[English] 日本語
 Yorodumi
Yorodumi- PDB-2bq4: Crystal structure of type I cytochrome c3 from Desulfovibrio africanus -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2bq4 | ||||||
|---|---|---|---|---|---|---|---|
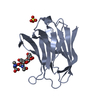
| Title | Crystal structure of type I cytochrome c3 from Desulfovibrio africanus | ||||||
 Components Components | BASIC CYTOCHROME C3 | ||||||
 Keywords Keywords | ELECTRON TRANSPORT / BASIC CYTOCHROME C3 / ELECTRON TRANSFER / SULFATE REDUCING BACTERIA / SAD / HEME / IRON | ||||||
| Function / homology |  Function and homology information Function and homology informationanaerobic respiration / periplasmic space / electron transfer activity / heme binding / metal ion binding Similarity search - Function | ||||||
| Biological species |  DESULFOVIBRIO AFRICANUS (bacteria) DESULFOVIBRIO AFRICANUS (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 1.68 Å SAD / Resolution: 1.68 Å | ||||||
 Authors Authors | Czjzek, M. / Pieulle, L. / Morelli, X. / Guerlesquin, F. / Hatchikian, E.C. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2005 Journal: J.Mol.Biol. / Year: 2005Title: The Type I / Type II Cytochrome C(3) Complex: An Electron Transfer Link in the Hydrogen-Sulfate Reduction Pathway. Authors: Pieulle, L. / Morelli, X. / Gallice, P. / Lojou, E. / Barbier, P. / Czjzek, M. / Bianco, P. / Guerlesquin, F. / Hatchikian, E.C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2bq4.cif.gz 2bq4.cif.gz | 74.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2bq4.ent.gz pdb2bq4.ent.gz | 57.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2bq4.json.gz 2bq4.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bq/2bq4 https://data.pdbj.org/pub/pdb/validation_reports/bq/2bq4 ftp://data.pdbj.org/pub/pdb/validation_reports/bq/2bq4 ftp://data.pdbj.org/pub/pdb/validation_reports/bq/2bq4 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| |||||||||||||||||||||||||||
| 2 | 
| |||||||||||||||||||||||||||
| Unit cell |
| |||||||||||||||||||||||||||
| Noncrystallographic symmetry (NCS) | NCS domain:
NCS domain segments:
NCS oper: (Code: given Matrix: (0.999442, 0.031835, -0.010076), Vector: |
- Components
Components
| #1: Protein | Mass: 12564.343 Da / Num. of mol.: 2 / Source method: isolated from a natural source Details: FOUR C-TYPE HEME-GROUPS COVALENTLY LINKED BY CYSTEINE RESIDUES Source: (natural)  DESULFOVIBRIO AFRICANUS (bacteria) / References: UniProt: P94691 DESULFOVIBRIO AFRICANUS (bacteria) / References: UniProt: P94691#2: Chemical | ChemComp-HEC / #3: Chemical | #4: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.3 Å3/Da / Density % sol: 47 % |
|---|---|
| Crystal grow | Details: 3.2M NAH2PO4/K2HPO4 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID14-2 / Wavelength: 0.933 / Beamline: ID14-2 / Wavelength: 0.933 |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Jan 31, 2003 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.933 Å / Relative weight: 1 |
| Reflection | Resolution: 1.68→48.22 Å / Num. obs: 106337 / % possible obs: 94.8 % / Observed criterion σ(I): 0 / Redundancy: 3.9 % / Rmerge(I) obs: 0.06 / Net I/σ(I): 20.2 |
| Reflection shell | Resolution: 1.68→1.75 Å / Redundancy: 2.1 % / Rmerge(I) obs: 0.06 / Mean I/σ(I) obs: 11.3 / % possible all: 71.8 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  SAD / Resolution: 1.68→48.22 Å / Cor.coef. Fo:Fc: 0.931 / Cor.coef. Fo:Fc free: 0.9 / SU B: 1.854 / SU ML: 0.063 / Cross valid method: THROUGHOUT / ESU R: 0.145 / ESU R Free: 0.134 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS. SAD / Resolution: 1.68→48.22 Å / Cor.coef. Fo:Fc: 0.931 / Cor.coef. Fo:Fc free: 0.9 / SU B: 1.854 / SU ML: 0.063 / Cross valid method: THROUGHOUT / ESU R: 0.145 / ESU R Free: 0.134 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 9.34 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.68→48.22 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller















































 PDBj
PDBj










