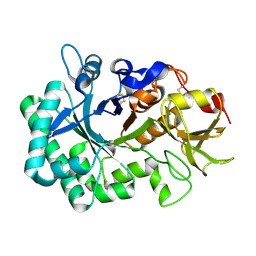 | |
7UEP
 
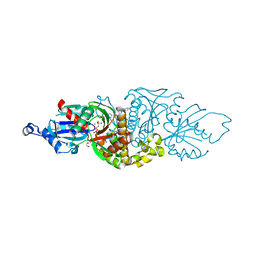 | | PANK3 complex structure with compound PZ-3860 | | Descriptor: | 1,2-ETHANEDIOL, 1-[(2R)-4-(6-chloropyridazin-3-yl)-2-methylpiperazin-1-yl]-2-(4-cyclopropylphenyl)ethan-1-one, MAGNESIUM ION, ... | | Authors: | White, S.W, Yun, M, Lee, R.E. | | Deposit date: | 2022-03-22 | | Release date: | 2023-03-29 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Development of Brain Penetrant Pyridazine Pantothenate Kinase Activators.
J.Med.Chem., 67, 2024
|
|
7UEO
 
 | | PANK3 complex structure with compound PZ-3977 | | Descriptor: | 1,2-ETHANEDIOL, 6-{4-[(4-cyclopropyl-2-fluorophenyl)acetyl]piperazin-1-yl}pyridazine-3-carbonitrile, MAGNESIUM ION, ... | | Authors: | White, S.W, Yun, M, Lee, R.E. | | Deposit date: | 2022-03-22 | | Release date: | 2023-03-29 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Development of Brain Penetrant Pyridazine Pantothenate Kinase Activators.
J.Med.Chem., 67, 2024
|
|
7UE4
 
 | | PANK3 complex structure with compound PZ-3855 | | Descriptor: | 1,2-ETHANEDIOL, 1-[4-(6-chloropyridazin-3-yl)piperazin-1-yl]-2-(4-cyclopropylphenyl)ethan-1-one, ACETATE ION, ... | | Authors: | White, S.W, Yun, M, Lee, R.E. | | Deposit date: | 2022-03-21 | | Release date: | 2023-03-29 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Development of Brain Penetrant Pyridazine Pantothenate Kinase Activators.
J.Med.Chem., 67, 2024
|
|
7UE3
 
 | | PANK3 complex structure with compound PZ-3804 | | Descriptor: | 1,2-ETHANEDIOL, 6-[4-({4-[(2R)-1-hydroxypropan-2-yl]phenyl}acetyl)piperazin-1-yl]pyridazine-3-carbonitrile, MAGNESIUM ION, ... | | Authors: | White, S.W, Yun, M, Lee, R.E. | | Deposit date: | 2022-03-21 | | Release date: | 2023-03-29 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Development of Brain Penetrant Pyridazine Pantothenate Kinase Activators.
J.Med.Chem., 67, 2024
|
|
7UEQ
 
 | | PANK3 complex structure with compound PZ-4061 | | Descriptor: | 1-[4-(6-chloropyridazin-3-yl)piperazin-1-yl]-2-[4-(dimethylamino)phenyl]ethan-1-one, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | White, S.W, Yun, M, Lee, R.E. | | Deposit date: | 2022-03-22 | | Release date: | 2023-03-29 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Development of Brain Penetrant Pyridazine Pantothenate Kinase Activators.
J.Med.Chem., 67, 2024
|
|
7UE5
 
