+ Open data
Open data
 Loading...
Loading...
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Small subunit of the Chlamydomonas reinhardtii mitoribosome | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Mitochondria / mitoribosome / alga / RIBOSOME | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationmitochondrial small ribosomal subunit / chloroplast / small ribosomal subunit / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / rRNA binding / structural constituent of ribosome / ribosome / translation / mitochondrial matrix ...mitochondrial small ribosomal subunit / chloroplast / small ribosomal subunit / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / rRNA binding / structural constituent of ribosome / ribosome / translation / mitochondrial matrix / ribonucleoprotein complex / mRNA binding / mitochondrion / RNA binding / nucleus / cytoplasm Similarity search - Function : / Mitochondrial glycoprotein / Mitochondrial glycoprotein superfamily / Mitochondrial glycoprotein / : / Ribosomal protein S23/S29, mitochondrial / Mitochondrial ribosomal death-associated protein 3 / Collagen triple helix repeat / Collagen triple helix repeat (20 copies) / 30S ribosomal protein Thx ...: / Mitochondrial glycoprotein / Mitochondrial glycoprotein superfamily / Mitochondrial glycoprotein / : / Ribosomal protein S23/S29, mitochondrial / Mitochondrial ribosomal death-associated protein 3 / Collagen triple helix repeat / Collagen triple helix repeat (20 copies) / 30S ribosomal protein Thx / 30S ribosomal protein Thx / 30S ribosomal protein / Ribosomal protein S19, bacterial-type / Ribosomal protein S6, plastid/chloroplast / Ribosomal protein S2, bacteria/mitochondria/plastid / Ribosomal protein S16 / Ribosomal protein S16 domain superfamily / Ribosomal protein S16 / Ribosomal protein S15, bacterial-type / Ribosomal protein S6 / Ribosomal protein S6 / Ribosomal protein S6 superfamily / Ribosomal protein S12, bacterial-type / Translation elongation factor EF1B/ribosomal protein S6 / Ribosomal protein S18 / Ribosomal protein S18 / Ribosomal protein S18 superfamily / Prokaryotic membrane lipoprotein lipid attachment site profile. / Ribosomal protein S3, C-terminal / Ribosomal protein S3, C-terminal domain / Ribosomal protein S3, C-terminal domain superfamily / Ribosomal protein S15/S19, conserved site / Ribosomal protein S19 signature. / Ribosomal protein S10 / Ribosomal protein S19/S15 / Ribosomal protein S19/S15, superfamily / Ribosomal protein S19 / Ribosomal protein S2 / Ribosomal protein S2, flavodoxin-like domain superfamily / Ribosomal protein S2 / Ribosomal protein S5 / S5 double stranded RNA-binding domain profile. / Ribosomal protein S5, N-terminal / Ribosomal protein S5, C-terminal / Ribosomal protein S5, N-terminal domain / Ribosomal protein S13 / 30s ribosomal protein S13, C-terminal / Ribosomal protein S13/S18 / Ribosomal protein S13 family profile. / Ribosomal protein S5, C-terminal domain / Ribosomal protein S14 / Ribosomal protein S14p/S29e / Ribosomal protein S13-like, H2TH / Ribosomal protein S10 domain / Ribosomal protein S10 domain superfamily / Ribosomal protein S10p/S20e / Ribosomal protein S9, conserved site / Ribosomal protein S9 signature. / Ribosomal protein S11 / Ribosomal protein S12 signature. / Ribosomal protein S11 / Ribosomal protein S5/S7 / Ribosomal protein S7 domain / Ribosomal protein S7 domain superfamily / Ribosomal protein S7p/S5e / Ribosomal protein S9 / Ribosomal protein S9/S16 / Ribosomal protein S12/S23 / Ribosomal protein S12/S23 / Ribosomal protein S17/S11 / Ribosomal protein S17 / Ribosomal protein S15 / Ribosomal_S15 / Ribosomal protein S15 / Ribosomal protein S11 superfamily / S15/NS1, RNA-binding / Ribosomal protein S5 domain 2-type fold, subgroup / Ribosomal protein S5 domain 2-type fold / Nucleic acid-binding, OB-fold / P-loop containing nucleoside triphosphate hydrolase Similarity search - Domain/homology Small ribosomal subunit protein uS10 domain-containing protein / Uncharacterized protein / Small ribosomal subunit protein uS15c / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Small ribosomal subunit protein uS7 domain-containing protein / Small ribosomal subunit protein uS3 C-terminal domain-containing protein / Mitochondrial ribosomal protein S16 / Mitochondrial glycoprotein ...Small ribosomal subunit protein uS10 domain-containing protein / Uncharacterized protein / Small ribosomal subunit protein uS15c / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Small ribosomal subunit protein uS7 domain-containing protein / Small ribosomal subunit protein uS3 C-terminal domain-containing protein / Mitochondrial ribosomal protein S16 / Mitochondrial glycoprotein / Small ribosomal subunit protein bS18c / S5 DRBM domain-containing protein / Small ribosomal subunit protein uS19c / Mitochondrial ribosomal protein S17 / Uncharacterized protein / Mitochondrial ribosomal protein S11 / Small ribosomal subunit protein mS29 / Uncharacterized protein / Uncharacterized protein / Small ribosomal subunit protein uS9c / Ribosomal protein S12, mitochondrial / S1 motif domain-containing protein / Small ribosomal subunit protein mS23 / Uncharacterized protein / : / Uncharacterized protein / : / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein Similarity search - Component | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||||||||
