-Search query
-Search result
Showing 1 - 50 of 1,084 items for (author: hu & gq)

EMDB-37515: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with antidepressant desipramine in KCl condition.

EMDB-37520: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with norepinephrine in nanodisc.

EMDB-39729: 
Cryo-EM structure of human norepinephrine transporter NET in the presence of the antidepressant atomoxetine in an outward-open state at resolution of 3.4 angstrom.

PDB-8wgr: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with antidepressant desipramine in KCl condition.

PDB-8wgx: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with norepinephrine in nanodisc.

PDB-8z1l: 
Cryo-EM structure of human norepinephrine transporter NET in the presence of the antidepressant atomoxetine in an outward-open state at resolution of 3.4 angstrom.

EMDB-44482: 
Cryo-EM structure of the HIV-1 JR-FL IDL Env trimer in complex with PGT122 Fab
EMDB-44484: 
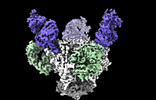
Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs
EMDB-44491: 
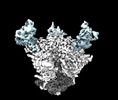
Cryo-EM structure of the HIV-1 WITO IDL Env trimer in complex with PGT122 Fab

PDB-9ber: 
Cryo-EM structure of the HIV-1 JR-FL IDL Env trimer in complex with PGT122 Fab

PDB-9bew: 
Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs

PDB-9bf6: 
Cryo-EM structure of the HIV-1 WITO IDL Env trimer in complex with PGT122 Fab
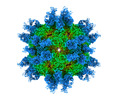

EMDB-32979: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)

PDB-7x35: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)

EMDB-37513: 
Cryo-EM structure of the red-shifted Fittonia albivenis PSI-LHCI

EMDB-38580: 
Structure of human class T GPCR TAS2R14-miniGs/gust complex with Aristolochic acid A.

EMDB-38582: 
Structure of human class T GPCR TAS2R14-DNGi complex with Aristolochic acid A.

EMDB-38583: 
Structure of human class T GPCR TAS2R14-Gi complex with Aristolochic acid A.

EMDB-38584: 
Structure of human class T GPCR TAS2R14-Gustducin complex with Aristolochic acid A.

EMDB-38586: 
Structure 2 of human class T GPCR TAS2R14-miniGs/gust complex with Flufenamic acid.

EMDB-38587: 
Structure of human class T GPCR TAS2R14-DNGi complex with Flufenamic acid.

EMDB-38588: 
Structure of human class T GPCR TAS2R14-Gi complex.

EMDB-39376: 
Structure of human class T GPCR TAS2R14-Ggustducin complex with agonist 28.1

PDB-8xql: 
Structure of human class T GPCR TAS2R14-miniGs/gust complex with Aristolochic acid A.

PDB-8xqn: 
Structure of human class T GPCR TAS2R14-DNGi complex with Aristolochic acid A.

PDB-8xqo: 
Structure of human class T GPCR TAS2R14-Gi complex with Aristolochic acid A.

PDB-8xqp: 
Structure of human class T GPCR TAS2R14-Gustducin complex with Aristolochic acid A.

PDB-8xqr: 
Structure 2 of human class T GPCR TAS2R14-miniGs/gust complex with Flufenamic acid.

PDB-8xqs: 
Structure of human class T GPCR TAS2R14-DNGi complex with Flufenamic acid.

PDB-8xqt: 
Structure of human class T GPCR TAS2R14-Gi complex.

PDB-8yky: 
Structure of human class T GPCR TAS2R14-Ggustducin complex with agonist 28.1

EMDB-35157: 
Cryo-EM structure of human norepinephrine transporter NET in the presence of the antidepressant escitalopram in an inward-open state at resolution of 2.85 angstrom.

PDB-8i3v: 
Cryo-EM structure of human norepinephrine transporter NET in the presence of the antidepressant escitalopram in an inward-open state at resolution of 2.85 angstrom.

EMDB-38460: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 spike protein(6P), 1-RBD-up state

EMDB-38463: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5 spike protein(6P), RBD-closed state

EMDB-38476: 
Cryo-EM structure of SARS-CoV-2 Omicron HV.1 spike protein(6P), RBD-closed state

EMDB-38488: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5.1 spike protein(6P), RBD-closed state

EMDB-38495: 
SARS-CoV-2 Omicron EG.5.1 RBD in complex with human ACE2 (local refined from the spike protein)

EMDB-38496: 
SARS-CoV-2 Omicron HV.1 RBD in complex with human ACE2 (local refinement from the spike protein)

EMDB-38498: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5.1 spike protein(6P) in complex with human ACE2

EMDB-38502: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 spike protein(6P) in complex with human ACE2

EMDB-38505: 
Cryo-EM structure of SARS-CoV-2 Omicron HV.1 spike protein(6P) in complex with human ACE2

EMDB-38826: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 spike protein in complex with human ACE2

EMDB-38827: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 RBD in complex with human ACE2 (local refinement from the spike protein)

EMDB-38937: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 spike protein

EMDB-38983: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 RBD in complex with human ACE2 and S309 Fab

PDB-8xlv: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 spike protein(6P), 1-RBD-up state

PDB-8xm5: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5 spike protein(6P), RBD-closed state

PDB-8xmg: 
Cryo-EM structure of SARS-CoV-2 Omicron HV.1 spike protein(6P), RBD-closed state

PDB-8xmt: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5.1 spike protein(6P), RBD-closed state
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
