-Search query
-Search result
Showing 1 - 50 of 83 items for (author: foerster & f)

EMDB-53127: 
Cryo-EM structure of natively purified Rubrerythrin isolated from OSIT assemblies of the anaerobic extremophile P. furiosus
Method: single particle / : Skalidis I, Song W, Howes S, Foerster F

PDB-9qg1: 
Natively purified Rubrerythrin 16-mer from the anaerobic extremophile P. furiosus
Method: single particle / : Skalidis I, Song W, Howes S, Foerster F

EMDB-13477: 
Small subunit of the Chlamydomonas reinhardtii mitoribosome
Method: single particle / : Waltz F, Soufari H, Hashem Y

EMDB-13481: 
Small subunit of the Chlamydomonas mitoribosome - head focus
Method: single particle / : Waltz F, Salinas-Giege T, Englmeier R, Meichel H, Soufari H, Kuhn L, Pfeffer S, Foerster F, Engel BD, Giege P, Drouard L, Hashem Y

EMDB-13578: 
C.reinhardtii mitochondrial ribosome (in situ)
Method: subtomogram averaging / : Waltz F, Salinas-Giege T, Englmeier R, Meichel H, Soufari H, Kuhn L, Pfeffer S, Foerster F, Engel BD, Giege P, Drouard L, Hashem Y

EMDB-13171: 
Human Signal Peptidase Complex Paralog A (SPC-A)
Method: single particle / : Liaci AM, Foerster F

EMDB-13172: 
Human Signal Peptidase Complex Paralog C (SPC-C)
Method: single particle / : Liaci AM, Foerster F

PDB-7p2p: 
Human Signal Peptidase Complex Paralog A (SPC-A)
Method: single particle / : Liaci AM, Foerster F

PDB-7p2q: 
Human Signal Peptidase Complex Paralog C (SPC-C)
Method: single particle / : Liaci AM, Foerster F

PDB-7akd: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 47D11 neutralizing antibody Fab fragment
Method: single particle / : Fedry J, Hurdiss DL, Wang C, Li W, Obal G, Drulyte I, Howes SC, van Kuppeveld FJM, Foerster F, Bosch BJ

PDB-7akj: 
Structure of the SARS-CoV spike glycoprotein in complex with the 47D11 neutralizing antibody Fab fragment
Method: single particle / : Fedry J, Hurdiss DL, Wang C, Li W, Obal G, Drulyte I, Howes SC, van Kuppeveld FJM, Foerster F, Bosch BJ

EMDB-12749: 
In situ cryo-electron tomogram of a pyrenoid inside a Chlamydomonas reinhardtii cell
Method: electron tomography / : Cuellar LK, Schaffer M, Strauss M, Martinez-Sanchez A, Plitzko JM, Foerster F, Engel BD

EMDB-11992: 
the molecular sociology at the HeLa cell nuclear periphery
Method: electron tomography / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-4306: 
Subtomogram average of unsorted ribosome-translocon complexes from STT3B(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4307: 
Subtomogram average of OST-free ribosome-translocon complexes from STT3B(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4308: 
Subtomogram average of OST-containing ribosome-translocon complexes from STT3B(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
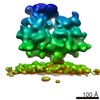
EMDB-4309: 
Subtomogram average of unsorted ribosome-translocon complexes from STT3A(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
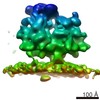
EMDB-4310: 
Subtomogram average of OST-free ribosome-translocon complexes from STT3A(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4311: 
Subtomogram average of ribosome-translocon complexes from STT3A(-/-) HEK cells containing an unidentified ER-lumenal component
Method: subtomogram averaging / : Pfeffer S, Foerster F
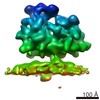
EMDB-4312: 
Subtomogram average of unsorted ribosome-translocon complexes from wild type HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
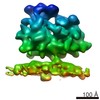
EMDB-4313: 
Subtomogram average of OST-free ribosome-translocon complexes from wild type HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
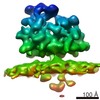
EMDB-4314: 
Subtomogram average of OST-containing ribosome-translocon complexes from wild type HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4315: 
Subtomogram average of OST-containing ribosome-translocon complexes from canine rough microsomal membranes
Method: subtomogram averaging / : Pfeffer S, Foerster F