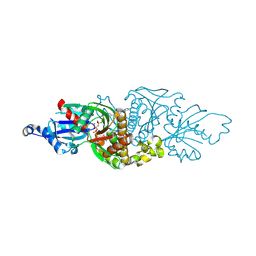 | | PANK3 complex structure with compound PZ-3741 | | Descriptor: | 6-[4-({4-[(2R)-1,1,1-trifluoropropan-2-yl]phenyl}acetyl)piperazin-1-yl]pyridazine-3-carbonitrile, 6-[4-({4-[(2S)-1,1,1-trifluoropropan-2-yl]phenyl}acetyl)piperazin-1-yl]pyridazine-3-carbonitrile, MAGNESIUM ION, ... | | Authors: | White, S.W, Yun, M, Lee, R.E. | | Deposit date: | 2022-03-21 | | Release date: | 2023-03-29 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Development of Brain Penetrant Pyridazine Pantothenate Kinase Activators.
J.Med.Chem., 67, 2024
|
|
7UE7
 
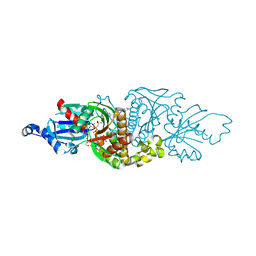 | | PANK3 complex structure with compound PZ-3883 | | Descriptor: | 1,2-ETHANEDIOL, 6-{4-[(4-cyclopropyl-3-fluorophenyl)acetyl]piperazin-1-yl}pyridazine-3-carbonitrile, ACETATE ION, ... | | Authors: | White, S.W, Yun, M, Lee, R.E. | | Deposit date: | 2022-03-21 | | Release date: | 2023-03-29 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Development of Brain Penetrant Pyridazine Pantothenate Kinase Activators.
J.Med.Chem., 67, 2024
|
|
7UE6
 
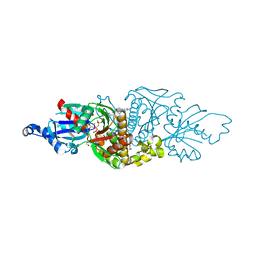 | | PANK3 complex structure with compound PZ-3802 | | Descriptor: | 1,2-ETHANEDIOL, 6-(4-{[5-fluoro-6-(propan-2-yl)pyridin-3-yl]acetyl}piperazin-1-yl)pyridazine-3-carbonitrile, ACETATE ION, ... | | Authors: | White, S.W, Yun, M, Lee, R.E. | | Deposit date: | 2022-03-21 | | Release date: | 2023-04-05 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Development of Brain Penetrant Pyridazine Pantothenate Kinase Activators.
J.Med.Chem., 67, 2024
|
|
5YZX
 
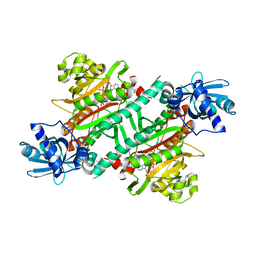 | | Crystal structure of E.coli LysU T146D mutant | | Descriptor: | BIS(ADENOSINE)-5'-TETRAPHOSPHATE, CALCIUM ION, Lysine--tRNA ligase, ... | | Authors: | Fang, P, Guo, M. | | Deposit date: | 2017-12-16 | | Release date: | 2018-12-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | A proposed role for MSC to reserve the canonical function in high eukaryotes prior to stimuli
To Be Published
|
|
5WVH
 
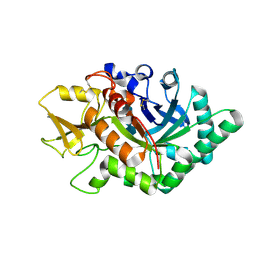 | |
5WVF
 
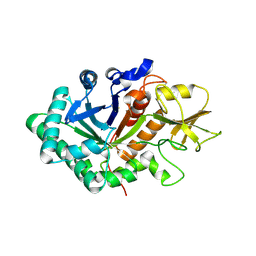 | |
5YXI
 
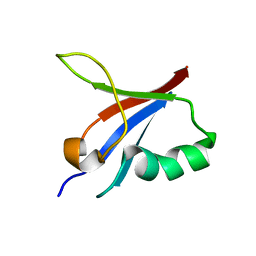 | | Designed protein dRafX6 | | Descriptor: | Designed protein dRafX6 | | Authors: | Liu, R. | | Deposit date: | 2017-12-05 | | Release date: | 2018-12-05 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | De novo sequence redesign of a functional Ras-binding domain globally inverted the surface charge distribution and led to extreme thermostability.
Biotechnol.Bioeng., 118, 2021
|
|
5ZTY
 
