-Search query
-Search result
Showing all 44 items for (author: wang & yw)

EMDB-34190: 
Structure of X18 UFO protomer in complex with F6 Fab VHVL domain

EMDB-34192: 
HIV-1 Env X18 UFO in complex with F6 Fab

EMDB-34193: 
HIV-1 Env X18 UFO in complex with 8ANC195 Fab

EMDB-34194: 
HIV-1 Env X16 UFO in complex with 8ANC195 Fab

EMDB-35598: 
cryo-EM structure of the middle part of the shrimp white spot syndrome virus nucleocapsid (wide type)

EMDB-35600: 
Cryo-Em structure of the middle part of the shrimp white spot syndrome virus nucleocapsid (narrow type)

EMDB-28915: 
SIRT6 bound to an H3K9Ac nucleosome

PDB-8f86: 
SIRT6 bound to an H3K9Ac nucleosome

EMDB-33871: 
Cryo-EM structure of the INSL5-bound human relaxin family peptidereceptor 4 (RXFP4)-Gi complex

EMDB-33888: 
Cryo-EM structure of the compound 4-bound human relaxin family peptide receptor 4 (RXFP4)-Gi complex

EMDB-33889: 
Cryo-EM structure of the DC591053-bound human relaxin family peptide receptor 4 (RXFP4)-Gi complex

EMDB-33901: 
Structure of hIAPP-TF-type2

EMDB-33902: 
Structure of hIAPP-TF-type1

EMDB-34585: 
Molecular recognition of two endogenous hormones by the human parathyroid hormone receptor-1

EMDB-34587: 
PTHrP-PTH1R-Gs complex

EMDB-34598: 
Human parathyroid hormone receptor-1 dimer

EMDB-25726: 
Structure of the human FPR2-Gi complex with compound C43

EMDB-25727: 
Structure of the human FPR1-Gi complex with fMLFII

EMDB-25728: 
Structure of the human FPR2-Gi complex with CGEN-855A

EMDB-25729: 
Structure of the human FPR2-Gi complex with fMLFII

EMDB-22677: 
Structure of outer-arm dyneins bound to microtubule with microtubule binding state 1(MTBS-1)

EMDB-22679: 
Structure of outer-arm dynein bound to microtubule doublet in microtubule binding state 2 (MTBS-2)

EMDB-22840: 
Structure of the free outer-arm dynein in pre-parallel state

EMDB-24066: 
16-nm repeat microtubule doublet

EMDB-31459: 
DNQX-bound GluK2-1xNeto2 complex, with asymmetric LBD

EMDB-31460: 
Kainate-bound GluK2-1xNeto2 complex, at the desensitized state

EMDB-31462: 
DNQX-bound GluK2-1xNeto2 complex

EMDB-31463: 
DNQX-bound GluK2-2xNeto2 complex

EMDB-31464: 
LBD-TMD focused reconstruction of DNQX-bound GluK2-1xNeto2 complex

EMDB-31387: 
Cryo-EM structure of an activated Cholecystokinin A receptor (CCKAR)-Gi complex

EMDB-31388: 
Cryo-EM structure of an activated Cholecystokinin A receptor (CCKAR)-Gs complex

EMDB-31389: 
Cryo-EM structure of an activated Cholecystokinin A receptor (CCKAR)-Gq complex

EMDB-0938: 
Structure of CLHM1 from Caenorhabditis Elegans

EMDB-0833: 
cryo-em structure of alpha-synuclein fiber mutation type E46K

EMDB-0566: 
Subtomogram average of the Legionella pneumophila Dot/Icm type IV secretion system with DotF fused to a superfolder GFP (aligning the OM-complex)

EMDB-6988: 
cryo-em structure of alpha-synuclein fiber

EMDB-4230: 
Single particle cryo em structure of Mycobacterium tuberculosis RNA polymerase in complex with Fidaxomicin

EMDB-8566: 
In vivo structure of the Legionella pneumophila Dot/Icm type IV secretion system (aligning the outer membrane complex)

EMDB-8567: 
In vivo structure of the Legionella pneumophila Dot/Icm type IV secretion system (aligning the inner membrane complex)

EMDB-8568: 
In vivo structure of the Legionella pneumophila Dot/Icm type IV secretion system core complex (aligning the outer membrane complex)

EMDB-8569: 
In vivo structure of the Legionella pneumophila Dot/Icm type IV secretion system core complex (aligning the inner membrane part)
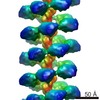
EMDB-2395: 
MuB is an AAA+ ATPase that forms helical filaments to control target selection for DNA transposition
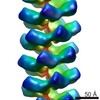
EMDB-2398: 
MuB is an AAA+ ATPase that forms helical filaments to control target selection for DNA transposition
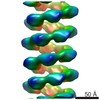
EMDB-2400: 
MuB is an AAA+ ATPase that forms helical filaments to control target selection for DNA transposition
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