PDB-6ftg: 
Subtomogram average of OST-containing ribosome-translocon complexes from canine rough microsomal membranes
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-3784: 
Structure of the membrane-bound human mitoribosome
Method: subtomogram averaging / : Englmeier R, Pfeffer S, Foerster F

EMDB-3855: 
CryoEM structure of recombinant CMV particles with Tetanus-epitope
Method: single particle / : Kotecha A, Stuart DI

PDB-5ow6: 
CryoEM structure of recombinant CMV particles with Tetanus-epitope
Method: single particle / : Kotecha A, Stuart DI, Backmann M

EMDB-3694: 
In situ subtomogram average of Rubisco within the Chlamydomonas pyrenoid
Method: subtomogram averaging / : Cuellar LK, Schaffer M, Strauss M, Martinez-Sanchez A, Plitzko JM, Foerster F, Engel BD

EMDB-3603: 
KaiCB circadian clock backbone model based on a Cryo-EM density
Method: single particle / : Schuller JM, Snijder J, Loessl P, Heck AJR, Foerster F

PDB-5n8y: 
KaiCBA circadian clock backbone model based on a Cryo-EM density
Method: single particle / : Schuller JM, Snijder J, Loessl P, Heck AJR, Foerster F

PDB-5mp9: 
26S proteasome in presence of ATP (s1)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

PDB-5mpa: 
26S proteasome in presence of ATP (s2)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

PDB-5mpb: 
26S proteasome in presence of AMP-PNP (s3)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

PDB-5mpc: 
26S proteasome in presence of BeFx (s4)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

PDB-5mpd: 
26S proteasome in presence of ATP (s1)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

PDB-5mpe: 
26S proteasome in presence of ATP (s2)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

EMDB-4143: 
Subtomogram average of the ER membrane-associated ribosome from human TRAPdelta-deficient fibroblasts
Method: subtomogram averaging / : Pfeffer S, Dudek J, Ng BG, Zimmermann R, Freeze HH, Foerster F

EMDB-4144: 
Subtomogram average of the ER membrane-associated ribosome from human TRAPgamma-deficient fibroblasts
Method: subtomogram averaging / : Pfeffer S, Dudek J, Ng BG, Zimmermann R, Freeze HH, Foerster F

EMDB-4145: 
Subtomogram average of the ER membrane-associated ribosome from FIB-milled C. reinhardtii cells
Method: subtomogram averaging / : Pfeffer S, Schaffer M, Albert S, Engel BD, Foerster F

EMDB-3418: 
Subtomogram average of 80S ribosomes obtained using the VPP
Method: subtomogram averaging / : Khoshouei M, Pfeffer S, Baumeister W, Foerster F, Danev R

EMDB-3419: 
Subtomogram average of 80S ribosomes obtained using CTEM at one defocus
Method: subtomogram averaging / : Khoshouei M, Pfeffer S, Baumeister W, Foerster F, Danev R

EMDB-3420: 
Subtomogram average of 80S ribosomes obtained using CTEM with a range of defocus values
Method: subtomogram averaging / : Khoshouei M, Pfeffer S, Baumeister W, Foerster F, Danev R

EMDB-8054: 
In situ sub-tomogram average of HeLa nuclear pore complex from a single cell
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-8055: 
In situ sub-tomogram average of HeLa nuclear pore complex
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-8056: 
In situ sub-tomogram average of HeLa ER-associated ribosomes
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-8057: 
In situ sub-tomogram average of HeLa cytosolic ribosomes
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-3323: 
p97 in the ADP state C6 symmetrized
Method: single particle / : Schuller JM, Beck F, Loessl P, Heck Albert JR, Foerster F

EMDB-3324: 
p97 in the AMPPNP state C6 symmetrized
Method: single particle / : Schuller JM, Beck F, Loessl P, Heck Albert JR, Foerster F

EMDB-3325: 
p97 in the ATPyS state C6 symmetrized
Method: single particle / : Schuller JM, Beck F, Loessl P, Heck Albert JR, Foerster F

EMDB-3326: 
p97 in the ADP state no symmetry applied
Method: single particle / : Schuller JM, Beck F, Loessl P, Heck Albert JR, Foerster F
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