 Authors Authors | Waltz F / Soufari H / Hashem Y | |||||||||||||||
| Funding support |  France, 4 items France, 4 items
| |||||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: The Chlamydomonas mitochondrial ribosome: how to build a ribosome from RNA fragments Authors: Waltz F / Salinas-Giege T / Englmeier R / Meichel H / Soufari H / Kuhn L / Pfeffer S / Foerster F / Engel BD / Giege P / Drouard L / Hashem Y | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13477.map.gz emd_13477.map.gz | 164.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13477-v30.xml emd-13477-v30.xml emd-13477.xml emd-13477.xml | 68.3 KB 68.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13477.png emd_13477.png | 69.7 KB | ||
| Filedesc metadata |  emd-13477.cif.gz emd-13477.cif.gz | 14.1 KB | ||
| Others |  emd_13477_half_map_1.map.gz emd_13477_half_map_1.map.gz emd_13477_half_map_2.map.gz emd_13477_half_map_2.map.gz | 140.4 MB 140.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13477 http://ftp.pdbj.org/pub/emdb/structures/EMD-13477 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13477 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13477 | HTTPS FTP |
-Related structure data
| Related structure data |  7pkqMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13477.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13477.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.13 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_13477_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
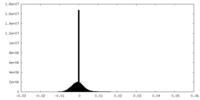
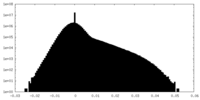
| Density Histograms |
-Half map: #2
| File | emd_13477_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
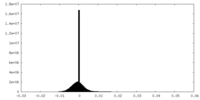
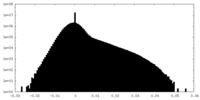
| Density Histograms |
- Sample components
Sample components
+Entire : Small subunit of the Chlamydomonas reinhardtii mitochondrial ribosome
| Entire | Name: Small subunit of the Chlamydomonas reinhardtii mitochondrial ribosome |
|---|---|
| Components |
|
+Supramolecule #1: Small subunit of the Chlamydomonas reinhardtii mitochondrial ribosome
| Supramolecule | Name: Small subunit of the Chlamydomonas reinhardtii mitochondrial ribosome type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#42 |
|---|---|
| Source (natural) | Organism:  |
+Macromolecule #1: mS35
| Macromolecule | Name: mS35 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 88.892773 KDa |
| Sequence | String: MTLSTQRRAL LAGFGKARGG QWTQEGIAAL HSTTTTNAQV ETQPDGELAE SSADDASRLF QELSRRNRPN SAGTEPGPSP RPVLAQLPA AAELMERAAG TPSSQALYDL PTYLSVTHPH ARVEPRNPAY DWRRSPQLEA GGPRRAALLL AVDAHMAAPE S REALLRMA ...String: MTLSTQRRAL LAGFGKARGG QWTQEGIAAL HSTTTTNAQV ETQPDGELAE SSADDASRLF QELSRRNRPN SAGTEPGPSP RPVLAQLPA AAELMERAAG TPSSQALYDL PTYLSVTHPH ARVEPRNPAY DWRRSPQLEA GGPRRAALLL AVDAHMAAPE S REALLRMA QLELAHLWYY QQHAAELPAA ASPSASAASA ASTATPDAAA AGQRRGGVAK QREAEPAAAS TSAAAGDKAQ AG AGTGTGA GAGAEAEAGD DDIFAAADRK ARERAAAVEA ATAAAASSAA KSLGRRGGRL PAELEARVRD MQLRYGMASR DAA SVLRLL ADSFSRRLPG GGRRLEGEAE AEAAVGLGEA EAEAGGDPTA ALTWALSGGG RGGSGGTISG LSRHLAKQAA QAKA ADRAI ITAMQSLADA VNTASSSSSS ATASSSTASS SGSSAYWAAH PALAYDQTLA ARAHRQGLAW RAAAEAGPDG AASLS AAIT ALSASLRRPT SSPTSSASAS SPPATPVLDL VLDYLQAKYA DLLTAWETAA RLREATERVS AAVARARATV PMSAVP PLP PALAAELERS WQNAAAAFRP ALLQPLLQPA GSGAAARNSL AAAAASGSPA LLQAQSSLTR PLDWAKAKAL IEQHYAK QQ QAAALAGATA AAEATAAAAA AASAAAAAGS PRGAVEALLQ PLLQRAADRH RALLAGGDAA AAEAAAAAAA AAAHGSST A AAGGAAPSEG AASVVCFRET LTLDASSAYA NSSFDSRVTL EFNVDRLAAA EPGLGGAWGA RFLERLLATP ALPGSGRRI RGGGGSSGMF GRARASPAAR AAANAAAYPV DVEHSFCRRT RTAALSTARY GSREANRKLL LEHYNELLRA AALAAAAGAS AAV UniProtKB: Uncharacterized protein |
+Macromolecule #2: mS45
| Macromolecule | Name: mS45 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 102.234461 KDa |
