[English] 日本語
 Yorodumi
Yorodumi- EMDB-44484: Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex wi... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs | |||||||||
 Map data Map data | sharpened | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CD4 / HIV-1 / SOSIP / Vaccine / gp120 / gp41 / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Gorman J / Kwong PD | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2024 Journal: Cell Rep / Year: 2024Title: Design of soluble HIV-1 envelope trimers free of covalent gp120-gp41 bonds with prevalent native-like conformation. Authors: Peng Zhang / Jason Gorman / Yaroslav Tsybovsky / Maolin Lu / Qingbo Liu / Vinay Gopan / Mamta Singh / Yin Lin / Huiyi Miao / Yuna Seo / Alice Kwon / Adam S Olia / Gwo-Yu Chuang / Hui Geng / ...Authors: Peng Zhang / Jason Gorman / Yaroslav Tsybovsky / Maolin Lu / Qingbo Liu / Vinay Gopan / Mamta Singh / Yin Lin / Huiyi Miao / Yuna Seo / Alice Kwon / Adam S Olia / Gwo-Yu Chuang / Hui Geng / Yen-Ting Lai / Tongqing Zhou / John R Mascola / Walther Mothes / Peter D Kwong / Paolo Lusso /   Abstract: Soluble HIV-1 envelope (Env) trimers may serve as effective vaccine immunogens. The widely utilized SOSIP trimers have been paramount for structural studies, but the disulfide bond they feature ...Soluble HIV-1 envelope (Env) trimers may serve as effective vaccine immunogens. The widely utilized SOSIP trimers have been paramount for structural studies, but the disulfide bond they feature between gp120 and gp41 constrains intersubunit mobility and may alter antigenicity. Here, we report an alternative strategy to generate stabilized soluble Env trimers free of covalent gp120-gp41 bonds. Stabilization was achieved by introducing an intrasubunit disulfide bond between the inner and outer domains of gp120, defined as interdomain lock (IDL). Correctly folded IDL trimers displaying a native-like antigenic profile were produced for HIV-1 Envs of different clades. Importantly, the IDL design abrogated CD4 binding while not affecting recognition by potent neutralizing antibodies to the CD4-binding site. By cryoelectron microscopy, IDL trimers were shown to adopt a closed prefusion configuration, while single-molecule fluorescence resonance energy transfer documented a high prevalence of native-like conformation. Thus, IDL trimers may be promising candidates as vaccine immunogens. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44484.map.gz emd_44484.map.gz | 155.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44484-v30.xml emd-44484-v30.xml emd-44484.xml emd-44484.xml | 27.3 KB 27.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_44484.png emd_44484.png | 86.1 KB | ||
| Masks |  emd_44484_msk_1.map emd_44484_msk_1.map | 166.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-44484.cif.gz emd-44484.cif.gz | 7.4 KB | ||
| Others |  emd_44484_additional_1.map.gz emd_44484_additional_1.map.gz emd_44484_additional_2.map.gz emd_44484_additional_2.map.gz emd_44484_half_map_1.map.gz emd_44484_half_map_1.map.gz emd_44484_half_map_2.map.gz emd_44484_half_map_2.map.gz | 43.1 MB 21.6 MB 154.5 MB 154.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44484 http://ftp.pdbj.org/pub/emdb/structures/EMD-44484 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44484 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44484 | HTTPS FTP |
-Related structure data
| Related structure data |  9bewMC  9berC  9bf6C C: citing same article ( M: atomic model generated by this map |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_44484.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44484.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0961 Å | ||||||||||||||||||||||||||||||||||||
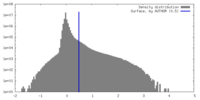
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_44484_msk_1.map emd_44484_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: unsharpened
| File | emd_44484_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_44484_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_44484_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A
| File | emd_44484_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM Structure of the HIV-1 BG505 IDL Env trimer in complex wi...
| Entire | Name: Cryo-EM Structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM Structure of the HIV-1 BG505 IDL Env trimer in complex wi...