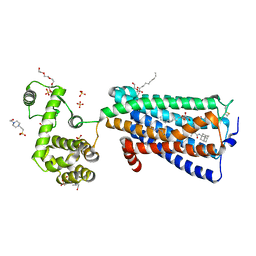 | | Crystal structure of human G protein coupled receptor | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Li, X.T, Hua, T, Wu, L.J, Liu, Z.J. | | Deposit date: | 2018-05-05 | | Release date: | 2019-01-30 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structure of the Human Cannabinoid Receptor CB2
Cell, 176, 2019
|
|
5WVB
 
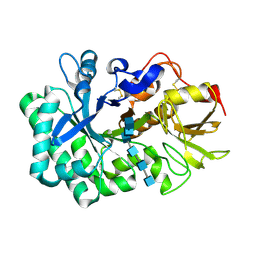 | |
6A0Q
 
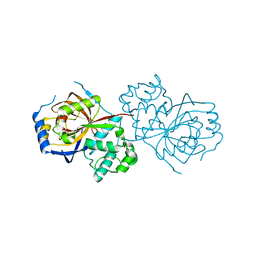 | | The crystal structure of Lpg2622_E64 complex | | Descriptor: | Lpg2622, N-[N-[1-HYDROXYCARBOXYETHYL-CARBONYL]LEUCYLAMINO-BUTYL]-GUANIDINE | | Authors: | Gong, X, Ge, H. | | Deposit date: | 2018-06-06 | | Release date: | 2018-09-12 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural characterization of the hypothetical protein Lpg2622, a new member of the C1 family peptidases from Legionella pneumophila
FEBS Lett., 592, 2018
|
|
2MVM
 
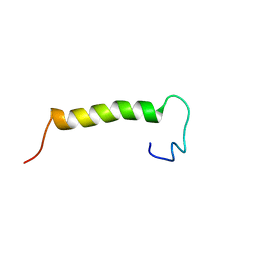 | | Solution structure of eEF1Bdelta CAR domain | | Descriptor: | Elongation factor 1-delta | | Authors: | Wu, H, Feng, Y. | | Deposit date: | 2014-10-09 | | Release date: | 2015-02-04 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Evolutionarily Conserved Binding of Translationally Controlled Tumor Protein to Eukaryotic Elongation Factor 1B.
J.Biol.Chem., 290, 2015
|
|
2MVN
 
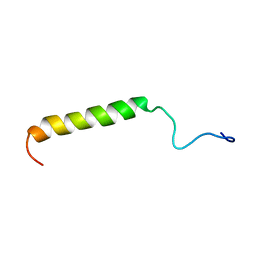 | |
5JI3
 
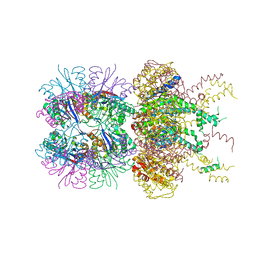 | | HslUV complex | | Descriptor: | 2'-DEOXYADENOSINE-5'-DIPHOSPHATE, ATP-dependent protease ATPase subunit HslU, ATP-dependent protease subunit HslV | | Authors: | Grant, R.A, Sauer, R.T, Schmitz, K.R, Baytshtok, V. | | Deposit date: | 2016-04-21 | | Release date: | 2016-12-07 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | A Structurally Dynamic Region of the HslU Intermediate Domain Controls Protein Degradation and ATP Hydrolysis.
Structure, 24, 2016
|
|