| Sequence | String: MRARLFQSLA IQVAEEGLLT TALSGYACRA FSANGVTKFG FQCSSVSTSA ASHEASSSRG SGDQGRVTGG RQERAGRQAP TGSDGPQRA DRAKQGYADG GKSRSLFRRD NNPDSGSRGG DGQRGGGKGA GTGSGYSGGS SGQRGGPWQQ QQGGRQHDRR G DDRRGGGG ...String: MRARLFQSLA IQVAEEGLLT TALSGYACRA FSANGVTKFG FQCSSVSTSA ASHEASSSRG SGDQGRVTGG RQERAGRQAP TGSDGPQRA DRAKQGYADG GKSRSLFRRD NNPDSGSRGG DGQRGGGKGA GTGSGYSGGS SGQRGGPWQQ QQGGRQHDRR G DDRRGGGG GGGPPPMLRD GRDGRDGRGG GGRPPFAPSG PGSGSRAPRA ARWDALDVAA SNAPQRLLQL VGDPRSGVAD DT SSPAFGE VTVELLHHQP SNEVYWYKYA FAGGQPGGRG GHRDGRLSDY SRNLAYVLRA KDPARWSVEA LADKFRVRPQ RML AALALK ELEAQRLESG ALLRGPLSAY SLRVSLADVH LDKASGAPRL RPASAPPAAS RELLLGRMLP RVVSATQRTQ AALL SALRP HGFDAAALRA ALQADVAAVE AAAGELDAAA AAAGGEGAEG GTAAGGVSAY LAPELAACRA AVEAVAAAVE VVVNA SPLL QEERQAAALR ARVEAALQAG AGQAPAAAAA VAGTGAGAGG GADGNSKDAV PAAAAAAATS PGGGEALEVL LQPLTQ DGS GAPLQRFLFS LPPPIRVALL RQLPRLAAQL AAAGVDLQRA GRDAAYRGPL APAAPAATPA GSKQQQQQRQ KQGGAAA ST RSGKLPRLTV DVLGFVPVSE AQAEALAAAV ESAAAAEAAL VAKLEALQVA AGAAPAPAAG AAAAAAAAAA TAPSAAEL R DARSLLRQYL STHSMWGELR LAARKAAGHS PDTRGKRRKL EVMAEADPAA LTRALQVEAH TQLRRSHDRA AAESLKAGV AAAAATEAAA AAAASGSEAA AKPPTPLSDP AGWDALAAWL DAEVAGRVYA RGSGERLVVR LPTYPAFEGY PLSELDKARE GELGALNRL VAARTAAAQW AAFRRDLLFN LGVRGEKLRD NDLPGAPAWP ANLRTELTRP VVVYGIGPDG ATQYPPVYVA A GDGAKRPL NEQEKVYQER RLPGPRLPYH MGRIRTLPEL N UniProtKB: Uncharacterized protein |
+Macromolecule #3: mS45-insert
| Macromolecule | Name: mS45-insert / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 36.187551 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) |
+Macromolecule #4: mS31/46
| Macromolecule | Name: mS31/46 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.847501 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #5: mS105
| Macromolecule | Name: mS105 / type: protein_or_peptide / ID: 5 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 28.879045 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)M ATRQRLMRGI AGLLHQQGQQ SSCSQLLMTS AAAASASASS VATGPSHQPA LLLRAGSALR GVTTSAPTF NGSLSSLLRE ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)M ATRQRLMRGI AGLLHQQGQQ SSCSQLLMTS AAAASASASS VATGPSHQPA LLLRAGSALR GVTTSAPTF NGSLSSLLRE EVDYERKNYE RPAQISGGPP APFKLTEAPG DTLLTLTRTF GAEEINVDLH VNSQPSPEYD GEDGDEGISV VAFNVSVAK GDRVLLFECE SDGNSVNINH VSLEPKEGLG SESMYSGPVF DELDDNLQGQ FGKYLEDRGI TAELGEYLRF L IYDKEQRE YQNWLSEVEA FVGGK UniProtKB: Mitochondrial glycoprotein |
+Macromolecule #6: mS106
| Macromolecule | Name: mS106 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 51.927285 KDa |
| Sequence | String: MDLIGRGGAP SASALGTDAL VLLCGPEAQA GISTCSAAAE SAASPQSSCS WVGTAQHSSA TRAEAAGPSA PCGPAPLLRN LRTPSQSGR LSGAPPTWIS AAAASLGNSP RFAASARCST ADVDVSRSGL EDASALGDCV DRWSTPAATV SGIALLHQYH N HHHNHHHI ...String: MDLIGRGGAP SASALGTDAL VLLCGPEAQA GISTCSAAAE SAASPQSSCS WVGTAQHSSA TRAEAAGPSA PCGPAPLLRN LRTPSQSGR LSGAPPTWIS AAAASLGNSP RFAASARCST ADVDVSRSGL EDASALGDCV DRWSTPAATV SGIALLHQYH N HHHNHHHI GHSSSSCGSV ASSTSSVPGS AAGSPFPPRP SLPSSATNSS RFLMQSQSQQ LRSVSFTAAT AAPKAAAKGA SA KAGSSSS SGAAAPSPAA AALRTPEWAR VPGHLHELQD LYRKRQKRMA ALRHGAQVEL EAREAVAWKT GGRGRAVAAA AKA ALDALP LPPPEDGGKA ELAAALAADT TFAAAHEALT GLLRRGVVLD AAEHLAPLLR KAGDAGQLAS ALGVAAANHL SQAA RQLGA HSPRHGPLHA VLVQECRRLK AAAPLVTLWE SLHEHGLAPE AADAAAAVRA AVELGDGGAA VRLLMLACMY GEAPL AGAA EAGAVLQLLQ QGNPDQAAQL RELLPKLGLR GA UniProtKB: Uncharacterized protein |
+Macromolecule #7: mS107
| Macromolecule | Name: mS107 / type: protein_or_peptide / ID: 7 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 43.833246 KDa |
| Sequence | String: MRKRELLNEA RALVPEGSGW LEAYTRNISP RQLTWRLGKR DSLAAMTEGW QLYQGKFDTV AMAALLRRLR HAQLQDPGFD PLAAQRLLD DLVPRLRSVG LRFGKLRDIT AYLHALAKLR SPAPSASSSA ASPRAGAAAA SLLTQPDALV LDLAVFATRN R TELLHASP ...String: MRKRELLNEA RALVPEGSGW LEAYTRNISP RQLTWRLGKR DSLAAMTEGW QLYQGKFDTV AMAALLRRLR HAQLQDPGFD PLAAQRLLD DLVPRLRSVG LRFGKLRDIT AYLHALAKLR SPAPSASSSA ASPRAGAAAA SLLTQPDALV LDLAVFATRN R TELLHASP QRLATLLWAL MRLLPPQLYG SEQLQVVLDR MALASLGRLQ NFAPLDLRWA ALAFATFGPH GPSSKATATT TA AAAGTRA AAGAGSGPAL PEWPRVVRGE AAAAATAGAQ GAEDVTARRQ SRNARVIKAL CDELAARSGN LALPQPEPRD LAL AAHALG LVAAASSSAG GAVAPPALLV KAAGVAARSL PALSGEEAVG LVETLAVWGL RQPALLEALR DAAGRWGEGE QAEA LRGRL QAAYTRLGVE L UniProtKB: Uncharacterized protein |
+Macromolecule #8: Unk1
| Macromolecule | Name: Unk1 / type: protein_or_peptide / ID: 8 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 2.826475 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK) |
+Macromolecule #9: Unk2
| Macromolecule | Name: Unk2 / type: protein_or_peptide / ID: 9 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.95396 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) |
+Macromolecule #10: Unk3
| Macromolecule | Name: Unk3 / type: protein_or_peptide / ID: 10 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 4.613678 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #11: Unk4
| Macromolecule | Name: Unk4 / type: protein_or_peptide / ID: 11 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 6.911511 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK) |
+Macromolecule #12: Unk5
| Macromolecule | Name: Unk5 / type: protein_or_peptide / ID: 12 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 9.209344 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) |
+Macromolecule #13: Unk6