| Supramolecule | Name: Cryo-EM Structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Macromolecule #1: Envelope glycoprotein gp120
| Macromolecule | Name: Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 54.12034 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETEKHNVWA THACVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHTDIIS LWCQSLKPC VKLTPLCVTL QCTNVTNNIT DDMRGELKNC SFNMTTELRD KKQKVYSLFY RLDVVQINEN QGNRSNNSNK E YRLINCNT ...String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETEKHNVWA THACVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHTDIIS LWCQSLKPC VKLTPLCVTL QCTNVTNNIT DDMRGELKNC SFNMTTELRD KKQKVYSLFY RLDVVQINEN QGNRSNNSNK E YRLINCNT SAITQACPKV SFEPIPIHYC APAGFAILKC KDKKFNGTGP CPNVSTVQCT HGIKPVVSTQ LLLNGSLAEE EV MIRSENI TNNAKNILVQ FNTPVQINCT RPNNNTRKSI RIGPGQAFYA TGDIIGDIRQ AHCNVSKATW NETLGKVVKQ LRK HFGNNT IIRFANSSGG DLEVTTHSFN CGGEFFYCNT SGLFNSTWIS NTSVQGSNST GSNDSITLPC RIKQIINMWQ GCGI GQAMY APPIQGVIRC VSNITGLILT RDGGSTNSTN ETFRPGGGDM RDNWRSELYK YKVVKIEPLG VAPTRAKRRV VGRRR RRR |
-Macromolecule #2: Envelope glycoprotein gp41
| Macromolecule | Name: Envelope glycoprotein gp41 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 17.114418 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVGIGAVFLG FLGAAGSTMG AASMTLTVQA RNLLSGIVQQ QSNLLRAPEA QQHLLKLTVW GIKQLQARVL AVERYLRDQQ LLGIWGCSG KLICATNVPW NSSWSNRNLS EIWDNMTWLQ WDKEISNYTQ IIYGLLEESQ NQQEKNEQDL LALD |
-Macromolecule #3: 3BNC117 heavy chain
| Macromolecule | Name: 3BNC117 heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.656484 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLLQSGAA VTKPGASVRV SCEASGYNIR DYFIHWWRQA PGQGLQWVGW INPKTGQPNN PRQFQGRVSL TRHASWDFDT YSFYMDLKA LRSDDTAVYF CARQRSDYWD FDVWGSGTQV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA ...String: QVQLLQSGAA VTKPGASVRV SCEASGYNIR DYFIHWWRQA PGQGLQWVGW INPKTGQPNN PRQFQGRVSL TRHASWDFDT YSFYMDLKA LRSDDTAVYF CARQRSDYWD FDVWGSGTQV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA LTSGVHTFPA VLQSSGLYSL SSVVTVPSSS LGTQTYICNV NHKPSNTKVD KKVEPKSC |
-Macromolecule #4: 3BNC117 light chain
| Macromolecule | Name: 3BNC117 light chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.022658 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIQMTQSPSS LSASVGDTVT ITCQANGYLN WYQQRRGKAP KLLIYDGSKL ERGVPSRFSG RRWGQEYNLT INNLQPEDIA TYFCQVYEF VVPGTRLDLK RTVAAPSVFI FPPSDEQLKS GTASVVCLLN NFYPREAKVQ WKVDNALQSG NSQESVTEQD S KDSTYSLS ...String: DIQMTQSPSS LSASVGDTVT ITCQANGYLN WYQQRRGKAP KLLIYDGSKL ERGVPSRFSG RRWGQEYNLT INNLQPEDIA TYFCQVYEF VVPGTRLDLK RTVAAPSVFI FPPSDEQLKS GTASVVCLLN NFYPREAKVQ WKVDNALQSG NSQESVTEQD S KDSTYSLS STLTLSKADY EKHKVYACEV THQGLSSPVT KSFNRGEC |
-Macromolecule #5: 10-1074 heavy chain
| Macromolecule | Name: 10-1074 heavy chain / type: protein_or_peptide / ID: 5 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.316314 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLQESGPG LVKPSETLSV TCSVSGDSMN NYYWTWIRQS PGKGLEWIGY ISDRESATYN PSLNSRVVIS RDTSKNQLSL KLNSVTPAD TAVYYCATAR RGQRIYGVVS FGEFFYYYSM DVWGKGTTVT VSSASTKGPS VFPLAPSSKS TSGGTAALGC L VKDYFPEP ...String: QVQLQESGPG LVKPSETLSV TCSVSGDSMN NYYWTWIRQS PGKGLEWIGY ISDRESATYN PSLNSRVVIS RDTSKNQLSL KLNSVTPAD TAVYYCATAR RGQRIYGVVS FGEFFYYYSM DVWGKGTTVT VSSASTKGPS VFPLAPSSKS TSGGTAALGC L VKDYFPEP VTVSWNSGAL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTQTYICNVN HKPSNTKVDK RVEPKSC |
-Macromolecule #6: 10-1074 light chain
| Macromolecule | Name: 10-1074 light chain / type: protein_or_peptide / ID: 6 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.18076 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: SYVRPLSVAL GETARISCGR QALGSRAVQW YQHRPGQAPI LLIYNNQDRP SGIPERFSGT PDINFGTRAT LTISGVEAGD EADYYCHMW DSRSGFSWSF GGATRLTVLG QPKAAPSVTL FPPSSEELQA NKATLVCLIS DFYPGAVTVA WKADSSPVKA G VETTTPSK ...String: SYVRPLSVAL GETARISCGR QALGSRAVQW YQHRPGQAPI LLIYNNQDRP SGIPERFSGT PDINFGTRAT LTISGVEAGD EADYYCHMW DSRSGFSWSF GGATRLTVLG QPKAAPSVTL FPPSSEELQA NKATLVCLIS DFYPGAVTVA WKADSSPVKA G VETTTPSK QSNNKYAASS YLSLTPEQWK SHRSYSCQVT HEGSTVEKTV APTECS |
-Macromolecule #12: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 12 / Number of copies: 39 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: PBS |
| Grid | Model: C-flat-1.2/1.3 / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Average exposure time: 10.0 sec. / Average electron dose: 63.9 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.4 µm / Nominal defocus min: 0.16 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)