-Search query
-Search result
Showing 1 - 50 of 2,102 items for (author: wu & q)

EMDB-32979: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)

PDB-7x35: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)

EMDB-38617: 
SARS-CoV-2 RBD + IMCAS-123 + IMCAS-72 Fab

EMDB-38618: 
SARS-CoV-2 RBD + IMCAS-364 + hACE2

EMDB-38619: 
SARS-CoV-2 RBD + IMCAS-364 (Local Refinement)

EMDB-38620: 
SARS-CoV-2 Omicron BA.4 RBD + IMCAS-316 + ACE2

EMDB-38621: 
SARS-CoV-2 spike + IMCAS-123

EMDB-38823: 
Cryo-EM structure of the 123-316 scDb/PT-RBD complex

PDB-8xse: 
SARS-CoV-2 RBD + IMCAS-123 + IMCAS-72 Fab

PDB-8xsf: 
SARS-CoV-2 RBD + IMCAS-364 + hACE2

PDB-8xsi: 
SARS-CoV-2 RBD + IMCAS-364 (Local Refinement)

PDB-8xsj: 
SARS-CoV-2 Omicron BA.4 RBD + IMCAS-316 + ACE2

PDB-8xsl: 
SARS-CoV-2 spike + IMCAS-123

PDB-8y0y: 
Cryo-EM structure of the 123-316 scDb/PT-RBD complex

EMDB-36756: 
CryoEM structure of the transketolase ANIP from Streptomyces hygrospinosus

EMDB-38580: 
Structure of human class T GPCR TAS2R14-miniGs/gust complex with Aristolochic acid A.

EMDB-38582: 
Structure of human class T GPCR TAS2R14-DNGi complex with Aristolochic acid A.

EMDB-38583: 
Structure of human class T GPCR TAS2R14-Gi complex with Aristolochic acid A.

EMDB-38584: 
Structure of human class T GPCR TAS2R14-Gustducin complex with Aristolochic acid A.

EMDB-38586: 
Structure 2 of human class T GPCR TAS2R14-miniGs/gust complex with Flufenamic acid.

EMDB-38587: 
Structure of human class T GPCR TAS2R14-DNGi complex with Flufenamic acid.

EMDB-38588: 
Structure of human class T GPCR TAS2R14-Gi complex.

EMDB-39376: 
Structure of human class T GPCR TAS2R14-Ggustducin complex with agonist 28.1

PDB-8xql: 
Structure of human class T GPCR TAS2R14-miniGs/gust complex with Aristolochic acid A.

PDB-8xqn: 
Structure of human class T GPCR TAS2R14-DNGi complex with Aristolochic acid A.

PDB-8xqo: 
Structure of human class T GPCR TAS2R14-Gi complex with Aristolochic acid A.

PDB-8xqp: 
Structure of human class T GPCR TAS2R14-Gustducin complex with Aristolochic acid A.

PDB-8xqr: 
Structure 2 of human class T GPCR TAS2R14-miniGs/gust complex with Flufenamic acid.

PDB-8xqs: 
Structure of human class T GPCR TAS2R14-DNGi complex with Flufenamic acid.

PDB-8xqt: 
Structure of human class T GPCR TAS2R14-Gi complex.

PDB-8yky: 
Structure of human class T GPCR TAS2R14-Ggustducin complex with agonist 28.1

EMDB-50580: 
SOLIST cryo-tomogram of native left ventricle mouse heart muscle #1

EMDB-50582: 
SOLIST native mouse heart muscle tomogram #2
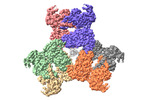
EMDB-36749: 
CryoEM structure of the NADP-dependent malic enzyme MaeB

EMDB-36757: 
CryoEM structure of a 2,3-hydroxycinnamic acid 1,2-dioxygenase MhpB in apo form

PDB-8jzo: 
CryoEM structure of the NADP-dependent malic enzyme MaeB

EMDB-36758: 
CryoEM structure of 3-phenylpropionate/cinnamic acid dioxygenase HcaE-HcaF complex

EMDB-36746: 
CryoEM structure of the Salmonella effector inositol phosphate phosphatase SopB

PDB-8jzl: 
CryoEM structure of the Salmonella effector inositol phosphate phosphatase SopB
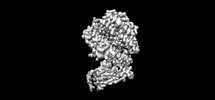
EMDB-38784: 
The structure of fox ACE2 and PT RBD complex

EMDB-38792: 
The structure of fox ACE2 and SARS-CoV RBD complex

PDB-8xyz: 
The structure of fox ACE2 and PT RBD complex

PDB-8xzb: 
The structure of fox ACE2 and SARS-CoV RBD complex

EMDB-37105: 
ATP-bound hMRP5 outward-open
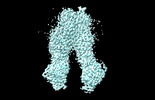
EMDB-37554: 
wt-hMRP5 inward-open
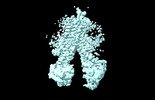
EMDB-37555: 
RD-hMRP5-inward open
EMDB-37556: 
ND-hMRP5-inward open

EMDB-37557: 
m6-hMRP5 inward open
EMDB-37558: 
M5PI-bound hMRP5

PDB-8kci: 
ATP-bound hMRP5 outward-open
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