| Macromolecule | Name: Unk6 / type: protein_or_peptide / ID: 13 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 48.613012 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #14: uS4m
| Macromolecule | Name: uS4m / type: protein_or_peptide / ID: 14 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 43.497859 KDa |
| Sequence | String: MQRTLRSLAT RCAGAITGST AQTGASGCIK AEAGVSTSAL SGLISQDAIH GLCPRGAAFA SLASSGVSAA AGAGPASQLR AGAPLSCLW LLALAPSRTG LAATASSSSS CGACGSHSST SSPFAPLPAS AAGLRHYAKA AKGGAAAAPA APSGPKRPRT S TTKLYSCR ...String: MQRTLRSLAT RCAGAITGST AQTGASGCIK AEAGVSTSAL SGLISQDAIH GLCPRGAAFA SLASSGVSAA AGAGPASQLR AGAPLSCLW LLALAPSRTG LAATASSSSS CGACGSHSST SSPFAPLPAS AAGLRHYAKA AKGGAAAAPA APSGPKRPRT S TTKLYSCR NDRLRHTHEQ IWPTLQLTEY EQAMFKRNSR LFVVDMGRSL SLRDKFRMGA YEPATAASTG AAAEEAGGAG AG DALVPAA GGASYRRVPY WQARSLLHES NLHLDALGEN PRYLRLRRVG SLFATKLQNV RKLRLLLGFQ RRGFVQKLYE HSL LARGSD RMWKMVCAME ATLPMTVTRM GLAEDVVGAA TAIRNDKIYV NGKQPVMPRK GLLEPGDVVG PAAGGAAYLR KRVA RSMEP LASVVTRDYV UniProtKB: Uncharacterized protein |
+Macromolecule #15: Mitochondrial ribosomal protein S6
| Macromolecule | Name: Mitochondrial ribosomal protein S6 / type: protein_or_peptide / ID: 15 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.661864 KDa |
| Sequence | String: MPLYELLMLA KPSLPRAEMV ALIKGLGDLV YRNGGFVTSL KSFGDQHLAY DLRRPFEKYD RAHIWQMDFV SSVDAIKPLD HGLHVNNTV LRWVLVKRPM SDKSAAAPAA ATQAVEAIAQ PQR UniProtKB: Uncharacterized protein |
+Macromolecule #16: bS16m
| Macromolecule | Name: bS16m / type: protein_or_peptide / ID: 16 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 10.058308 KDa |
| Sequence | String: MEKRQARNSG VRKEDVGWWD PNPAADGNMH LGLNFDRLKY WLTAGAKPTD KVAELLGHAG VLPKVPQPPH YNPRDPKDDT KWRPNEDK UniProtKB: Mitochondrial ribosomal protein S16 |
+Macromolecule #17: bS18m
| Macromolecule | Name: bS18m / type: protein_or_peptide / ID: 17 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 36.901188 KDa |
| Sequence | String: MPACSGLHAH ADQQSSLLQL LAQRKGAYSG RPDINQQSRG FASSAGSDNS SSGQQAAGLE SERLAPTTRL QDLVAGATER IAQKQVGTL GQLPEQLAAE AASLEHQLAS SSGRSGHTHD LDGAGALADR LASAAAAGGP DGRAGGADGA AVTAGQRLLH N KYGYAALG ...String: MPACSGLHAH ADQQSSLLQL LAQRKGAYSG RPDINQQSRG FASSAGSDNS SSGQQAAGLE SERLAPTTRL QDLVAGATER IAQKQVGTL GQLPEQLAAE AASLEHQLAS SSGRSGHTHD LDGAGALADR LASAAAAGGP DGRAGGADGA AVTAGQRLLH N KYGYAALG GVVPDGGEMM AAAGPAAVLR SVTAAQPRLH PRRRFQPGQT YEPQDLNPYS SASPESRQRG VGVAVPRPSV AE VLEKADY KNVAFLTRWF LSPAGRLLPR RQTRLPVAVH KHVSRQVRLA RHMGLIAGEA RLDKTHVQAL REVEAAQLLA GRL VAGAEA GAQQQAAAGA GAGAGAAPRQ RYNELLDFGA UniProtKB: Small ribosomal subunit protein bS18c |
+Macromolecule #18: mS26
| Macromolecule | Name: mS26 / type: protein_or_peptide / ID: 18 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.474727 KDa |
| Sequence | String: MQSGTVQKLL PSAEMLSALL PGLFVAQTRS KTYTSNVMDE TRTKRIIDHF AYHKARKQYR ASLHNLYSSW RAELVAKQLA DPGAQRRLK AEEAREEAAR MAELLREKRV ATLVEQVRKA ELEHKLAQVQ LQRAVRRQEE LHRREEARAA RYQRLLEASR N WIRQEDLE ...String: MQSGTVQKLL PSAEMLSALL PGLFVAQTRS KTYTSNVMDE TRTKRIIDHF AYHKARKQYR ASLHNLYSSW RAELVAKQLA DPGAQRRLK AEEAREEAAR MAELLREKRV ATLVEQVRKA ELEHKLAQVQ LQRAVRRQEE LHRREEARAA RYQRLLEASR N WIRQEDLE AAINRALDNP EPFGFVTSLK INRGF UniProtKB: UNIPROTKB: A8J4B1 |
+Macromolecule #23: bS1m
| Macromolecule | Name: bS1m / type: protein_or_peptide / ID: 23 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.942551 KDa |
| Sequence | String: MRARRSPAIN TLDAQLQQAD ASPASAPSPS SPSSAPLAPS DLDDVLCRHT LRLPKPASTG QEAALRLLSR FNGALPSLRG QTILAKVLS VDSERVTVDT GYNGVAEVPR ADVSIAHVHT ADGAAPVRPT TSDVRPGDLL RLRVDAPYTP YGDMQLTAVR E ESEFKRRL ...String: MRARRSPAIN TLDAQLQQAD ASPASAPSPS SPSSAPLAPS DLDDVLCRHT LRLPKPASTG QEAALRLLSR FNGALPSLRG QTILAKVLS VDSERVTVDT GYNGVAEVPR ADVSIAHVHT ADGAAPVRPT TSDVRPGDLL RLRVDAPYTP YGDMQLTAVR E ESEFKRRL AWGELRRAME AGQPVSGRVL NDCVGGFAVG VAGFVAFLPS AMAARDTTQR VGSLQSFRIV AMDEARRRIT LR DAGVAAA GFRRVG UniProtKB: S1 motif domain-containing protein |
+Macromolecule #24: uS2m
| Macromolecule | Name: uS2m / type: protein_or_peptide / ID: 24 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 53.354316 KDa |
| Sequence | String: MLTRLAALGR PAVLDAAAAA VVVAVEPSAS ASAAWRDGGR DASTSYGVHS LPLAWGAQRR ALGALASGCA STAPTPMVAS AASAAAQAR TGLPFSTAAS TPAPVNAPPP PVNAPHLSNT PAELEEAELA GLDMESLMRA IVHPLPLGRV RNGALLAQAL G TGAPVVQL ...String: MLTRLAALGR PAVLDAAAAA VVVAVEPSAS ASAAWRDGGR DASTSYGVHS LPLAWGAQRR ALGALASGCA STAPTPMVAS AASAAAQAR TGLPFSTAAS TPAPVNAPPP PVNAPHLSNT PAELEEAELA GLDMESLMRA IVHPLPLGRV RNGALLAQAL G TGAPVVQL ENGPGGRDAA LFFDPAATLS STMRALHVIR QVLEKDGHVV VVNSNPRMRP LLREAAHLCL NSNVWFWADD WL PGCLAET DARGGHCPLL SDKAQPNRQI MAEKGLVFRN PLVPDRGNPA AVLGGPAPRL SAADMQRLLA GARSPMGLTH SVY KREQTA SHRGCRKALA ALTAARTAEA RLLLPGSQTG AGRGRFRQPA LVVVLDLSYG GEAVREAAAR NVPTVALLNG HSDA SAVTF PVYASESHLG YHHFFLEWLL RVVNIDPKVL HTLRNAARAA PSAGATGATG ATGATGATGA TGATGATGAT GATGA TGAT GATGATGATG ARAAHAGAGA GAAAAPAAGS GKAAGKAGQQ GGRGKR UniProtKB: Uncharacterized protein |
