-検索条件
-検索結果
検索 (著者・登録者: fernandez & c)の結果582件中、1から50件目までを表示しています

EMDB-44372: 
In-cell Saccharomyces cerevisiae nuclear pore complex with single nuclear ring

EMDB-44377: 
In-cell Saccharomyces cerevisiae nuclear pore complex with double nuclear ring and basket

EMDB-44379: 
In-cell Mus musculus nuclear pore complex with nuclear basket

EMDB-44381: 
In-cell Toxoplasma gondii nuclear pore complex

EMDB-45197: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with double nuclear ring and basket

EMDB-45198: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with single nuclear ring

EMDB-45199: 
In-cell Saccharomyces cerevisiae nuclear pore complex cytoplasmic ring focused refinement

EMDB-45200: 
In-cell Saccharomyces cerevisiae nuclear pore complex inner ring focused refinement

EMDB-45201: 
In-cell Saccharomyces cerevisiae nuclear pore complex single nuclear ring focused refinement

EMDB-45202: 
In-cell Saccharomyces cerevisiae nuclear pore complex double nuclear ring focused refinement

EMDB-45203: 
In-cell Saccharomyces cerevisiae nuclear pore complex nuclear basket focused refinement

EMDB-45204: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for single nuclear ring

EMDB-45205: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for double nuclear ring

EMDB-45216: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map

EMDB-45219: 
In-cell Mus musculus nuclear pore complex cytoplasmic ring focused refinement

EMDB-45220: 
In-cell Mus musculus nuclear pore complex inner ring focused refinement

EMDB-45222: 
In-cell Mus musculus nuclear pore complex nuclear ring focused refinement

EMDB-45223: 
In-cell Mus musculus nuclear pore complex basket focused refinement

EMDB-45227: 
In-cell Mus musculus nuclear pore complex membrane focused refinement

EMDB-45228: 
In-cell Toxoplasma gondii symmetry-expanded nuclear pore complex
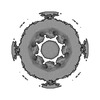
EMDB-45255: 
In-cell Saccharomyces cerevisiae C8-symmetrised nuclear pore complex consensus map

EMDB-45256: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex consensus map

EMDB-45257: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map

EMDB-45258: 
In-cell Mus musculus symmetry-expanded nuclear pore complex with nuclear basket consensus map

EMDB-45259: 
In-cell Toxoplasma gondii C8-symmetrised nuclear pore complex consensus map
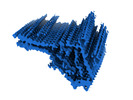

EMDB-50272: 
Additional cryo-EM structure of cardiac amyloid AL59 - mixed polymorph

EMDB-50271: 
Additional cryo-EM structure of cardiac amyloid AL59 - bent polymorph

EMDB-51031: 
Cryo-electron tomogram of AL59 amyloids interacting with collagen VI

EMDB-51032: 
Cryo-electron tomogram of AL59 amyloids interacting with collagen VI

EMDB-51033: 
Cryo-electron tomogram of AL59 amyloids interacting with collagen VI

EMDB-51038: 
Cryo-electron tomogram of AL59 amyloids interacting with collagen VI

EMDB-18689: 
Maps of Collagen VI half- and full-beads

EMDB-50270: 
Cryo-EM structure of cardiac collagen-associated amyloid AL59

EMDB-17418: 
CryoEM structure of a C7-symmetrical GroEL7-GroES7 cage in presence of ADP-BeFx

EMDB-17420: 
CryoEM structure of a GroEL7-GroES7 cage with encapsulated disordered substrate MetK in the presence of ADP-BeFx

EMDB-17421: 
CryoEM structure of a GroEL7-GroES7 cage with encapsulated ordered substrate MetK in the presence of ADP-BeFx

EMDB-17422: 
Density for MetK encapsulated in the GroEL7-GroES7 cage

EMDB-17423: 
Symmetry-averaged GroEL7-GroES7 chamber with encapsulated disordered substrate MetK obtained by in vitro cryo electron tomography
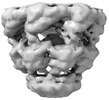
EMDB-17424: 
Symmetry-averaged GroEL7-GroES7 chamber with encapsulated ordered substrate MetK obtained by in vitro cryo electron tomography

EMDB-17425: 
Structure average of GroEL14 complexes found in the cytosol of Escherichia coli overexpressing GroEL obtained by cryo electron tomography

EMDB-17426: 
In situ structure average of GroEL14-GroES14 complexes in Escherichia coli cytosol obtained by cryo electron tomography

EMDB-17534: 
Cryo-EM structure of a D7-symmetrical GroEL14-GroES14 complex in presence of ADP-BeFx

EMDB-17535: 
Cryo-EM structure of a C7-symmetrical GroEL14-GroES7 complex in presence of ADP-BeFx

EMDB-17559: 
In situ structure average of GroEL7-GroES7 chamber with no or disordered substrate in Escherichia coli cytosol obtained by cryo electron tomography

EMDB-17560: 
In situ structure average of GroEL7-GroES7 chamber with encapsulated, ordered substrate in Escherichia coli cytosol obtained by cryo electron tomography
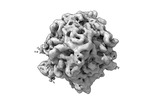
EMDB-17561: 
Cryo-ET subtomogram of 70S ribosomes in Escherichia coli cells at 37 and 46 degrees centigrade and in Escherichia coli cells overexpressing GroELS and MetK

EMDB-17562: 
Cryo-ET subtomogram of 70S ribosomes in Escherichia coli cells overexpressing GroEL

EMDB-17563: 
CryoEM structure of a GroEL14-GroES7 cage with encapsulated ordered substrate MetK in the presence of ADP-BeFx

EMDB-17564: 
CryoEM structure of a GroEL14-GroES7 cage with encapsulated disordered substrate MetK in the presence of ADP-BeFx

EMDB-17565: 
CryoEM structure of a GroEL14-GroES14 cage with two encapsulated disordered MetK substrates in the presence of ADP-BeFx
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