+Macromolecule #25: S5 DRBM domain-containing protein
| Macromolecule | Name: S5 DRBM domain-containing protein / type: protein_or_peptide / ID: 25 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.876633 KDa |
| Sequence | String: MLRGSILSAQ STGAFSCGAA AVMPSLRSAN SLHGSFAASA AFSSPISTTT SCLDVSQNQE RGLSTEATGE GESEDDPTYL RGIKTQSGL RNSDGDEEPP DRRKAYEFEP GGRGRGGGGG GGGAPFQRRP ADAAAAGTDM LELVATAREL FTRNVDSGSG G GGAGGGGG ...String: MLRGSILSAQ STGAFSCGAA AVMPSLRSAN SLHGSFAASA AFSSPISTTT SCLDVSQNQE RGLSTEATGE GESEDDPTYL RGIKTQSGL RNSDGDEEPP DRRKAYEFEP GGRGRGGGGG GGGAPFQRRP ADAAAAGTDM LELVATAREL FTRNVDSGSG G GGAGGGGG GGGGVGPLPP LPRELREALY GHRWDRVIVD VRRTHTPVRG RGKREEYTAM VATGNMRGVF GLGIGVAESA QL ATARAHL DSLSRLAAVP LYRGHTLYTH VDHTFHRLSI RMMPRPAGWG LRCPDLLYEL CGLVGLRDVS IRVRGQRGSR FFV AQCFQE ALQKQTTPHD GVEATGMYIR EVFRGGPGGR LPCGLRRGVD VM UniProtKB: S5 DRBM domain-containing protein |
+Macromolecule #26: uS15m
| Macromolecule | Name: uS15m / type: protein_or_peptide / ID: 26 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 34.706652 KDa |
| Sequence | String: MAGRLLSRLG LIELGASAAC AARQPWPATT TAALPMAVLS ALHPWRLAWV TPCGALLLST SSSSGGAAGA TEAVSTSAPF VEPSPSTAA GASAGSSSAD IAGPSGDSKP DLVEKLLSSD VLGKGERRRY ERQALIREFQ RHPADTGSPQ VMVAVWTHRV R ELVKHFDA ...String: MAGRLLSRLG LIELGASAAC AARQPWPATT TAALPMAVLS ALHPWRLAWV TPCGALLLST SSSSGGAAGA TEAVSTSAPF VEPSPSTAA GASAGSSSAD IAGPSGDSKP DLVEKLLSSD VLGKGERRRY ERQALIREFQ RHPADTGSPQ VMVAVWTHRV R ELVKHFDA NRKELRPMRE LEIMMNQRRR MLTWLRRADF ESYAYVISKL GLQDIYTSVG ISDRYREGLR PSDPV(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)KDDS VNRLRFNFHK QYKQKKTNLW QRLRPQLLAE DPVLAAQAVA TAAAEKQRQT QQRNATSQRL PAQ UniProtKB: Small ribosomal subunit protein uS15c, Small ribosomal subunit protein uS15c |
+Macromolecule #27: mS34
| Macromolecule | Name: mS34 / type: protein_or_peptide / ID: 27 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.287194 KDa |
| Sequence | String: MAFVELLRSA PKRTLSDLLR KLPNHGVGAI VTRDTWHPES KKYWEVVEVA PSTADPTKLS AWGYQYFKGE RQHPAPKRIA SVWKYGWLL KEQPGAGLSA AVLQQLSERQ AAAQAAGLVA GGSEAAAGTG AAST UniProtKB: UNIPROTKB: A8J254 |
+Macromolecule #28: mS23
| Macromolecule | Name: mS23 / type: protein_or_peptide / ID: 28 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 20.813639 KDa |
| Sequence | String: MEFWRLVVRP TRRANLVAKL MRDLESGHFA APKALDVLTQ YRTEQLSYSL TGIPRVTLPE KPLIRAFLQK YPEARAEPVA LDSFTPPLA RQFAQRQLQL MQAGAAREQA FTQAEQELAG RLQALRSRLL GSAATALSEG AQAAVPGPAA SGVRGMVELL Q QEEQEALD AGLEALASSA QQQQQQLSAN SR UniProtKB: Small ribosomal subunit protein mS23 |
+Macromolecule #29: uS17m
| Macromolecule | Name: uS17m / type: protein_or_peptide / ID: 29 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.409297 KDa |
| Sequence | String: MGGMEFIGRV VSNRMQKTVV VAVSYVVWVP KYKVYQKRVA RHKARDESQA SVIGDIVRIR KSRKFSREVS YSVVDTLRKA HVYDPAAAA ALVATRDAAR AAAAASTGTR AGAAAAGEGQ AAQLEGAGFA ASGVATSSAG GRSDPWVAEA ARRLDASAAR L AALRELYE ...String: MGGMEFIGRV VSNRMQKTVV VAVSYVVWVP KYKVYQKRVA RHKARDESQA SVIGDIVRIR KSRKFSREVS YSVVDTLRKA HVYDPAAAA ALVATRDAAR AAAAASTGTR AGAAAAGEGQ AAQLEGAGFA ASGVATSSAG GRSDPWVAEA ARRLDASAAR L AALRELYE KEVGPGVPLS GAVLAVANSG PGGTGSVRGK AAAAAAGAAE GPGDAKAQQD KQ UniProtKB: Mitochondrial ribosomal protein S17 |
+Macromolecule #30: uS12m
| Macromolecule | Name: uS12m / type: protein_or_peptide / ID: 30 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.082508 KDa |
| Sequence | String: MRVWSEVQAA HMATVGQLLG GARKVAKTRK CRIRQLDGSP FKKGICLRVY TTSPKKPNSA NRKLAKVQLS NGLKALAYIP GEGHNLQEH SMVLVSGNGV RDLPGVKLRI VRGRYDCAGV KDRKKSSED UniProtKB: Ribosomal protein S12, mitochondrial |
+Macromolecule #31: uS8m
| Macromolecule | Name: uS8m / type: protein_or_peptide / ID: 31 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 44.616223 KDa |
| Sequence | String: MAAPLLDPLV SKLRQTTATA ARAAEVMRAA FPGATHETAG RNTIAVQLPR KDVPTYVMAN QRPQPWELLP MKAAAMTQYP NFFNNSCTF FGSIKRDVVN GVPFCLLRPS RLALDMAKVV RNLGIVDGFE VVQRRSRLGA HDFVWLPEQQ PQEPEHLYDT S LFRQRLIR ...String: MAAPLLDPLV SKLRQTTATA ARAAEVMRAA FPGATHETAG RNTIAVQLPR KDVPTYVMAN QRPQPWELLP MKAAAMTQYP NFFNNSCTF FGSIKRDVVN GVPFCLLRPS RLALDMAKVV RNLGIVDGFE VVQRRSRLGA HDFVWLPEQQ PQEPEHLYDT S LFRQRLIR LHLRTDLFSR LPGAPGSGAG GPQPASAQLA PAVGLLPLSV KNISKASQPV LMYPRQLEEA AARLPAGVFM CY HPQLGLI TDAMAQQYDV PALVAAHVGL PLSQAAAIRG ALRVKAAEEA GKELRHVTQL KDWNMMELLR QRMVERRAAL EAG MGVGGE VAARLQELRE AGLRLRDEAS DRVTSALNVA QDLEDGALAW QLVHSRALGA APGAAAAGVD EGAGGEEQAG AGEG RTSPR GQPRRRR UniProtKB: Uncharacterized protein |
+Macromolecule #32: uS11m
| Macromolecule | Name: uS11m / type: protein_or_peptide / ID: 32 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.121113 KDa |
| Sequence | String: MAQASKGLIG ALQASLGGLL STSGRLHGGL GPSAGPAALV SGAWGLSACA AFTRSVATTS AGASATAPAE PGPASTASDA GSGAAQEAE AISSGSNSGG GGFSTGFRKV VSERRAGQLG ALEGIVHIQN TLNNIILTLT DKQGLIKTYT SAGVVGYKGS R KSQPVAAE ...String: MAQASKGLIG ALQASLGGLL STSGRLHGGL GPSAGPAALV SGAWGLSACA AFTRSVATTS AGASATAPAE PGPASTASDA GSGAAQEAE AISSGSNSGG GGFSTGFRKV VSERRAGQLG ALEGIVHIQN TLNNIILTLT DKQGLIKTYT SAGVVGYKGS R KSQPVAAE KAAEELARRA LKLGYSSVQV RLKGAGSNKQ YAVTSLAAAG LTITSLADVT PVPYNGCRLP KKRRV UniProtKB: Mitochondrial ribosomal protein S11 |
+Macromolecule #33: Ribosomal_S10 domain-containing protein
| Macromolecule | Name: Ribosomal_S10 domain-containing protein / type: protein_or_peptide / ID: 33 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 61.617887 KDa |
| Sequence | String: MQPVVLRGIS SLTWQHARGA LPSALQQALA ASAASSSSSL TAVDCDAGAD MQASCSGRGV ECARHRLAWG PLSGVAGLST SARAASPAA AAMPSSSAAS AASPAADAAP PGAYSVEIAV QGFEKRYVDM ACNTLGDIII LAFAPKSYAA LPTGAPSPSD A PVNLAFGP ...String: MQPVVLRGIS SLTWQHARGA LPSALQQALA ASAASSSSSL TAVDCDAGAD MQASCSGRGV ECARHRLAWG PLSGVAGLST SARAASPAA AAMPSSSAAS AASPAADAAP PGAYSVEIAV QGFEKRYVDM ACNTLGDIII LAFAPKSYAA LPTGAPSPSD A PVNLAFGP ERRDVRLPWR RTRITLIRGP HIDKTGMEQF ERREYKSVLT AATNCPQELA RLLEAVKLYQ FTGVQIKMQL AS AQRVDLP ADVSALLRGG GGATAASTGS SSSSSSSSSS SSSLVGSPVV (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)EIPAPA SADLAFAEEV GRVLGVLRPA VWAG LSRRR AVLAGSADHE TWLKAAAGGL PPSASATRGA GRGGGAAALP PPAVVAGWDA EAEVAGAAAE GGDAFGRLLR LRLRQ QAQL QDAQASTSAS GTSTPAAASA AAYGLDPALA ARLLARIDAD LLHAHEAAAA GLPPPAALAA GTAAAAVAAP PPATAV LDA SKMSPAEYAY AVLKYVQYVD SLYESSGAAA AAARGDAALQ LAAPALAVRL LQLWWEATTV EFKRALGLPP AEEERRM LE RLLELQKKKD ELKGAGRGGG R UniProtKB: Small ribosomal subunit protein uS10 domain-containing protein, Small ribosomal subunit protein uS10 domain-containing protein |
+Macromolecule #34: Mitochondrial ribosomal protein S13
| Macromolecule | Name: Mitochondrial ribosomal protein S13 / type: protein_or_peptide / ID: 34 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.758888 KDa |
| Sequence | String: MAQIQRVALF PHATFHIALA RVFGLGKSTS LAVSEACGIS KDLKVKDVKD SYIQKVTAYI EANFVTGDTL KRQVRENVLE EVAINSRRG VRHTLGLPVS SAQTRSNGVN ARKLRPLLLY DTSRR UniProtKB: Uncharacterized protein |
+Macromolecule #35: Mitochondrial ribosomal protein S14
| Macromolecule | Name: Mitochondrial ribosomal protein S14 / type: protein_or_peptide / ID: 35 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.009181 KDa |
| Sequence | String: MKFRLPQYAY IIGTRGNVKD HMRRMLVEQH EVDRSVFKAV TTDKSVPLEH RLQVQRLFET EVPRDSAANR VVNRCVLTGR ARGVHRFAR LSRIMIRQLA HSGFLPGVSK ASW UniProtKB: Uncharacterized protein |
+Macromolecule #36: Plastid-specific ribosomal protein 4
| Macromolecule | Name: Plastid-specific ribosomal protein 4 / type: protein_or_peptide / ID: 36 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.437036 KDa |
| Sequence | String: MNQSTSSLVR LASASFAAIG ECTSNLARQL FLSSTLTQFR YTTDSVHQPV NEVQSSSTGP ASTICGKGDN RTRRGKIFRG TYREQLPSH GPQLHPYPGG PVVQRPGGGS RGAFRYPGEA PPTAPPGASH TVPPAQR UniProtKB: Uncharacterized protein |
+Macromolecule #37: uS19m
| Macromolecule | Name: uS19m / type: protein_or_peptide / ID: 37 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.951077 KDa |
| Sequence | String: MPRSTWKGPY VAVSLLQEVV ALARKHPNWW NKGRYIGQKA PEVINTYSRA SVILPDFIAC RFGVHNGKSF VGMEVQEAMV GHRLGEFAP TKATVQHKAK EVNTQKKKIN PKTGKAA UniProtKB: Small ribosomal subunit protein uS19c |
+Macromolecule #38: uS7m,Ribosomal_S7 domain-containing protein,Ribosomal_S7 domain-c...
| Macromolecule | Name: uS7m,Ribosomal_S7 domain-containing protein,Ribosomal_S7 domain-containing protein,Ribosomal_S7 domain-containing protein type: protein_or_peptide / ID: 38 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 32.755266 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)MLSSA RRLLATVAEQ RALAKADAPA AAAQLRARA GLASVSCCGS TTATITTRLG ATGSWRHMSS AADSTSSSSS SSSSQQQQPP AQQQQQPEAS TSSAAASSPD G PMSEVVQQ LTKASPTGRL LDLFTPSQQQ IQTLQRSRGP GRGSRSGTGP AGPAAGTGVQ AAEAAAPLTA DSDVLEVLMR AV DNCKPLM KVIQSKAGTR VVYQPRPLNP DQSTNFAVKW ILQAAQKRRA GAKGVAGAAS MAEALAVELL LAAQRKGGAR ARR DEVHKL ALDNRANLRR UniProtKB: Small ribosomal subunit protein uS7 domain-containing protein |
+Macromolecule #39: 30S ribosomal protein S9, chloroplastic
| Macromolecule | Name: 30S ribosomal protein S9, chloroplastic / type: protein_or_peptide / ID: 39 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 45.610211 KDa |
| Sequence | String: MVRGGVATVG PSSLALAASA LLGTLPQQTA SHSGASLWVF GSRHLAAPAA AGGASNQDRT QPSAKPSAAE PELKLNFSLR PSVLSPLFP AWEQTRQINQ RREHLMALLD AHQQLAEGLR ITKDGVAAST DGADAGTTSS FAAGGVRAGV TRSARRRARN L QQVASQLR ...String: MVRGGVATVG PSSLALAASA LLGTLPQQTA SHSGASLWVF GSRHLAAPAA AGGASNQDRT QPSAKPSAAE PELKLNFSLR PSVLSPLFP AWEQTRQINQ RREHLMALLD AHQQLAEGLR ITKDGVAAST DGADAGTTSS FAAGGVRAGV TRSARRRARN L QQVASQLR ALLDATPGHP RGVTGLPTAC PSPAHAVAYC GALEAALASP DAASTGARDF GAVARGDVSP WTHYAKHFAG RY AGAPQLQ PLEVIYSRPD PAEVVRQLRR AAREQEALGI SSSGGGSGEA GSGLAFPATS TSPEPAGPHI DLTGVTHARG KRK ASAASV QLVPAGGAGI TVNGVPLSSY FRDPACREKV LAPLALAGAE TAYDVAARVK GGGMMGQAQA IATALARALC VQQP SLGAT AGDGGNGEGP LAAMTRWDGR VVERQKPGQA GSRRGFQWVK R UniProtKB: Small ribosomal subunit protein uS9c |
+Macromolecule #40: mS29
| Macromolecule | Name: mS29 / type: protein_or_peptide / ID: 40 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 49.635844 KDa |
| Sequence | String: MTSALLVASR RARQAQGLPR CLLHAIGIHA GTRAEFASVA LQEAGTTPST SGQEQPSSAQ LTPAHLRSYY PLNLALLPEA ARGSAGAFY TPRDPGHERR GGCKALQQEM EATGRASILY RPIMAALNGA VAAGQQPRLL LTGPAGCGKS LALLGLVEWA R QQGWLVVY ...String: MTSALLVASR RARQAQGLPR CLLHAIGIHA GTRAEFASVA LQEAGTTPST SGQEQPSSAQ LTPAHLRSYY PLNLALLPEA ARGSAGAFY TPRDPGHERR GGCKALQQEM EATGRASILY RPIMAALNGA VAAGQQPRLL LTGPAGCGKS LALLGLVEWA R QQGWLVVY VPSCLALVRG GYFARRGRGA AGGWDTLTSA QQLLKGVMDA HGPLLQSLPV LPVPGRAARR QQQQQHEPRQ AD KPAKVEE GQGQGQGQAG GSGLLEEGSG SASGAGGGGG RTLQDVALRG LSSDDNAQLA VDSALQLIRQ LQLLGSGAAQ PPD SQPGQP PRVLFALDDY NYLYGPTDYG VQPPSASPLQ GRRRVLDAGE LILARGLRLL ESELGTNPVA AAAAGGGAGG VGGA VVVAA TTATPALPAP RSLALEVPHT VVEVPGFDEA ETAAALAHYA ATGAATRAAS AAEARHLFAL TGGNGRELRA KAGAL GVRV G UniProtKB: Small ribosomal subunit protein mS29 |
+Macromolecule #41: Ribosomal_S3_C domain-containing protein
| Macromolecule | Name: Ribosomal_S3_C domain-containing protein / type: protein_or_peptide / ID: 41 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 50.704199 KDa |
| Sequence | String: MDDMVLVLAR NLSGKAKGKP APKGAPAPPP PPVASSAVTA NPATRSAGAR IRRNQPRLDP LTLLTPQQRE LAESLAHLSR QAAQPPSAT AAGADALTPA AEAAADAAAE AAAAPLRGLG EVPLPLRRRE VAALVDAALT GAAGGQQVHF QPFVFNDIFQ S APAAATYV ...String: MDDMVLVLAR NLSGKAKGKP APKGAPAPPP PPVASSAVTA NPATRSAGAR IRRNQPRLDP LTLLTPQQRE LAESLAHLSR QAAQPPSAT AAGADALTPA AEAAADAAAE AAAAPLRGLG EVPLPLRRRE VAALVDAALT GAAGGQQVHF QPFVFNDIFQ S APAAATYV AEALESGSAW GRIERFFLAA AEADAGKLVA AVRVEVAGRI GRKADMAATK SWAWGDLRLA SLTSKLDYGT AA ANTRMGV LGIKVWIRYQ PAAAPDLYFP ATGGSGLHAP LPAGAAPSMS LDELYAAALE RSGSSSSSSS SSSVGGAGGS STR YGAVGW WSRPAGQQPA ANRVHEVFTP EFNPKAGDRL DGYLRRAAES RRVFRSAANR EEYMATVGRF LVKQRQAAKA RAAA EAYAA ALSATKPATI PTTSSSSSSS SANSSAASSV PARPRLGRRP HRPLARFPVE AFSPLGRRGP VLLRWRLQAA AAVGK GKGG NGERRA UniProtKB: Small ribosomal subunit protein uS3 C-terminal domain-containing protein |
+Macromolecule #42: mS33
| Macromolecule | Name: mS33 / type: protein_or_peptide / ID: 42 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.289773 KDa |
| Sequence | String: MSASTLSRLL PAGSLAWGLP GVLLRSASLL VHHQPCAPST SGRVQESQLQ WSCYYATSHG TGAPKAASTQ SSSTTSAASA AASATSAAR PAAVSSEAGG DGASSAPFYA GLAARYPDPD TVPDHVMHRL RNTLFGRAVR PRETTGRRAL ARPLQGKALT D WYWMPPNE ...String: MSASTLSRLL PAGSLAWGLP GVLLRSASLL VHHQPCAPST SGRVQESQLQ WSCYYATSHG TGAPKAASTQ SSSTTSAASA AASATSAAR PAAVSSEAGG DGASSAPFYA GLAARYPDPD TVPDHVMHRL RNTLFGRAVR PRETTGRRAL ARPLQGKALT D WYWMPPNE SPGFHSEEDE YELRRALNRR HTKEAEAEAA DTGADKKKKR K UniProtKB: Uncharacterized protein |
+Macromolecule #19: S1 rRNA
| Macromolecule | Name: S1 rRNA / type: rna / ID: 19 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 32.590293 KDa |
| Sequence | String: AAAUUAGAGU UUGGUGCUGG CUCAGCUUUC AUGCGCCGAG CGGUGGUUUA UACACAUGCG AGCUUAAAAG AUUAAGGCCA CGAGUGCGU AAUACGCGAG UU GENBANK: GENBANK: X54860.1 |
+Macromolecule #20: S4 rRNA
| Macromolecule | Name: S4 rRNA / type: rna / ID: 20 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 133.231078 KDa |
| Sequence | String: AGCUCUUGCA UUGCUGAAUU UUUUUUUUUU UUUUUUUUUU AAAAAAAAAA AAAAUUUUUU UUUUUCAAGU CAUCAUGGGG CUUAUAGAG UGGGCUACAG GCGUAUUACA UUGGACACCC ACAAGUUGCC AAAACUGUCC GAAUAUACGG AUUGGAGUAA A AAAAAAAA ...String: AGCUCUUGCA UUGCUGAAUU UUUUUUUUUU UUUUUUUUUU AAAAAAAAAA AAAAUUUUUU UUUUUCAAGU CAUCAUGGGG CUUAUAGAG UGGGCUACAG GCGUAUUACA UUGGACACCC ACAAGUUGCC AAAACUGUCC GAAUAUACGG AUUGGAGUAA A AAAAAAAA AUUUUUUAAA AUUUUUUUUU UUUUUUGCUG AAACUAGCCU CCAUGAAGAA GGAAUCGCGA GUAAUCGUAG AU CAUUAGC GCUACGGUGA AGGUAACCUC UAUUGUGCAC ACAUUGCCCG UCACCUCCGA UAAUAGUAUU GUACAGGAAG AAC UAUGGC UACACUUAGU CGCGGCCUGG AACGUAUGCG UGAUAUUAGA GUUGGAGUAA GUCGUAACAG GUUGGGGUAG GGGA ACCUG CUCCAGAGUC |
+Macromolecule #21: S2 rRNA
| Macromolecule | Name: S2 rRNA / type: rna / ID: 21 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 68.527445 KDa |
| Sequence | String: UGCUAGGUGA CCCAAUACGG GCAUCGGGCA AAACUCGCGU AGGAUUAGCG AGCUGGGUCG CUUUUUUUUU UCAGCGGCCC AUGGCUUAU CCUUAGCCUG UCUUAACGGU ACUAGGCCAC GGUGGCACUG AAAAGGGGCC ACGGUUCUUA UGAACCCAGC A GUGUUGAA ...String: UGCUAGGUGA CCCAAUACGG GCAUCGGGCA AAACUCGCGU AGGAUUAGCG AGCUGGGUCG CUUUUUUUUU UCAGCGGCCC AUGGCUUAU CCUUAGCCUG UCUUAACGGU ACUAGGCCAC GGUGGCACUG AAAAGGGGCC ACGGUUCUUA UGAACCCAGC A GUGUUGAA UUUUGGACAA UCGCUUGAAC GGCGGAUCCA GAUGCUGCUU ACAAA GENBANK: GENBANK: X54860.1 |
+Macromolecule #22: S3 rRNA
| Macromolecule | Name: S3 rRNA / type: rna / ID: 22 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 126.928117 KDa |
| Sequence | String: AUUGUUUGAA CACCCCCCAA GCACGUGCCA GAAGGGUCGG UAAAACGUGC GGUGUCAGUA UAAAGCGUCU UGACUAGGCA GGCAGCGCG UCUGAGCGUG UGAACACUUU AAACGUUGGG UAAUAUUCGG AGGAUCGGUC AAAUGAGAAU AUUCCGGAUG G AAAGCCGA ...String: AUUGUUUGAA CACCCCCCAA GCACGUGCCA GAAGGGUCGG UAAAACGUGC GGUGUCAGUA UAAAGCGUCU UGACUAGGCA GGCAGCGCG UCUGAGCGUG UGAACACUUU AAACGUUGGG UAAUAUUCGG AGGAUCGGUC AAAUGAGAAU AUUCCGGAUG G AAAGCCGA AGGCGAAAGC ACCAACAUCA GAGUCACUAA AGCUUCAACG CGUAAGUUUG GGUAGCGAAC CGGAUUAGAG AC CCGGGUA GUCCAAACCG UCAACACAUU AUAGUAAUCU AUAACGCCUG GUGAUACGGU GGCAACACUA UAAAUCAAAG CAA UUGGCA GCGAUAGAGA UGCGCGGUGG AAUAUGCUGU UUAAAUCGAA UUUACGCGCA AAAUCUUACC ACUUUUU GENBANK: GENBANK: X54860.1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 7.6 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Number classes used: 1 / Resolution.type: BY AUTHOR / Resolution: 4.2 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 40131 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT / Software - Name: RELION |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION |
| Final 3D classification | Number classes: 6 / Software - Name: RELION |
+ About Yorodumi
About Yorodumi
-News
-Feb 9, 2022. New format data for meta-information of EMDB entries
New format data for meta-information of EMDB entries
- Version 3 of the EMDB header file is now the official format.
- The previous official version 1.9 will be removed from the archive.
Related info.: EMDB header
EMDB header
External links: wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
-Aug 12, 2020. Covid-19 info
Covid-19 info

URL: https://pdbj.org/emnavi/covid19.php
New page: Covid-19 featured information page in EM Navigator.
Related info.: Covid-19 info /
Covid-19 info /  Mar 5, 2020. Novel coronavirus structure data
Mar 5, 2020. Novel coronavirus structure data
+Mar 5, 2020. Novel coronavirus structure data
Novel coronavirus structure data
- International Committee on Taxonomy of Viruses (ICTV) defined the short name of the 2019 coronavirus as "SARS-CoV-2".
- In the structure databanks used in Yorodumi, some data are registered as the other names, "COVID-19 virus" and "2019-nCoV". Here are the details of the virus and the list of structure data.
Related info.: Yorodumi Speices /
Yorodumi Speices /  Aug 12, 2020. Covid-19 info
Aug 12, 2020. Covid-19 info
External links: COVID-19 featured content - PDBj /
COVID-19 featured content - PDBj /  Molecule of the Month (242):Coronavirus Proteases
Molecule of the Month (242):Coronavirus Proteases
+Jan 31, 2019. EMDB accession codes are about to change! (news from PDBe EMDB page)
EMDB accession codes are about to change! (news from PDBe EMDB page)
- The allocation of 4 digits for EMDB accession codes will soon come to an end. Whilst these codes will remain in use, new EMDB accession codes will include an additional digit and will expand incrementally as the available range of codes is exhausted. The current 4-digit format prefixed with “EMD-” (i.e. EMD-XXXX) will advance to a 5-digit format (i.e. EMD-XXXXX), and so on. It is currently estimated that the 4-digit codes will be depleted around Spring 2019, at which point the 5-digit format will come into force.
- The EM Navigator/Yorodumi systems omit the EMD- prefix.
Related info.: Q: What is EMD? /
Q: What is EMD? /  ID/Accession-code notation in Yorodumi/EM Navigator
ID/Accession-code notation in Yorodumi/EM Navigator
External links: EMDB Accession Codes are Changing Soon! /
EMDB Accession Codes are Changing Soon! /  Contact to PDBj
Contact to PDBj
+Jul 12, 2017. Major update of PDB
Major update of PDB
- wwPDB released updated PDB data conforming to the new PDBx/mmCIF dictionary.
- This is a major update changing the version number from 4 to 5, and with Remediation, in which all the entries are updated.
- In this update, many items about electron microscopy experimental information are reorganized (e.g. em_software).
- Now, EM Navigator and Yorodumi are based on the updated data.
External links: wwPDB Remediation /
wwPDB Remediation /  Enriched Model Files Conforming to OneDep Data Standards Now Available in the PDB FTP Archive
Enriched Model Files Conforming to OneDep Data Standards Now Available in the PDB FTP Archive
-Yorodumi
Thousand views of thousand structures
- Yorodumi is a browser for structure data from EMDB, PDB, SASBDB, etc.
- This page is also the successor to EM Navigator detail page, and also detail information page/front-end page for Omokage search.
- The word "yorodu" (or yorozu) is an old Japanese word meaning "ten thousand". "mi" (miru) is to see.
Related info.: EMDB /
EMDB /  PDB /
PDB /  SASBDB /
SASBDB /  Comparison of 3 databanks /
Comparison of 3 databanks /  Yorodumi Search /
Yorodumi Search /  Aug 31, 2016. New EM Navigator & Yorodumi /
Aug 31, 2016. New EM Navigator & Yorodumi /  Yorodumi Papers /
Yorodumi Papers /  Jmol/JSmol /
Jmol/JSmol /  Function and homology information /
Function and homology information /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)